Abstract
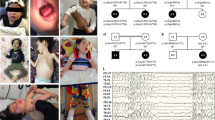
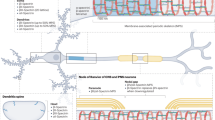
SPTBN1 encodes βII-spectrin, the ubiquitously expressed β-spectrin that forms micrometer-scale networks associated with plasma membranes. Mice deficient in neuronal βII-spectrin have defects in cortical organization, developmental delay and behavioral deficiencies. These phenotypes, while less severe, are observed in haploinsufficient animals, suggesting that individuals carrying heterozygous SPTBN1 variants may also show measurable compromise of neural development and function. Here we identify heterozygous SPTBN1 variants in 29 individuals with developmental, language and motor delays; mild to severe intellectual disability; autistic features; seizures; behavioral and movement abnormalities; hypotonia; and variable dysmorphic facial features. We show that these SPTBN1 variants lead to effects that affect βII-spectrin stability, disrupt binding to key molecular partners, and disturb cytoskeleton organization and dynamics. Our studies define SPTBN1 variants as the genetic basis of a neurodevelopmental syndrome, expand the set of spectrinopathies affecting the brain and underscore the critical role of βII-spectrin in the central nervous system.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout







Similar content being viewed by others
Data availability
The whole-genome and -exome sequencing or transcriptomic data will not be made publicly available as they contain information that could compromise research participant privacy/consent. Source data are provided with this paper. Information on the DNA- and RNA-sequencing raw data and other analyses supporting the findings of this study is available from the corresponding authors upon request.
References
Bennett, V. & Lorenzo, D. N. Spectrin- and ankyrin-based membrane domains and the evolution of vertebrates. Curr. Top. Membr. 72, 1–37 (2013).
Bennett, V. & Lorenzo, D. N. An adaptable spectrin/ankyrin-based mechanism for long-range organization of plasma membranes in vertebrate tissues. Curr. Top. Membr. 77, 143–184 (2016).
Lorenzo, D. N. Cargo hold and delivery: ankyrins, spectrins, and their functional patterning of neurons. Cytoskeleton 77, 129–148 (2020).
Ikeda, Y. et al. Spectrin mutations cause spinocerebellar ataxia type 5. Nat. Genet. 38, 184–190 (2006).
Saitsu, H. et al. Dominant-negative mutations in alpha-II spectrin cause West syndrome with severe cerebral hypomyelination, spastic quadriplegia, and developmental delay. Am. J. Hum. Genet. 86, 881–891 (2010).
Wang, C. C. et al. βIV spectrinopathies cause profound intellectual disability, congenital hypotonia, and motor axonal neuropathy. Am. J. Hum. Genet. 102, 1158–1168 (2018).
Jacob, F. D., Ho, E. S., Martinez-Ojeda, M., Darras, B. T. & Khwaja, O. S. Case of infantile onset spinocerebellar ataxia type 5. J. Child Neurol. 28, 1292–1295 (2013).
Parolin Schnekenberg, R. et al. De novo point mutations in patients diagnosed with ataxic cerebral palsy. Brain 138, 1817–1832 (2015).
Nuovo, S. et al. Between SCA5 and SCAR14: delineation of the SPTBN2 p.R480W-associated phenotype. Eur. J. Hum. Genet. 26, 928–929 (2018).
Nicita, F. et al. Heterozygous missense variants of SPTBN2 are a frequent cause of congenital cerebellar ataxia. Clin. Genet. 96, 169–175 (2019).
Mizuno, T. et al. Infantile-onset spinocerebellar ataxia type 5 associated with a novel SPTBN2 mutation: a case report. Brain Dev. 41, 630–633 (2019).
Accogli, A. et al. Heterozygous missense pathogenic variants within the second spectrin repeat of SPTBN2 lead to infantile-onset cerebellar ataxia. J. Child Neurol. 35, 106–110 (2019).
Lise, S. et al. Recessive mutations in SPTBN2 implicate β-III spectrin in both cognitive and motor development. PLoS Genet. 8, e1003074 (2012).
Yıldız Bölükbaşı, E. et al. Progressive SCAR14 with unclear speech, developmental delay, tremor, and behavioral problems caused by a homozygous deletion of the SPTBN2 pleckstrin homology domain. Am. J. Med. Genet. A 173, 2494–2499 (2017).
Al-Muhaizea, M. et al. A novel homozygous mutation in SPTBN2 leads to spinocerebellar ataxia in a consanguineous family: report of a new infantile-onset case and brief review of the literature. Cerebellum 17, 276–285 (2018).
Writzl, K. et al. Early onset West syndrome with severe hypomyelination and coloboma-like optic discs in a girl with SPTAN1 mutation. Epilepsia 53, e106–e110 (2012).
Hamdan, F. F. et al. Identification of a novel in-frame de novo mutation in SPTAN1 in intellectual disability and pontocerebellar atrophy. Eur. J. Hum. Genet. 20, 796–800 (2012).
Nonoda, Y. et al. Progressive diffuse brain atrophy in West syndrome with marked hypomyalination due to SPTAN1 gene mutation. Brain Dev. 35, 280–283 (2013).
Tohyama, J. et al. SPTAN1 encephalopathy: distinct phenotypes and genotypes. J. Hum. Genet. 60, 167–173 (2015).
Syrbe, S. et al. Delineating SPTAN1 associated phenotypes: from isolated epilepsy to encephalopathy with progressive brain atrophy. Brain 140, 2322–2336 (2017).
Beijer, D. et al. Nonsense mutations in alpha-II spectrin in three families with juvenile onset hereditary motor neuropathy. Brain 142, 2605–2616 (2019).
Knierim, E. et al. A recessive mutation in beta-IV-spectrin (SPTBN4) associates with congenital myopathy, neuropathy, and central deafness. Hum. Genet. 136, 903–910 (2017).
Häusler, M. G. et al. A novel homozygous splice-site mutation in the SPTBN4 gene causes axonal neuropathy without intellectual disability. Eur. J. Med. Genet. 63, 103826 (2020).
Xu, K., Zhong, G. & Zhuang, X. Actin, spectrin, and associated proteins form a periodic cytoskeletal structure in axons. Science 339, 452–456 (2013).
Cheney, R., Hirokawa, N., Levine, J. & Willard, M. Intracellular movement of fodrin. Cell Motil. 3, 649–655 (1983).
Lorenzo, D. N. et al. βII-spectrin promotes mouse brain connectivity through stabilizing axonal plasma membranes and enabling axonal organelle transport. Proc. Natl Acad. Sci. USA 116, 15686–15695 (2019).
Jaganathan, K. et al. Predicting splicing from primary sequence with deep learning. Cell 176, 535–548 (2019).
Karczewski, K. J. et al. The mutational constraint spectrum quantified from variation in 141,456 humans. Nature 581, 434–443 (2020).
Huang, N., Lee, I., Marcotte, E. M. & Hurles, M. E. Characterising and predicting haploinsufficiency in the human genome. PLoS Genet. 6, e1001154 (2010).
Zech, M. et al. Monogenic variants in dystonia: an exome-wide sequencing study. Lancet Neurol. 19, 908–918 (2020).
Willsey, A. J. et al. De novo coding variants are strongly associated with Tourette disorder. Neuron 94, 486–499 (2017).
Firth, H. V. et al. DECIPHER: database of chromosomal imbalance and phenotype in humans using ensembl resources. Am. J. Hum. Genet. 84, 524–533 (2009).
Iossifov, I. et al. The contribution of de novo coding mutations to autism spectrum disorder. Nature 515, 216–221 (2014).
Schultz-Rogers, L. et al. Haploinsufficiency as a disease mechanism in GNB1-associated neurodevelopmental disorder. Mol. Genet. Genomic Med. 8, e1477 (2020).
Richards, S. et al. Standards and guidelines for the interpretation of sequence variants: a joint consensus recommendation of the American College of Medical Genetics and Genomics and the Association for Molecular Pathology. Genet. Med. 17, 405–424 (2015).
Abou Tayoun, A. N. et al. Recommendations for interpreting the loss of function PVS1 ACMG/AMP variant criterion. Hum. Mutat. 39, 1517–1524 (2018).
Strande, N. T. et al. Evaluating the clinical validity of gene-disease associations: an evidence-based framework developed by the Clinical Genome Resource. Am. J. Hum. Genet. 100, 895–906 (2017).
Brnich, S. E. et al. Recommendations for application of the functional evidence PS3/BS3 criterion using the ACMG/AMP sequence variant interpretation framework. Genome Med. 12, 3 (2019).
Molina, S. G., Beltran, A. A. & Beltran, A. S. Generation of an integration-free induced pluripotent stem cell line (UNC001-A) from blood of a healthy individual. Stem Cell Res. 49, 102015 (2020).
Speicher, D. W., Weglarz, L. & DeSilva, T. M. Properties of human red cell spectrin heterodimer (side-to-side) assembly and identification of an essential nucleation site. J. Biol. Chem. 267, 14775–14782 (1992).
Li, X. & Bennett, V. Identification of the spectrin subunit and domains required for formation of spectrin/adducin/actin complexes. J. Biol. Chem. 271, 15695–15702 (1996).
Davis, L. et al. Localization and structure of the ankyrin-binding site on β2-spectrin. J. Biol. Chem. 284, 6982–6987 (2009).
Bignone, P. A. & Baines, A. J. Spectrin alpha II and beta II isoforms interact with high affinity at the tetramerization site. Biochem. J. 374, 613–624 (2003).
Hyvönen, M. et al. Structure of the binding site for inositol phosphates in a PH domain. EMBO J. 14, 4676–4685 (1995).
Korenbaum, E. & Rivero, F. Calponin homology domains at a glance. J. Cell Sci. 115, 3543–3545 (2002).
Yin, L. M., Schnoor, M. & Jun, C. D. Structural characteristics, binding partners and related diseases of the calponin homology (CH) domain. Front. Cell. Dev. Biol. 8, 342 (2020).
Keep, N. H. et al. Crystal structure of the actin-binding region of utrophin reveals a head-to-tail dimer. Structure 7, 1539–1546 (1999).
Bañuelos, S., Saraste, M. & Djinović Carugo, K. Structural comparisons of calponin homology domains: implications for actin binding. Structure 6, 1419–1431 (1998).
Avery, A. W. et al. Structural basis for high-affinity actin binding revealed by a β-III-spectrin SCA5 missense mutation. Nat. Commun. 8, 1350 (2017).
Vajda, S. et al. New additions to the ClusPro server motivated by CAPRI. Proteins 85, 435–444 (2017).
Kozakov, D. et al. The ClusPro web server for protein-protein docking. Nat. Protoc. 12, 255–278 (2017).
Galiano, M. R. et al. A distal axonal cytoskeleton forms an intra-axonal boundary that controls axon initial segment assembly. Cell 149, 1125–1139 (2012).
Sleigh, J. N., Rossor, A. M., Fellows, A. D., Tosolini, A. P. & Schiavo, G. Axonal transport and neurological disease. Nat. Rev. Neurol. 15, 691–703 (2019).
Susuki, K. et al. Glial βII spectrin contributes to paranode formation and maintenance. J. Neurosci. 38, 6063–6075 (2018).
Gobius, I. et al. Astroglial-mediated remodeling of the interhemispheric midline is required for the formation of the corpus callosum. Cell Rep. 17, 735–747 (2016).
Goebbels, S. et al. Genetic targeting of principal neurons in neocortex and hippocampus of NEX-Cre mice. Genesis 44, 611–621 (2006).
Fame, R. M., MacDonald, J. L. & Macklis, J. D. Development, specification, and diversity of callosal projection neurons. Trends Neurosci. 34, 41–50 (2011).
Satterstrom, F. K. et al. Autism spectrum disorder and attention deficit hyperactivity disorder have a similar burden of rare protein-truncating variants. Nat. Neurosci. 22, 1961–1965 (2019).
Yang, R. et al. ANK2 autism mutation targeting giant ankyrin-B promotes axon branching and ectopic connectivity. Proc. Natl Acad. Sci. USA 116, 15262–15271 (2019).
Lorenzo, D. N. et al. A PIK3C3–ankyrin-B–dynactin pathway promotes axonal growth and multiorganelle transport. J. Cell Biol. 207, 735–752 (2014).
Zhong, G. et al. Developmental mechanism of the periodic membrane skeleton in axons. eLife 3, e04581 (2014).
Zhou, R., Han, B., Xia, C. & Zhuang, X. Membrane-associated periodic skeleton is a signaling platform for RTK transactivation in neurons. Science 365, 929–934 (2019).
Lek, M. et al. Analysis of protein-coding genetic variation in 60,706 humans. Nature 536, 285–291 (2016).
Park, J. et al. Mutational characteristics of ANK1 and SPTB genes in hereditary spherocytosis. Clin. Genet. 90, 69–78 (2016).
Baek, H. J. et al. Transforming growth factor-β adaptor, β2-spectrin, modulates cyclin dependent kinase 4 to reduce development of hepatocellular cancer. Hepatology 53, 1676–1684 (2011).
Derbala, M. H., Guo, A. S., Mohler, P. J. & Smith, S. A. The role of βII spectrin in cardiac health and disease. Life Sci. 192, 278–285 (2018).
Sobreira, N., Schiettecatte, F., Valle, D. & Hamosh, A. GeneMatcher: a matching tool for connecting investigators with an interest in the same gene. Hum. Mutat. 36, 928–930 (2015).
Zepeda-Mendoza, C. J. et al. An intragenic duplication of TRPS1 leading to abnormal transcripts and causing trichorhinophalangeal syndrome type I. Cold Spring Harb. Mol. Case Stud. 5, a004655 (2019).
Kalari, K. R. et al. MAP-RSeq: Mayo analysis pipeline for RNA sequencing. BMC Bioinformatics 15, 224 (2014).
Kim, D. et al. TopHat2: accurate alignment of transcriptomes in the presence of insertions, deletions and gene fusions. Genome Biol. 14, R36 (2013).
Langmead, B. Aligning short sequencing reads with Bowtie. Curr. Protoc. Bioinformatics 11, 7 (2010).
Anders, S., Pyl, P. T. & Huber, W. HTSeq—a Python framework to work with high-throughput sequencing data. Bioinformatics 31, 166–169 (2015).
Quinlan, A. R. BEDTools: the Swiss-army tool for genome feature analysis. Curr. Protoc. Bioinformatics 47, 1–34 (2014).
Lorenzo, D. N. & Bennett, V. Cell-autonomous adiposity through increased cell surface GLUT4 due to ankyrin-B deficiency. Proc. Natl Acad. Sci. USA 114, 12743–12748 (2017).
Burridge, K., Kelly, T. & Mangeat, P. Nonerythrocyte spectrins: actin-membrane attachment proteins occurring in many cell types. J. Cell Biol. 95, 478–486 (1982).
Snouwaert, J. N. et al. A mutation in the Borcs7 subunit of the lysosome regulatory BORC complex results in motor deficits and dystrophic axonopathy in mice. Cell Rep. 24, 1254–1265 (2018).
García-Alvarez, B., Bobkov, A., Sonnenberg, A. & de Pereda, J. M. Structural and functional analysis of the actin binding domain of plectin suggests alternative mechanisms for binding to F-actin and integrin β4. Structure 11, 615–625 (2003).
Zhang, Y. I-TASSER server for protein 3D structure prediction. BMC Bioinformatics 9, 40 (2008).
von der Ecken, J., Heissler, S. M., Pathan-Chhatbar, S., Manstein, D. J. & Raunser, S. Cryo-EM structure of a human cytoplasmic actomyosin complex at near-atomic resolution. Nature 534, 724–728 (2016).
Källberg, M., Margaryan, G., Wang, S., Ma, J. & Xu, J. RaptorX server: a resource for template-based protein structure modeling. Methods Mol. Biol. 1137, 17–27 (2014).
Jurrus, E. et al. Improvements to the APBS biomolecular solvation software suite. Protein Sci. 27, 112–128 (2018).
Shindyalov, I. N. & Bourne, P. E. Protein structure alignment by incremental combinatorial extension (CE) of the optimal path. Protein Eng. 11, 739–747 (1998).
Acknowledgements
We thank all the families who participated in this study. We thank M. Rasband and K.-A. Nave for the gift of the βII-spectrin conditional null and the Nex-Cre mice, respectively. We thank N. V. Riddick for her assistance with the behavioral studies. We thank B. Koller and K. Mohlke for their insightful comments on this manuscript and J. Bear for helpful discussions. M.A.C., L.E.S.-R. and E.W.K. were supported by the Center for Individualized Medicine at the Mayo Clinic. D.N.L. was supported by the University of North Carolina at Chapel Hill (UNC-CH) School of Medicine as a Simmons Scholar, by the National Ataxia Foundation and by the US National Institutes of Health (NIH) grant no. R01NS110810. E.E.E. was supported by the NIH grant no. MH101221. Microscopy was performed at the UNC-CH Neuroscience Microscopy Core Facility, supported, in part, by funding from the NIH-NINDS Neuroscience Center Grant no. P30 NS045892 and the NIH-NICHD Intellectual and Developmental Disabilities Research Center Support Grant no. U54 HD079124, which also supported the behavioral studies. Research reported in this manuscript was supported by the NIH Common Fund, through the Office of Strategic Coordination/Office of the NIH Director under Award Number U01HG007672 (Duke University to V. Shashi). The content is solely the responsibility of the authors and does not necessarily represent the official views of the NIH. This research was made possible through access to the data and findings generated by the 100,000 Genomes Project. The 100,000 Genomes Project is managed by Genomics England Limited (a wholly owned company of the Department of Health and Social Care). The 100,000 Genomes Project is funded by the National Institute for Health Research and NHS England. The Wellcome Trust, Cancer Research UK and the Medical Research Council have also funded research infrastructure. The 100,000 Genomes Project uses data provided by patients and collected by the National Health Service as part of their care and support.
Author information
Authors and Affiliations
Consortia
Contributions
M.A.C. and D.N.L. conceived and planned the study with input from Q.K.-G.T. and R.C.S. M.A.C. managed the collection, analysis and interpretation of patient clinical data with Q.K.-G.T., R.C.S. and D.N.L. D.N.L. designed the cell biology, histology and biochemistry studies; performed these; and analyzed the data together with B.A.C., K.A.B., S.D., D.A., R.J.E., S. Afriyie., J.C.B. and L.F.R. A.A.B., L.J.M. and A.S.B. generated and characterized the iPSCs. S.T., M.T.Z., B.T. and D.N.L. performed the structural modeling. K.M.H. and S.S.M. performed the mouse behavioral studies. M.C.S. contributed reagents. M.A.C. and D.N.L. wrote the manuscript with contributions from B.A.C., R.C.S., S.S.M., M.T.Z. and B.T. E.W.K. and D.N.L. supervised the study. All other authors including Q.K.-G.T. and R.C.S. contributed clinical data. All authors approved the final manuscript.
Corresponding authors
Ethics declarations
Competing interests
E.T., R.E.P., Y.S., E.A.N. and A.B. are employees of GeneDx, Inc.
Additional information
Peer review Information Nature Genetics thanks Karun Singh and the other, anonymous, reviewer(s) for their contribution to the peer review of this work. Peer reviewer reports are available.
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
Extended Data Fig. 1 Expression of SPTBN1 variants alters protein expression, cellular distribution and morphology.
a, Western blot of total lysates from HEK 293 T/17 cells co-transfected with GFP-βIISp and mCherry plasmids and blotted with anti-GFP and anti-mCherry antibodies. Results are representative of three independent experiments. b, Western blot of Triton-X100 soluble and insoluble fractions from HEK 293 T/17 cell lysates transfected with GFP-βIISp plasmids and blotted with anti-GFP antibody. Images are representative of three independent experiments. c, Partition of indicated GFP-βIISp proteins expressed in HEK 293 T/17 cells between Triton-X100 soluble and insoluble fractions relative to total GFP-βIISp levels. Data in c were collected from n = 3 biological replicates in three independent experiments. Data represent mean ± SEM. One-way ANOVA with Dunnett’s post hoc analysis test for multiple comparisons, ***P = 0.001, ****P < 0.0001. d, Western blot of total lysates from primary mouse cortical neurons from βIISp-KO mice transduced with lentivirus expressing RFP-PP-βIISp proteins driven by the neuronal-specific synapsin I promoter and blotted with anti-RFP and anti-βIII-tubulin antibodies. Red arrowheads and boxes mark the presence of an additional 70-kDa GFP-positive fragment in HEK 293 T/17 (a) and mouse neuron (d) lysates expressing variants that result in GFP-positive aggregates. Blots are representative of three separate experiments. e, Western blot of total lysates from human iPSC lines reprogrammed from PBMCs carrying the indicated variants and blotted with anti-βII-spectrin and anti-α-tubulin antibodies. A red asterisk indicates the presence of a truncated 205-kDa βII-spectrin fragment in lysates from iPSCs reprogrammed from P27 (p.W1787*, c.5361 G > A). Blots are representative of four independent experiments. Western blot images were cropped from Source Data Extended Data Fig. 1. f, Analysis of sequencing reads from RNA-seq of blood RNA obtained from P27 (p.W1787*, c.5361 G > A) indicate allelic expression bias, suggesting some level of nonsense mediated decay of the SPTBN1 allele transcript harboring the nonsense variant, and increased abundance of the major c.5361 G SPTBN1 allele. g, Quantification of the percent of GFP-positive HEK 293 T/17 cells with GFP aggregates for each of the indicated variants. Data were collected from n = 20 cells/genotype pooled from three independent experiments and the following number of transfection replicates: WT (n = 10), T59I (n = 3), I59Q_160Δ (n = 5), C183* (n = 6), Y190_R216Δ (n = 3), G205D (n = 4), G205S (n = 6), L247H (n = 5), L250R (n = 7), D255E (n = 5), T268A (n = 4), T268N (n = 4), T268S (n = 6), V271M (n = 4), H275R (n = 6), F344L (n = 4), R411W (n = 3), E491Q (n = 4), A850G (n = 3), E892* (n = 3), R1003W (n = 9), A1086T (n = 4), E1110D (n = 4), G1398S (n = 3), W1787* (n = 3), E1886Q (n = 4). h,i, Quantification of cell length (h) and filopodia density normalized to cell length (i) of GFP-positive HEK 293 T/17 cells expressing the indicated variants. Data in h and i were collected from WT (n = 23), T59I (n = 13), I59Q_160Δ (n = 12), C183* (n = 12), Y190_R216Δ (n = 26), G205D (n = 11), G205S (n = 11), L247H (n = 22), L250R (n = 14), D255E (n = 18), T268A (n = 13), T268N (n = 15), T268S (n = 12), V271M (n = 10), H275R (n = 13), F344L (n = 12), R411W (n = 10), E491Q (n = 11), A850G (n = 11), E892* (n = 12), R1003W (n = 15), A1086T (n = 10), E1110D (n = 12), G1398S (n = 10), W1787* (n = 12), and E1886Q (n = 12) cells pooled from six independent experiments. All data represent mean ± SEM. One-way ANOVA with Dunnett’s post hoc analysis test for multiple comparisons. (g) ****P < 0.0001, ns P > 0.05. (h) *P = 0.0119 (T268S), *P = 0.0376 (H275R), *P = 0.0184 (R411W), *P = 0.0492 (A850G); **P = 0.0029 (T59I), **P = 0.0083 (V271M); ***P = 0.0009 (E1110D), ***P = 0.0005 (E1886Q); ****P < 0.0001. (i) *P = 0.0141 (G205D); **P = 0.0079 (C183*), **P = 0.0023 (Y190_R216Δ), **P = 0.0027 (G205S), **P = 0.0083 (R411W); ***P = 0.0006 (E491Q), ***P = 0.0002 (E1886Q); ****P < 0.0001. See statistics summary in Source Data Extended Data Fig. 5.
Extended Data Fig. 2 SPTBN1 variants alter interaction with critical cytoskeleton partners.
a, Immunofluorescence images, representative of three independent experiments, show HEK 293 T/17 cells transfected with mCherry-αIISp and with either WT or mutant GFP-βIISp plasmids. Cells were stained for actin (phalloidin) and DAPI. Scale bar, 10 μm. b, Immunofluorescence images of DIV8 mouse βIISp-WT (top) and βIISp-Het (bottom) cortical neurons transfected with indicated GFP-βIISp plasmids. Scale bar, 10 μm. GFP-positive aggregates are detected in neurons expressing these subsets of CH domain variants regardless of the level of endogenous βII-spectrin. Images are representative of n = 15 neurons per transfection derived from three independent experiments. c, Western blot from a binding assay to assess interaction between mCherry-αIISp and GFP-βII-spectrin proteins representative of n = 3 biological replicates from three independent experiments. Lysates from HEK 293 T/17 cells expressing mCherry-αII-spectrin were incubated with GFP-βII-spectrin proteins coupled to GFP beads. The presence of mCherry-αII-spectrin in eluates from GFP beads was evaluated by blotting with anti-GFP and anti-mCherry antibodies. d, Coomassie blue staining showing the presence of purified full-length βII-spectrin and F-actin in the supernatant (S) and pellet (P) fractions from an actin co-sedimentation assay. Blot is representative of three independent experiments each with n = 1 biological replicate. e, Co-IP assay in HEK 293 T/17 cells to assess interaction between 220-kDa ankyrin-B (AnkB)-2HA and GFP-βII-spectrin proteins. The presence of 220-kDa AnkB-3xHA and GFP-βII-spectrin proteins in initial lysates and eluates from beads coupled to rabbit IgG isotype control of a rabbit anti-GFP antibody was detected by blotting with anti-GFP and anti-HA antibodies. Blot is representative of four independent experiments, each with n = 1 biological replicate. Western blot images were cropped from Source Data Extended Data Fig. 2.
Extended Data Fig. 3 Modeling effects of SPTBN1 variants.
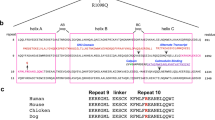
a-c, Potential coevolution of the closed conformation of the tandem calponin homology domains (CH1-CH2) of βII-spectrin (SPTBN1) (CH1 domain (teal), CH2 domain (red)), actinin-4 (ACTN4) (brown), and utrophin (UTRN) (purple)47. d,e, Top hits from docking simulations of βII-spectrin’s CH1 (d) and CH248 (e) onto F-actin (gray). Domains in dark blue correspond to cryo-EM structure of the CH1 domain of βIII-spectrin bound to F-actin49. f, Correct length of simulated interdomain linker (dark blue) in agreement with the orientation of the docked CH2 domain (red). g,h, Spatial distributions of the missense variants in βII-spectrin implicate disease mechanisms. g, Linear conformation of the entire 3D protein model is shown with the calponin homology (CH) domains (CH1 and CH2) in the N-terminus (red), the spectrin repeats (SR) (green) and the pleckstrin homology (PH) domain in the C-terminus (purple). h, The 17 SR domains are superimposed with a minimal cartoon representation to emphasize the consistency of the 3D architecture despite high sequence diversity. The positions of the amino acid residues representing the missense variants are marked by gold-colored spheres.
Extended Data Fig. 4 Effects of SPTBN1 variants on axonal growth.
a, Images of DIV8 βII-SpWT, βII-SpHet, βII-SpKO, and GFP-βIISp rescued βII-SpKO neurons transfected at DIV3 with mCherry. Staining with an antibody specific for AnkG was used to label the AIS (yellow arrowhead) and to identify axonal processes. Scale bar, 30 μm. Images are representative of three independent experiments.
Extended Data Fig. 5 Effects of SPTBN1 variants on dendrites.
a, Images of DIV18 βII-SpWT, βII-SpHet, βII-SpKO, and GFP-βIISp rescued βII-SpKO neurons stained with an anti-GFP antibody. Scale bar, 30 μm. b,c, Quantification of length of primary dendrites (b) and of total number of primary and secondary dendrites (c) of βII-SpWT, βII-SpHet, βII-SpKO, and rescued βII-SpKO DIV18 neurons (n = 6-16 neurons/genotype) compiled from three independent experiments. Data represent mean ± SEM. One-way ANOVA with Dunnett’s post hoc analysis test for multiple comparisons, *P < 0.05, **P < 0.01, ***P < 0.001, ****P < 0.0001. See statistics summary in Source Data Extended Data Fig. 5.
Extended Data Fig. 6 Effects of βII-spectrin deficiency on neuronal morphology and brain development.
a, Image, representative from three independent experiments show DIV8 βII-SpKO cortical neurons rescued with WT GFP-βIISp or with GFP-βIISp bearing variants within the distal portion of the CH2 domain. Neurons were stained for actin (phalloidin) and endogenous αII-spectrin. Yellow dotted lines demark the cell edge. Scale bar, 5 μm. b, Images of PND25 βII-SpNexWT and βII-SpNexKO brains stained for neurofilament to label axons and DAPI. Staining for βII-spectrin show specific loss of the protein in axons from callosal projection neurons from βII-SpNexKO mice. Scale bar, 50 μm. White dotted lines denote the position and boundaries of the corpus callosum (CC). Brains were collected from two separate litters and processed for staining and imaging as part of one independent experiment.
Extended Data Fig. 7 Developmental and behavioral phenotypes of βII-spectrin deficient mice.
a, Images of male PND25 wildtype (βII-SpNexWT) mice and mice lacking βII-spectrin only in cortical and hippocampal projection neurons (βII-SpNexKO) driven by Nex-Cre. b-e, Magnitude of acoustic startle responses (b,d) and percent of prepulse inhibition (c,e) in βII-SpWT mice and mice with partial (βII-SpHet) and complete (βII-SpKO) loss of βII-spectrin in neural progenitors driven by Nestin-Cre. Trials included no stimulus (NoS) trials and acoustic startle stimulus (AS; 120 dB) alone trials. Data in b and c represent mean ± SEM (n = 15 βII-SpWT and n = 5 βII-SpKO male mice). Data in d and e represent mean ± SEM (n = 12 male mice/genotype). Fisher’s PLSD tests following repeated measures ANOVA. b,*P < 0.05, **P < 0.01. c-e, P > 0.05. f, Latency to fall from an accelerating rotarod. Trials 4 and 5 were given 48 h after the first three trials. g,h, Latencies to find the hidden escape platform during acquisition (g) and reversal (h) learning phases of the Morris water maze test for βII-SpWT and βII-SpHet mice. Data represent mean ± SEM of four trials per day. Fisher’s PLSD tests following repeated measures ANOVA. f-h, P > 0.05. i,j, Mice were given a one-minute probe trial without the platform following the acquisition and reversal phases of the Morris water maze test. Target indicates the site where the platform had been located in each phase. Measures were taken of swim path crossings over the target location or corresponding areas in the other quadrants. Within-genotype repeated measures ANOVA, effect of quadrant (the repeated measure), **P = 0.0012, ****P < 0.0001. k,l, Preference for social novelty during a three-chamber choice task. Within-genotype repeated measures ANOVA, *P = 0.0145, **P = 0.0052. Data in f-l represent mean ± SEM (n = 12 male mice/genotype). m, Lack of significant genotype effects on anxiety-like behavior in the elevated plus maze, marble-burying assay, and open field; sensory ability in the buried food test for olfactory function and hot plate test for thermal sensitivity; and vision and swimming ability in the Morris water maze. Data represent mean ± SEM (n = 12 male mice/genotype). Within-genotype repeated measures ANOVA, P > 0.05.
Supplementary information
Supplementary Information
Supplementary Note
Supplementary Tables
Supplementary Tables 1–5
Source data
Source Data Fig. 2
Statistical Source Data
Source Data Fig. 3
Statistical Source Data
Source Data Fig. 5
Statistical Source Data
Source Data Fig. 6
Statistical Source Data
Source Data Fig. 7
Statistical Source Data
Source Data Extended Data Fig. 1
Statistical Source Data
Source Data Extended Data Fig. 1
Unprocessed Western Blots
Source Data Extended Data Fig. 2
Unprocessed Western Blots
Source Data Extended Data Fig. 5
Statistical Source Data
Rights and permissions
About this article
Cite this article
Cousin, M.A., Creighton, B.A., Breau, K.A. et al. Pathogenic SPTBN1 variants cause an autosomal dominant neurodevelopmental syndrome. Nat Genet 53, 1006–1021 (2021). https://doi.org/10.1038/s41588-021-00886-z
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41588-021-00886-z
This article is cited by
-
Consensus reporting guidelines to address gaps in descriptions of ultra-rare genetic conditions
npj Genomic Medicine (2024)
-
Spectrins: molecular organizers and targets of neurological disorders
Nature Reviews Neuroscience (2023)
-
Spectrin-beta 2 facilitates the selective accumulation of GABAA receptors at somatodendritic synapses
Communications Biology (2023)
-
Essential genes: a cross-species perspective
Mammalian Genome (2023)
-
Recent advances in novel mutation genes of Parkinson's disease
Journal of Neurology (2023)