Abstract
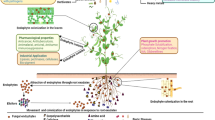
The diversity of actinobacteria associated with marine ascidian Phallusia nigra from Andaman Islands was investigated. A total of 10 actinobacteria were isolated and based on the biochemical and molecular characterization, the isolates were assigned to 7 different actinobacterial genera. Eight putatively novel species belonging to genera Rhodococcus, Kineococcus, Kocuria, Janibacter, Salinispora and Arthrobacter were identified based on 16S rDNA sequence similarity with the NCBI database. The organic extracts of ten isolates displayed considerable bioactivity against test pathogens, which were Gram-positive and Gram-negative in nature. PCR-based screening for type I and type II polyketide synthases (PKS-I, PKS-II) and nonribosomal peptide synthetases (NRPS) revealed that, 10 actinobacterial isolates encoded at least one type of polyketide synthases biosynthesis gene. Majority of the isolates found to produce industrially important enzymes; amylase, protease, gelatinase, lipase, DNase, cellulase, urease, phosphatase and l-asparaginase. The present study emphasized that, ascidians are a prolific resource for novel bioactive actinobacteria with potential for novel drug discovery. This result expands the scope to functionally characterize the novel ascidian associated marine actinobacteria and their metabolites could be a source for the novel molecules of commercial interest.




Similar content being viewed by others
References
Abdelmohsen UR, Bayer K, Hentschel U (2014) Diversity, abundance and natural products of marine sponge-associated actinomycetes. Nat Prod Rep 31:381–399. https://doi.org/10.1039/c3np70111e
Andreo MA, Jimenez PC, Siebra JB, Costa-Lotufo LV, Vessecchi R, Niehues M, Lopes JL, Lopes NP (2012) Systematic UPLC-ESI-MS/MS study on the occurrence of staurosporine and derivatives in associated marine microorganisms from Eudistoma vannamei. J Braz Chem Soc 23:335–343. https://doi.org/10.1590/S0103-50532012000200021
Asolkar RN, Kirkland TN, Jensen PR, Fenical W (2010) Arenimycin, an antibiotic effective against rifampin and methicillin-resistant Staphylococcus aureus from the marine actinomycete Salinispora arenicola. J Antibiot 63:37–39. https://doi.org/10.1038/ja.2009.114
Ayuso A, Clark D, Gonzalez I, Salazar O, Anderson A, Genilloud O (2005) A novel actinomycete strain de-replication approach based on the diversity of polyketide synthase and nonribosomal peptide synthetase biosynthetic pathways. Appl Microbiol Biotechnol 67:795–806. https://doi.org/10.1007/s00253-004-1828-7
Ayuso-Sacido A, Genilloud O (2005) New PCR primers for the screening of NRPS and PKS-I systems in actinomycetes: detection and distribution of these biosynthetic gene sequences in major taxonomic groups. Microb Ecol 49:10–24. https://doi.org/10.1007/s00248-004-0249-6
Baltz RH (2007) Antimicrobials from Actinomycetes: back to the future. Microbe 2:125–131. https://doi.org/10.4236/ns.2013.54A005
Bernfield P (1955) Amylases, α and β. Methods in enzymology, 1st edn. Academic Press, New York, pp 149–158
Bian GK, Feng ZZ, Qin S, Xing K, Wang Z, Cao CL, Liu CH, Dai CC, Jiang JH (2012) Kineococcus endophytica sp. nov., a novel endophytic actinomycete isolated from a coastal halophyte in Jiangsu, China. Antonie Van Leeuwenhoek 102:621–628. https://doi.org/10.1007/s10482-012-9757-4
Biehle JR, Cavalieri SJ, Felland T, Zimmer BL (1996) Novel method for rapid identification of Nocardia species by detection of performed enzymes. J Clin Microbiol 34:103–107. https://doi.org/10.1128/jcm.34.1.103-107.1996
Blunt JW, Copp BR, Keyzers RA, Munro MHG, Prinsep MR (2013) Marine natural products. Nat Prod Rep 30:237–323. https://doi.org/10.1039/D0NP00089B
Bright M, Bulgheresi S (2010) A complex journey: transmission of microbial symbionts. Nat Rev Microbiol 8:218–230. https://doi.org/10.1038/nrmicro2262
Carillo P, Mardarz C, Pitta-Alvarez S (1996) Isolation and selection of biosurfactant producing bacteria. World J Microbiol Biotechnol 12:82–84. https://doi.org/10.1007/BF00327807
Chen YG, Tang SK, Zhang YQ, Li ZY, Yi LB, Wang YX, Li WJ, Cui XL (2009) Arthrobacter halodurans sp. nov., a new halotolerant bacterium isolated from sea water. Antonie Van Leeuwenhoek 96:63–70. https://doi.org/10.1007/s10482-009-9336-5
Davidson L, Brear DR, Wingard P, Hawkins J, Kitto GB (1977) Purification and properties of an L-glutaminase–L-asparaginase from Pseudomonas acidovorans. J Bacteriol 129:1379–1386. https://doi.org/10.1128/jb.129.3.1379-1386.1977
Douglas A (2010) The symbiotic habit, 1st edn. Princeton University Press, Princeton, NJ
Duangmal K, Supattra M, Ratchanee M, Thanakorn Y, Nantana S, Vichien K, Atsuko M, Yoko T (2016) Kineococcus mangrovi sp. nov., isolated from mangrove sediment. Int J Syst Evol Microbiol 66:1230–1235. https://doi.org/10.1099/ijsem.0.000860
Ellaiah P, Kalyan D, Rao VS, Rao BV (1996) Isolation and characterization of bioactive actinomycetes from marine sediments. Hindustan Antibiot Bull 38:48–52. https://doi.org/10.1016/j.jgeb.2017.02.004
Erwin PM, Thacker RW (2008) Phototrophic nutrition and symbiont diversity of two Caribbean sponge–cyanobacteria symbioses. Mar Ecol Prog Ser 362:139–147. https://doi.org/10.3354/meps07464
Fenical W, Jensen PR (2006) Developing a new resource for drug discovery: marine actinomycete bacteria. Nat Chem Biol 2:666–673. https://doi.org/10.1038/nchembio841
Fenical W, Jensen PR, Palladino MA, Lam KS, Lloyd GK, Potts BC (2009) Discovery and development of the anticancer agent salinosporamide A (NPI-0052). Bioorg Med Chem 17:2175–2180. https://doi.org/10.1016/j.bmc.2008.10.075
Gandhimathi R, Seghal Kiran G, Hema TA, Selvin J, Rajeetha R, Shanmughapriya S (2009) Production and characterization of lipopeptide biosurfactant by a sponge-associated marine actinomycetes Nocardiopsis alba MSA10. Bioprocess Biosyst Eng 32:825–835. https://doi.org/10.1007/s00449-009-0309-x
Gulati R, Saxena RK, Gupta R (1997) A rapid plate assay for screening L-asparaginase producing micro-organisms. Lett Appl Microbiol 24:23–26. https://doi.org/10.1046/j.1472-765x.1997.00331.x
Hodges TW, Slattery M, Olson JB (2012) Unique actinomycetes from marine caves and coral reef sediments provide novel PKS and NRPS biosynthetic gene clusters. Mar Biotechnol 14:270–280. https://doi.org/10.1007/s10126-011-9410-7
Holt JG, Krieg NR, Sneath PHA, Staley JT, Williams ST (1994) Bergey’s manual of determinative bacteriology, 9th edn. Williams and Wilkins, Baltimore, pp 605–701
Ilic SB, Kontantinovic SS, Todorovic ZB (2005) UV/VIS analysis and antimicrobial activity of Streptomyces isolates. Facta Univ Med Biol 12:44–46
Imada A, Igarasi S, Nakahama K, Isono M (1973) Asparaginase and Glutaminase activities of microorganisms. J Gen Microbiol 76:85–99. https://doi.org/10.1099/00221287-76-1-85
Jang HD, Chen KS (2003) Production and purification of thermostable cellulases from Streptomyces transformant T3–1. World J Microbiol Biotechnol 19:263–268. https://doi.org/10.1023/A:1023641806194
Jensen PR, Fenical W (1994) Strategies for the discovery of secondary metabolites from marine bacteria: ecological perspectives. Annu Rev Microbiol 48:559–584. https://doi.org/10.1146/annurev.mi.48.100194.003015
Joulli MM, Leonard MS, Portonovo P, Liang B, Ding X, La Clair JJ (2003) Chemical defense in ascidians of the didemnidae family. J Bioconj Chem 14:30–37. https://doi.org/10.1021/bc025576n
Kageyama A, Takahashi Y, Yasumoto-Hirose M, Kasai H, Shizuri Y, Omura S (2007) Janibacter corallicola sp. nov., isolated from coral in Palau. J Gen Appl Microbiol 53:185–189. https://doi.org/10.2323/jgam.53.185
Kuster E, Williams S (1964) Selection of media for the isolation of Streptomyces. Nature 202:928–929. https://doi.org/10.1038/202928a0
Kwan JC, Tianero MDB, Donia MS, Wyche TP, Bugni TS, Schmidt EW (2014) Host control of symbiont natural product chemistry in cryptic populations of the tunicate Lissoclinum patella. PLoS ONE 9:e95850. https://doi.org/10.1371/journal.pone.0095850
Leal MC, Sheridan C, Osinga R, Dionisio G, Rocha RJM, Silva B, Rosa R, Calado R (2014) Marine microorganism-invertebrate assemblages: perspectives to solve the “supply problem” in the initial steps of drug discovery. Mar Drugs 12:3929–3952. https://doi.org/10.3390/md12073929
Lei C, Jin-Shuang H, Jia-Lei X, Chang-Lun S, Guang-Yu W (2018) Biological and chemical diversity of ascidian-associated microorganisms. Mar Drugs 16:362. https://doi.org/10.3390/md16100362
Lemos ML, Toranzo AE, Barja JL (1985) Antibiotic activity of epiphytic bacteria isolated from intertidal seaweeds. Microb Ecol 11:149–163. https://doi.org/10.1007/BF02010487
Leon J, Liza L, Soto I, Cuadra D, Patino L, Zerpa R (2007) Bioactives actinomycetes of marine sediment from the central coast of Peru. Revi Peru Boil 14:259–270
Lowry OH, Rosenbrough NJ, Farr AL, Randall RJ (1951) Protein estimation with the folin-phenol reagent. J Biol Chem 193:265–275. https://doi.org/10.1016/S0021-9258(19)52451-6
Meena B, Anburajan L, Vinithkumar NV, Kirubagaran R (2013) Novel marine actinobacteria from emerald Andaman & Nicobar Islands: a prospective source for industrial and pharmaceutical byproducts. BMC Microbiol 13:145. https://doi.org/10.1186/1471-2180-13-145
Meyer CA (2017) The Global Marine Pharmaceuticals Pipeline. http://marinepharmacology.midwestern.edu/clinPipeline.htm. Accessed 4 Apr 2017
Miller GL (1959) Use of dinitrosalicylic acid reagent for determination of reducing sugars. Anal Chem 31:426–428. https://doi.org/10.1021/ac60147a030
Mincer TJ, Fenical W, Jensen PR (2005) Culture-dependent and culture-independent diversity within the obligate marine actinomycete genus Salinispora. Appl Environ Microbiol 71:7019–7028. https://doi.org/10.1128/AEM.71.11.7019-7028.2005
Morikawa M, Daido H, Takao T, Marato S, Shimonishi Y, Imanaka T (1993) A new lipopeptide biosurfactant produced by Arthrobacter sp. strain MIS 38. J Bacteriol 175:6459–6466. https://doi.org/10.1128/jb.175.20.6459-6466.1993
Narayana KJ, Kumar KG, Vijayalakshmi M (2008) L-Asparaginase production by Streptomyces albidoflavus. Indian J Microbiol 48:331–336. https://doi.org/10.1007/s12088-008-0018-1
Paraszkiewicz K, Kanwal A, Dlugonski J (1992) Emulsifier production by steroid transforming filamentous fungus Curvularia lunata. Growth and product characterization. J Biotechnol 92:287–294. https://doi.org/10.1016/s0168-1656(01)00376-5
Park EJ, Kim MS, Roh SW, Jung MJ, Bae JW (2010) Kocuria atrinae sp. nov., isolated from traditional Korean fermented seafood. Int J Syst Evol Microbiol 60:914–918. https://doi.org/10.1099/ijs.0.014506-0
Peterson RE, Ciegler A (1969) L-Asparaginase production by Erwinia aroideae. J Appl Microbiol 18:64–67. https://doi.org/10.1128/am.18.1.64-67.1969
Prauser H (1976) Nocardioides, a new genus of the order actinomycetales. Int J Syst Bacteriol 26:58–65. https://doi.org/10.1099/00207713-26-1-58
Qinyuan L, Guiding L, Xiu C, Fangji X, Yong L, LiHua X, Yi J, Chenglin J (2015) Kineococcus gypseus sp. nov., isolated from saline sediment. Int J Syst Evol Microbiol 65:3703–3708. https://doi.org/10.1099/ijsem.0.000478
Ramesh S, Rajesh M, Mathivanan N (2009) Characterization of a thermostable alkaline protease produced by marine Streptomyces fungicidicus MML1614. Bioprocess Biosyst Eng 32:791–800. https://doi.org/10.1007/s00449-009-0305-1
Reddy GS, Prakash JS, Prabahar V, Matsumoto GI, Stackebrandt E, Shivaji S (2003) Kocuria polaris sp. nov., an orange-pigmented psychrophilic bacterium isolated from an Antarctic cyanobacterial mat sample. Int J Syst Evol Microbiol 53:183–187. https://doi.org/10.1099/ijs.0.02336-0
Schmidt EW (2014) The secret to a successful relationship: lasting chemistry between ascidians and their symbiotic bacteria. Invertebr Biol 134:88–102. https://doi.org/10.1111/ivb.12071
Selvam K, Vishnupriya B, Subash Chandra Bose V (2011) Screening and quantification of marine actinomycetes producing industrial enzymes amylase, cellulase and lipase from South cost of India. Int J Pharma Biol 2:1481–1487
Shirling EB, Gottileb D (1966) Methods for characterization of Streptomyces species. Int J Syst Bactriol 16:312–340. https://doi.org/10.1099/00207713-16-3-313
Sikorska J, Parker-Nance S, Davies-Coleman MT, Vining OB, Sikora AE, McPhail KL (2012) Antimicrobial rubrolides from a South African species of synoicum tunicate. J Nat Prod 75:1824–1827. https://doi.org/10.1021/np300580z
Suthindhiran K, Kannabiran K (2010) Diversity and exploration of bioactive actinomycetes in the Bay of Bengal of the Puducherry cost of India. Indian J Microbiol 50:76–82. https://doi.org/10.1007/s12088-010-0048-3
Tianero MDB, Kwan JC, Wyche TP, Presson AP, Koch M, Barrows LR, Bugni TS, Schmidt EW (2014) Species specificity of symbiosis and secondary metabolism in ascidians. ISME J 9:615–628. https://doi.org/10.1038/ismej.2014.152
Usama RA, Chen Y, Hannes H, Dina H, Timothy R, Ute H (2014) Actinomycetes from Red Sea sponges: sources for chemical and phylogenetic diversity. Mar Drugs 12:2771–2789. https://doi.org/10.3390/md12052771
Waksman SA (1961) The actinomycetes classification identification and description of genera and species. Williams & Wilkins company, Baltimore, pp 261–292. https://doi.org/10.1002/jps.2600501130
Wang YX, Wang HB, Zhang YQ, Xu LH, Jiang CL, Li WJ (2008) Rhodococcus kunmingensis sp. nov., an actinobacterium isolated from a rhizosphere soil. Int J Syst Evol Microbiol 58:1467–1471. https://doi.org/10.1099/ijs.0.65673-0
Williams ST, Cross T (1971) Actinomycetes, methods in microbiology, vol 4. Academic Press, New York
Wriston JC, Yellin TO (1973) L-Asparaginase: a review. Adv Enz 39:185–248. https://doi.org/10.1002/9780470122846.ch3
Wyche TP, Hou Y, Vazquez-Rivera E, Braun D, Bugni TS (2012) Peptidolipins B-F, antibacterial lipopeptides from an ascidian-derived Nocardia sp. J Nat Prod 75:735–740. https://doi.org/10.1021/np300016r
Youssef NH, Dunacn KE, Nagle DP, Savage KN, Knapp RM, McInerney MJ (2004) Comparision of methods to detect biosurfactant production by diverse microorganism. J Microbiol Methods 56:339–347. https://doi.org/10.1016/j.mimet.2003.11.001
Zhang Y, Adnani N, Braun DR, Ellis GA, Barns KJ, Parker-Nance S, Guzei IA, Bugni TS (2016) Micromonohalimanes A and B: Antibacterial halimane-type diterpenoids from a marine micromonospora species. J Nat Prod 79:2968–2972. https://doi.org/10.1021/acs.jnatprod.6b00555
Acknowledgements
The author’s great fully acknowledge the financial support given by the Ministry of Earth Sciences, Government of India, New Delhi to conduct the survey and research. The authors are thankful to Dr. M. A. Atmanand, Director, National Institute of Ocean Technology, Chennai for providing necessary facilities to perform this research.
Author information
Authors and Affiliations
Contributions
The research concept and the experiments were executed by BM, LA and KN. NVV and GD analysed the data and reviewed the manuscript. All of the authors approved the final manuscript.
Corresponding authors
Ethics declarations
Conflict of interest
The authors declare that they have no competing interests.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
About this article
Cite this article
Meena, B., Anburajan, L., Nitharsan, K. et al. Existence in cellulose shelters: industrial and pharmaceutical leads of symbiotic actinobacteria from ascidian Phallusia nigra, Andaman Islands. World J Microbiol Biotechnol 37, 120 (2021). https://doi.org/10.1007/s11274-021-03090-7
Received:
Accepted:
Published:
DOI: https://doi.org/10.1007/s11274-021-03090-7