Abstract
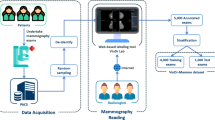
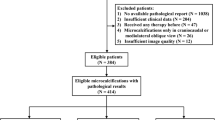
Recently, computer-aided diagnosis (CAD) systems powered by deep learning (DL) algorithms have shown excellent performance in the evaluation of digital mammography for breast cancer diagnosis. However, such systems typically require pixel-level annotations by expert radiologists which is prohibitively time-consuming and expensive. Medical institutes would wonder if a high-performance breast cancer CAD system can be trained by exploring their own huge amount of historical imaging data and corresponding diagnosis reports, without additional annotations workload of their radiologists. In this study, we show that a DL classification model trained on historical mammograms with only image-level pathology labels (which can be automatically extracted from medical reports) can achieve surprisingly good diagnostic performance on newly incoming exams compared with experienced radiologists. A DL model called DenseNet was trained and cross-validated with 5979 historical exams acquired before September 2017 with biopsy-verified pathology and tested with 1194 newly obtained cases after that. For both cross-validation and test sets, the ROCs generated by DL predictions were above the ROCs generated by ratings from radiologists. For the suspicious cases which radiologists suggest biopsy (BI-RADS category 4 and 5), the DL model can reject 60% of false biopsies on benign breasts while keeping 95% sensitivity. For the mammograms based on which radiologists were not able to make a diagnosis (BI-RADS 0), the DL model still achieved an AUC score of 79%. Moreover, the model is able to localize lesions on mammograms although such information was not provided in the training phase. Finally, the impact of input image resolution and different DL model architectures on the diagnostic accuracy were also presented and analyzed.






Similar content being viewed by others
References
Jemal, A., Siegel, R., Ward, E., Hao, Y., Xu, J., Murray, T., Thun, M.J.: Cancer statistics, 2008. CA Cancer J. Clin. 58(2), 71–96 (2008)
DeSantis, C., et al.: Breast cancer statistics, 2013. CA Cancer J. Clin. 64(1), 52–62 (2014)
Lehman, C.D., Arao, R.F., Sprague, B.L., Lee, J.M., Buist, D.S., Kerlikowske, K., et al.: National performance benchmarks for modern screening digital mammography: update from the Breast Cancer Surveillance Consortium. Radiology 283(1), 49–58 (2016)
Sickles, E.A., Wolverton, D.E., Dee, K.E.: Performance parameters for screening and diagnostic mammography: specialist and general radiologists. Radiology 224(3), 861–869 (2002)
Wu, N., et al.: Deep neural networks improve radiologists’ performance in breast cancer screening. IEEE Trans. Med. Imaging 39(4), 1184–1194 (2019)
Ribli, D., et al.: Detecting and classifying lesions in mammograms with deep learning. Sci. Rep. 8(1), 1–7 (2018)
Shen, L., et al.: Deep learning to improve breast cancer detection on screening mammography. Sci. Rep. 9(1), 1–12 (2019)
Rodríguez-Ruiz, A., et al.: Detection of breast cancer with mammography: effect of an artificial intelligence support system. Radiology 290(2), 305–314 (2019)
Gardezi, S.J.S., et al.: Breast cancer detection and diagnosis using mammographic data: Systematic review. J. Med. Internet Res. 21(7), e14464 (2019)
Wang, X., et al.: Inconsistent performance of deep learning models on mammogram classification. J. Am. Coll. Radiol. 17(6), 796–803 (2020)
Krizhevsky, A., Sutskever, I., Hinton, G.E.: Imagenet classification with deep convolutional neural networks. Adv. Neural. Inf. Process. Syst. 25(2), 1097–1105 (2012)
Simonyan, K., Zisserman, A.: Very deep convolutional networks for large-scale image recognition. Ppreprint at arXiv:1409.1556 (2014)
He, K., Zhang, X., Ren, S., & Sun, J.: Deep residual learning for image recognition. In: Proceedings of the IEEE conference on computer vision and pattern recognition, pp. 770–778 (2016)
Huang, G., Liu, Z., Van Der Maaten, L., Weinberger, K. Q.: Densely connected convolutional networks. In: Proceedings of the IEEE conference on computer vision and pattern recognition, pp. 4700–4708 (2017)
Kingma, D.P., Ba, J.A.: A method for stochastic optimization. arXiv preprint arXiv:1412.6980 (2014)
Deng, J., Dong, W., Socher, R., Li, L.J., Li, K., Fei-Fei, L.: Imagenet: a large-scale hierarchical image database. In: 2009 IEEE conference on computer vision and pattern recognition, pp. 248–255 (2009)
Paszke, A., Gross, V., Chintala, S., Chanan, G., Yang, E., DeVito, Z., Lin, Z., Desmaison, A., Antiga, L., Lerer, A.: NIPS 2017 Workshop on Autodiff (2017 )
Wu, Y., He, K.: Group normalization. In: Proceedings of the European Conference on Computer Vision (ECCV) (2018)
Becker, A.S., Marcon, M., Ghafoor, S., Wurnig, M.C., Frauenfelder, T., Boss, A.: Deep learning in mammography: diagnostic accuracy of a multipurpose image analysis software in the detection of breast cancer. Invest. Radiol. 52(7), 434–440 (2017)
Rodriguez-Ruiz, A., Lång, K., Gubern-Merida, A., Broeders, M., Gennaro, G., Clauser, P., et al.: Stand-alone artificial intelligence for breast cancer detection in mammography: comparison with 101 radiologists. J Natl. Cancer Inst. 111(9), 916–922 (2019)
Jiang, Y., Metz, C.E.: BI-RADS data should not be used to estimate ROC curves. Radiology 256(1), 29–31 (2010)
Zhou, B., Khosla, A., Lapedriza, A., Oliva, A., Torralba, A.: Learning deep features for discriminative localization. In: Proceedings of the IEEE conference on computer vision and pattern recognition, pp. 2921–2929 (2016)
Rajpurkar, P., Irvin, J., Zhu, K., Yang, B., Mehta, H., Duan, T., et al.: Chexnet: radiologist-level pneumonia detection on chest x-rays with deep learning. arXiv preprint arXiv:1711.05225 (2017)
Lee, R.S., et al.: A curated mammography data set for use in computer-aided detection and diagnosis research. Sci. Data 4(1), 1–9 (2017)
Moreira, I.C., et al.: Inbreast: toward a full-field digital mammographic database. Acad. Radiol. 19(2), 236–248 (2012)
Kooi, T., Litjens, G., Van Ginneken, B., Gubern-Mérida, A., Sánchez, C.I., Mann, R., et al.: Large scale deep learning for computer aided detection of mammographic lesions. Med. Image Anal. 35, 303–312 (2017)
Rodriguez-Ruiz, A., Lång, K., Gubern-Merida, A., Broeders, M., Gennaro, G., Clauser, P., et al.: Stand-alone artificial intelligence for breast cancer detection in mammography: comparison with 101 radiologists. J. Natl. Cancer Inst. 111(9), 916–922 (2019)
Aboutalib, S.S., Mohamed, A.A., Berg, W.A., Zuley, M.L., Sumkin, J.H., Wu, S.: Deep learning to distinguish recalled but benign mammography images in breast cancer screening. Clin. Cancer Res. 24(23), 5902–5909 (2018)
Zhu, W., et al.: Deep multi-instance networks with sparse label assignment for whole mammogram classification. In: International conference on medical image computing and computer-assisted intervention. Springer, Cham (2017)
Liang, Y., et al.: Urbanfm: inferring fine-grained urban flows. In: Proceedings of the 25th ACM SIGKDD international conference on knowledge discovery & data mining (2019)
Ouyang, K., et al.: Fine-grained urban flow inference. IEEE Trans. Knowl. Data Eng. (2020).
Yan, C., et al.: Deep multi-view enhancement hashing for image retrieval. IEEE Trans. Pattern Anal. Mach. Intell. (2020).
Yan, C., et al.: 3D room layout estimation from a single RGB image. IEEE Trans. Multimedia 22(11), 3014–3024 (2020)
Funding
This work is supported by National Natural Science Foundation of China (Grand No. 61702453), Natural Science Foundation of Zhejiang Province (Grand No. Q17F030003).
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors declare no conflicts of interest.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
About this article
Cite this article
Qu, J., Zhao, X., Chen, P. et al. Deep learning on digital mammography for expert-level diagnosis accuracy in breast cancer detection. Multimedia Systems 28, 1263–1274 (2022). https://doi.org/10.1007/s00530-021-00823-4
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00530-021-00823-4