Abstract
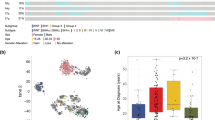
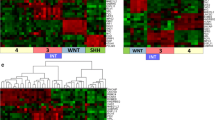
Medulloblastoma is a common pediatric malignant brain tumor. There were four consensus molecular subgroups (WNT, SHH, Group3 and Group4). Group 3 and Group 4 tumors exhibited a great degree of transcriptional overlap, and were neither derived from exact pathway aberration. We investigated transcriptional and chromatin accessibility of medulloblastoma by multi-omics single-cell analysis. Our work identified inter- and intra-tumoral heterogeneity within the Group 3, Group 4 and Group 3/4 intermediate subgroups. Unsupervised cluster of each tumor identified 9 cell clusters with transcriptional profiles and 6 cell clusters with chromatin accessibility profiles. OTX2 had the highest activity and expression level across the clusters in a special cluster based on open chromatin single-cell profilings. We identified multiple genes as a significant targeted gene within the OTX2 target genes, which made sense in prognosis. We analyzed the copy-number-variations which presented with expected subgroup distribution from transcriptional and chromatin accessibility profiles. Collectively, these data provide novel insights into Group 3 and Group 4 medulloblastoma and provide a potential therapeutic target.




Similar content being viewed by others
References
Archer TC, Mahoney EL, Pomeroy SL (2017) Medulloblastoma: molecular classification-based personal therapeutics. Neurotherapeutics 14(2):265–273
Wang J et al (2018) Medulloblastoma: from molecular subgroups to molecular targeted therapies. Annu Rev Neurosci 41:207–232
Ramaswamy V et al (2016) Medulloblastoma subgroup-specific outcomes in irradiated children: who are the true high-risk patients? Neuro Oncol 18(2):291–297
Taylor MD et al (2012) Molecular subgroups of medulloblastoma: the current consensus. Acta Neuropathol 123(4):465–472
Louis DN et al (2016) The 2016 World Health Organization classification of tumors of the central nervous system: a summary. Acta Neuropathol 131(6):803–820
Kool M et al (2008) Integrated genomics identifies five medulloblastoma subtypes with distinct genetic profiles, pathway signatures and clinicopathological features. PLoS ONE 3(8):e3088
Hovestadt V et al (2019) Resolving medulloblastoma cellular architecture by single-cell genomics. Nature 572(7767):74–79
Tanay A, Regev A (2017) Scaling single-cell genomics from phenomenology to mechanism. Nature 541(7637):331–338
Chen S, Lake BB, Zhang K (2019) High-throughput sequencing of the transcriptome and chromatin accessibility in the same cell. Nat Biotechnol 37(12):1452–1457
Rai V et al (2020) Single-cell ATAC-Seq in human pancreatic islets and deep learning upscaling of rare cells reveals cell-specific type 2 diabetes regulatory signatures. Mol Metab 32:109–121
Zhu C et al (2019) An ultra high-throughput method for single-cell joint analysis of open chromatin and transcriptome. Nat Struct Mol Biol 26(11):1063–1070
Uhlén M et al (2015) Proteomics. Tissue-based map of the human proteome. Science 347(6220):1260419
Zhang L et al (2019) Single-cell transcriptomics in medulloblastoma reveals tumor-initiating progenitors and oncogenic cascades during tumorigenesis and relapse. Cancer Cell 36(3):302-318.e7
Zheng R et al (2019) Cistrome data browser: expanded datasets and new tools for gene regulatory analysis. Nucleic Acids Res 47(D1):D729-d735
Mei S et al (2017) Cistrome Data Browser: a data portal for ChIP-Seq and chromatin accessibility data in human and mouse. Nucleic Acids Res 45(D1):D658-d662
Cavalli FMG et al (2017) Intertumoral Heterogeneity within Medulloblastoma Subgroups. Cancer Cell 31(6):737-754.e6
Liu L et al (2019) An integrated chromatin accessibility and transcriptome landscape of human pre-implantation embryos. Nat Commun 10(1):364
Takahashi K, Yamanaka S (2006) Induction of pluripotent stem cells from mouse embryonic and adult fibroblast cultures by defined factors. Cell 126(4):663–676
Buenrostro JD et al (2013) Transposition of native chromatin for fast and sensitive epigenomic profiling of open chromatin, DNA-binding proteins and nucleosome position. Nat Methods 10(12):1213–1218
Lu Y et al (2017) OTX2 expression contributes to proliferation and progression in Myc-amplified medulloblastoma. Am J Cancer Res 7(3):647–656
Davie K et al (2015) Discovery of transcription factors and regulatory regions driving in vivo tumor development by ATAC-seq and FAIRE-seq open chromatin profiling. PLoS Genet 11(2):e1004994
Morrissy AS et al (2017) Spatial heterogeneity in medulloblastoma. Nat Genet 49(5):780–788
Vladoiu MC et al (2019) Childhood cerebellar tumours mirror conserved fetal transcriptional programs. Nature 572(7767):67–73
Jones DT et al (2012) Dissecting the genomic complexity underlying medulloblastoma. Nature 488(7409):100–105
Schwalbe EC et al (2017) Novel molecular subgroups for clinical classification and outcome prediction in childhood medulloblastoma: a cohort study. Lancet Oncol 18(7):958–971
Bunt J et al (2012) OTX2 directly activates cell cycle genes and inhibits differentiation in medulloblastoma cells. Int J Cancer 131(2):E21-32
Boulay G et al (2017) OTX2 activity at distal regulatory elements shapes the chromatin landscape of group 3 medulloblastoma. Cancer Discov 7(3):288–301
Figueira Muoio VM et al (2019) OTX1 and OTX2 genes in medulloblastoma. World Neurosurg 127:e58–e64
Di C et al (2005) Identification of OTX2 as a medulloblastoma oncogene whose product can be targeted by all-trans retinoic acid. Cancer Res 65(3):919–924
Gumireddy K et al (2003) All-trans-retinoic acid-induced apoptosis in human medulloblastoma: activation of caspase-3/poly(ADP-ribose) polymerase 1 pathway. Clin Cancer Res 9(11):4052–4059
Hallahan AR et al (2003) BMP-2 mediates retinoid-induced apoptosis in medulloblastoma cells through a paracrine effect. Nat Med 9(8):1033–1038
Bai R et al (2010) Evaluation of retinoic acid therapy for OTX2-positive medulloblastomas. Neuro Oncol 12(7):655–663
Corbo JC et al (2010) CRX ChIP-seq reveals the cis-regulatory architecture of mouse photoreceptors. Genome Res 20(11):1512–1525
Acknowledgements
This research was funded by the National Natural Science Foundation of China (No. 81870834), Intergovernmental cooperation on international scientific and technological innovation (2017YFE0121200) and Special project of pediatrics of collaborative development center of Beijing hospital administration (No. XTYB201817).
Author information
Authors and Affiliations
Contributions
All authors contributed to the study conception and design. Material preparation, data collection and analysis were performed by ZL, YW, YS, and LT. The first draft of the manuscript was written by ZL and all authors commented on previous versions of the manuscript. All authors read and approved the final manuscript.
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest. All authors consent for publication.
Ethics approval
This research with approved by the research ethics committee of the Beijing Tiantan Hospital, Capital Medical University, Beijing and had been performed in accordance with the ethical standards laid down in the 1964 Declaration of Helsinki and its later amendments. All persons gave their informed consent prior to their inclusion in the study. Details that might disclose the identity of the subjects under study should be omitted.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Li, Z., Wei, Y., Shao, Y. et al. Multi-omics analysis of intertumoral heterogeneity within medulloblastoma uncharted-pathway subtypes. Brain Tumor Pathol 38, 234–242 (2021). https://doi.org/10.1007/s10014-021-00400-7
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10014-021-00400-7