Abstract
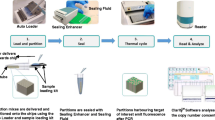
An integrated droplet digital polymerase chain reaction (ddPCR) gene chip that can quantify nucleic acid absolutely was constructed to avoid problems such as droplet fusion, fragmentation and sample contamination during pipetting. The integrated ddPCR gene chip was fabricated by photolithography and cyclic olefin copolymer (COC) injection mold. The 20 μL sample was distributed to about 60,000 water-in-oil droplets with a diameter of about 87 μm within 3 min by flow focus microstructure, and the volume of each droplet was about 0.27 nL. The generation of bubbles was reduced by reducing the width of the collection chamber and increasing the deceleration step. In addition, the efficiency of PCR was improved using COC materials. The sample of human epidermal growth factor receptor (EGFR) exon18 gene was quantitatively detected by this chip. Experimental results show that the detection signal has a good linear relationship with the DNA concentration from 101 to 105 copies/μL (R2 = 0.999) so that the integrated ddPCR gene chip can realize absolute quantification of nucleic acid. It is verified that the integration of droplet generation, PCR amplification and fluorescence detection are realized on one chip, reduce bubble generation and improve PCR efficiency. Compared with the current independent split chip, the integrated ddPCR gene chip ensures a high degree of anti-pollution performance, avoids the operation of droplet movement, reduces human interference, and standardizes the results. It has wide application prospects in nucleic acid quantification and trace detection.








Similar content being viewed by others
Data availability
The data that support the findings of this study are available from the corresponding author upon reasonable request.
References
Ahrberg CD, Manz A, Chung BG (2016) Polymerase chain reaction in microfluidic devices. Lab Chip 16(20):3866
Anna SL, Bontoux N, Stone HA (2003) Formation of dispersions using “flow focusing” in microchannels. Adv Mech 82(3):364–366
Basu AS (2017) Digital assays part I: partitioning statistics and digital PCR. Slas Technol 22(1):2472630317705680
Cao L, Cui X, Hu J, Li Z, Choi JR, Yang Q et al (2016) Advances in digital polymerase chain reaction (dPCR) and its emerging biomedical applications. Biosens Bioelectron 90:459–474
Chen H, Li J, Zhang H, Li M, Rosengarten G, Nordon RE (2011) Microwell perfusion array for high-throughput, long-term imaging of clonal growth. Biomicrofluidics 5(4):345
Chen H, Cornwell J, Zhang H, Lim T, Resurreccion R, Port T et al (2013) Cardiac-like flow generator for long-term imaging of endothelial cell responses to circulatory pulsatile flow at microscale. Lab Chip 13(15):2999–3007
Chen J, Luo Z, Li L et al (2018) Capillary-based integrated digital PCR in picoliter droplets. Lab Chip 18(3):412–421
Ciardiello F, Tortora G (2008) Drug therapy: EGFR antagonists in cancer treatment. N Engl J Med 358(11):1160–1174
Dreyfus R, Tabeling P, Willaime H (2003) Ordered and Disordered patterns in two-phase flows in microchannels. Phys Rev Lett 90(14):144505
Fan Y, Dong D, Li Q et al (2018) Fluorescent analysis of bioactive molecules in single cells based on microfluidic chips. Lab Chip 18(8):1151–1173
Garstecki P, Gitlin I, Diluzio W, Whitesides GM, Kumacheva E, Stone HA (2004) Formation of monodisperse bubbles in a microfluidic flow-focusing device. Appl Phys Lett 85(13):2649–2651
Hatch AC, Fisher JS, Tovar AR, Hsieh AT, Lin R, Pentoney SL et al (2011) 1-Million droplet array with wide-field fluorescence imaging for digital PCR. Lab Chip 11(22):3838–3845
Jin Q, Zhao J, Jing F et al (2015) A microfluidic droplet digital PCR for simultaneous detection of pathogenic Escherichia coli O157 and Listeria monocytogenes. Biosens Bioelectron 74:770–777
Jingmin Li, Chao L, Shuai W et al (2017) Using serrated edges to control fluid front motion in microfluidic devices. Microsyst Technol 23(10):4733–4740
Kanagal-Shamanna R (2016) Digital PCR: principles and applications. In: Clinical applications of PCR. Springer, Berlin, Germany
Kukhtin AC, Sebastian T, Golova J, Perov A, Knickerbocker C, Linger Y et al (2019) Lab-on-a-Film disposable for genotyping multidrug-resistant Mycobacterium tuberculosis from sputum extracts Electronic supplementary information (ESI) available. See. https://doi.org/10.1039/c8lc01404c.LabonAChip
Madic J, Zocevic A, Senlis V, Fradet E, Droniou ME (2016) Three-color crystal digital PCR. Biomol Detect Quantifi 10:34–46
Nunes PS, Ohlsson PD, Ordeig O, Kutter JP (2010) Cyclic olefin polymers: emerging materials for lab-on-a-chip applications. Microfluid Nanofluid 9(2):145–161
Pekin D, Skhiri Y, Baret JC, Corre DL, Mazutis L, Salem CB et al (2011) Quantitative and sensitive detection of rare mutations using droplet-based microfluidics. Lab Chip 11(13):2156–2166
Ping W, Fengxiang J, Gang L et al (2015) Absolute quantification of lung cancer related microRNA by droplet digital PCR. Biosens Bioelectron 74:836–842
Schuler F, Trotter M, Geltman M, Schwemmer F, Wadle S, Domínguez-Garrido E et al (2015) Digital droplet PCR on disk. Lab Chip 16(1):208–216
Schuler F, Siber C, Hin S, Wadle S, Paust N, Zengerle R, Von Stetten F (2016) Digital droplet LAMP as a microfluidic app on standard laboratory devices. Anal Methods 8(13):2750–2755
Sharma SV, Bell DW, Settleman J, Haber DA (2007) Epidermal growth factor receptor mutations in lung cancer. Nat Rev Cancer 7(3):169–181
Sreejith KR, Ooi CH, Jin J, Dao DV, Nguyen NT (2018) Digital polymerase chain reaction technology—recent advances and future perspectives. Lab Chip 18(24):3717–3732
Stavis SM (2012) A glowing future for lab on a chip testing standards. Lab Chip 12(17):3008–3011
Suo T, Liu X, Feng J, Guo M, Chen Y (2020) ddPCR: a more accurate tool for SARS-CoV-2 detection in low viral load specimens. Emerg Microbes Infect 9(75296):1–30
Takeuchi S (2010) An axisymmetric flow-focusing microfluidic device. Adv Mater 17(8):1067–1072
Yoon KB, Choi JS, Lee DH, Hur Y, Noh SK (2006) Synthesis and optical properties of cycloolefin copolymers for plastic substrates. Mol Cryst Liq Cryst 459(1):27/[307]-334/[314]
Zhang J, Jia C, Mao H et al (2015) Absolute quantification of lung cancer related microRNA by droplet digital PCR. Biosens Bioelectron 74:836–842
Zhang H, Xiao L, Li Q, Qi X, Zhou A (2018) Microfluidic chip for non-invasive analysis of tumor cells interaction with anti-cancer drug doxorubicin by AFM and Raman spectroscopy. Biomicrofluidics 12(2):024119
Zhang H, Zhang W, Xiao L, Liu Y, Gilbertson T, Zhou A (2019) Use of Surface-Enhanced Raman Scattering (SERS) probes to detect fatty acid receptor activity in a microfluidic device. Sensors 19(7):1663
Zhang H, Guzman AR, Wippold JA, Li Y, Dai J, Huang C, Han A (2020) An ultra high-efficiency droplet microfluidics platform using automatically synchronized droplet pairing and merging. Lab Chip 20(21):3948–3959
Zida L, Luoquan L, Meixiang L, Liqun H, Ping W (2019) Multiple splitting of droplets using multi-furcating microfluidic channels. Biomicrofluidics 13(2):024112
Acknowledgements
This work described in this paper was supported by the Jilin Scientific and Technological Development Program (CN) (Grant Number 20191102003YY).
Funding
This study was funded by the Jilin Scientific and Technological Development Program (CN) (Grant Number 20191102003YY).
Author information
Authors and Affiliations
Contributions
GJ contributed to the conception of the study, YY and PG performed the experiment, XM contributed data analysis and wrote the manuscript.
Corresponding author
Ethics declarations
Conflict of interest
The authors declare no financial or commercial conflict of interest.
Ethical approval
The article does not contain any studies with human participants or animals performed by any authors.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
About this article
Cite this article
Meng, X., Yu, Y., Gong, P. et al. An integrated droplet digital PCR gene chip for absolute quantification of nucleic acid. Microfluid Nanofluid 25, 62 (2021). https://doi.org/10.1007/s10404-021-02465-4
Received:
Accepted:
Published:
DOI: https://doi.org/10.1007/s10404-021-02465-4