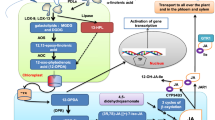
Abstract
The extracellular matrix of plants can contain the hydrophobic biopolymers lignin, suberin and/or cutin, which provide mechanical strength and limit water loss and pathogen invasion. Due to their remarkable chemical resistance, these polymers have a high potential in various biotechnological applications and can replace petrol-based resources, for example, in the packing industry. However, despite the importance of these polymers, the regulation of their precursor biosynthesis is far from being fully understood. This is particularly true for suberin and cutin, which hinders efforts to engineer their formation in plants and produce customised biopolymers. This review brings attention to knowledge gaps in the current research and highlights some of the most recent findings on transcription factors that regulate lignin, suberin and cutin precursor biosynthesis. Finally, we also briefly discuss how some of the remaining knowledge gaps can be closed.

Similar content being viewed by others
References
Agarwal T, Grotewold E, Doseff AI, Gray J (2016) MYB31/MYB42 syntelogs exhibit divergent regulation of phenylpropanoid genes in maize, sorghum and rice. Sci Rep 6:1–17
Ambavaram MM, Krishnan A, Trijatmiko KR, Pereira A (2011) Coordinated activation of cellulose and repression of lignin biosynthesis pathways in rice. Plant Physiol 155:916–931
Bang SW, Lee DK, Jung H, Chung PJ, Kim YS, Choi YD, Suh JW, Kim JK (2019) Overexpression of OsTF1L, a rice HD-Zip transcription factor, promotes lignin biosynthesis and stomatal closure that improves drought tolerance. Plant Biotechnol J 17:118–131
Bertella S, Luterbacher JS (2020) Lignin functionalization for the production of novel materials. Trends Chem 2:440–453
Cesarino I (2019) Structural features and regulation of lignin deposited upon biotic and abiotic stresses. Curr Obin Biotechnol 56:209–214
Cosgrove DJ (2005) Growth of the plant cell wall. Nat Rev Mol Cell Biol 6:850–861
del Río JC, Rencoret J, Prinsen P, Martínez ÁT, Ralph J, Gutiérrez A (2012) Structural characterization of wheat straw lignin as revealed by analytical pyrolysis, 2D-NMR, and reductive cleavage methods. J Agric Food Chem 60:5922–5935
Dixon RA, Barros J (2019) Lignin biosynthesis: old roads revisited and new roads explored. Open Biol 9:190215
Dubos C, Stracke R, Grotewold E, Weisshaar B, Martin C, Lepiniec L (2010) MYB transcription factors in Arabidopsis. Trends Plant Sci 15:573–581
Eder M, Lütz-Meindl U (2008) Pectin-like carbohydrates in the green alga Micrasterias characterized by cytochemical analysis and energy filtering TEM. J Microsc 231:201–214
Eder M, Tenhaken R, Driouich A, Lütz-Meindl U (2008) Occurrence and characterisation of arabinogalactan-like proteins and hemicelluloses in Micrasterias (Streptophyta). J Phycol 44:1221–1234
Eudes A, George A, Mukerjee P, Kim JS, Pollet B, Benke PI et al (2012) Biosynthesis and incorporation of side-chain-truncated lignin monomers to reduce lignin polymerization and enhance saccharification. P Biotechnol J 10:609–620
Figueiredo R, Araújo P, Llerena JPP, Mazzafera P (2019) Suberin and hemicellulose in sugarcane cell wall architecture and crop digestibility: A biotechnological perspective. Food Energy Secur 8(3). https://doi.org/10.1002/fes3.163
Figueiredo R, Llerena JPP, Kiyota E, Ferreira SS, Cardeli BR, de Souza SCR (2020) The sugarcane ShMYB78 transcription factor activates suberin biosynthesis in Nicotiana benthamiana. Plant Mol Biol 104:411–427
Fordyce PM, Gerber D, Tran D, Zheng J, Li H, DeRisi JL, Quake SR (2010) De novo identification and biophysical characterization of transcription-factor binding sites with microfluidic affinity analysis. Nat Biotechnol 28:970–975
Fornalé S, Lopez E, Salazar-Henao JE, Fernández-Nohales P, Rigau J, Caparros-Ruiz D (2014) AtMYB7, a new player in the regulation of UV-sunscreens in Arabidopsis thaliana. Plant Cell Physiol 55:507–516
Franke R, Briesen I, Wojciechowski T, Faust A, Yephremov A, Nawrath C, Schreiber L (2005) Apoplastic polyesters in Arabidopsis surface tissues–a typical suberin and a particular cutin. Phytochem 66:2643–2658
Garcia H, Ferreira R, Martins C, Sousa AF, Freire CS, Silvestre AJ et al (2014) Ex situ reconstitution of the plant biopolyester suberin as a film. Biomacromol 15:1806–1813
Gui J, Luo L, Zhong Y, Sun J, Umezawa T, Li L (2019) Phosphorylation of LTF1, an MYB transcription factor in populus, acts as a sensory switch regulating lignin biosynthesis in wood cells. Mol Plant 12:1325–1337
He J, Liu Y, Yuan D, Duan M, Liu Y, Shen Z et al (2020) An R2R3 MYB transcription factor confers brown planthopper resistance by regulating the phenylalanine ammonia-lyase pathway in rice. Proc Nat Acad Sci USA 117:271–277
Heredia-Guerrero JA, Heredia A, Domínguez E, Cingolani R, Bayer IS, Athanassiou A, Benítez JJ (2017) Cutin from agro-waste as a raw material for the production of bioplastics. J Exp Bot 68:5401–5410
Hong L, Brown J, Segerson NA, Rose JK, Roeder AH (2017) CUTIN SYNTHASE 2 Maintains Progressively Developing Cuticular Ridges in Arabidopsis Sepals. Mol Plant 10:560–574
Hu WJ, Harding SA, Lung J, Popko JL, Ralph J, Stokke DD et al (1999) Repression of lignin biosynthesis promotes cellulose accumulation and growth in transgenic trees. Nature Biotechnol 17:808–812
Jakobson L, Lindgren LO, Verdier G, Laanemets K, Brosché M, Beisson F, Kollist H (2016) BODYGUARD is required for the biosynthesis of cutin in Arabidopsis. New Phytol 211:614–626
Kai D, Jiang S, Low ZW, Loh XJ (2015) Engineering highly stretchable lignin-based electrospun nanofibers for potential biomedical applications. J Mat Chem B 3:6194–6204
Kang X, Kirui A, Widanage MCD, Mentink-Vigier F, Cosgrove DJ, Wang T (2019) Lignin-polysaccharide interactions in plant secondary cell walls revealed by solid-state NMR. Nat Commun 10:1–9
Kannangara R, Branigan C, Liu Y, Penfield T, Rao V, Mouille G et al (2007) The transcription factor WIN1/SHN1 regulates cutin biosynthesis in Arabidopsis thaliana. Plant Cell 19:1278–1294
Karaba A (2007) Improvement of water use efficiency in rice and tomato using Arabidopsis wax biosynthetic genes and transcription factors. PhD thesis. Wageningen University, Wageningen, The Netherlands
Karlen SD, Zhang C, Peck ML, Smith RA, Padmakshan D, Helmich K et al (2016) Monolignol ferulate conjugates are naturally incorporated into plant lignins. Sci. Adv. 2:e1600393
Kim SH, Lam PY, Lee MH, Jeon HS, Tobimatsu Y, Park OK (2020) The Arabidopsis R2R3 MYB Transcription Factor MYB15 Is a Key Regulator of Lignin Biosynthesis in Effector-Triggered Immunity. Front Plant Sci 11:1456
Klempnauer KH, Gonda TJ, Bishop JM (1982) Nucleotide sequence of the retroviral leukemia gene v-myb and its cellular progenitor c-myb: the architecture of a transduced oncogene. Cell 31:453–463
Kosma DK, Murmu J, Razeq FM, Santos P, Bourgault R, Rowland MI, O, (2014) AtMYB41 activates ectopic suberin synthesis and assembly in multiple plant species and cell types. Plant J 80:216–229
Lashbrooke J, Cohen H, Levy-Samocha D, Tzfadia O, Panizel I, Zeisler V et al (2016) MYB107 and MYB9 homologs regulate suberin deposition in angiosperms. Plant Cell 28:2097–2116
Legay S, Guerriero G, Deleruelle A, Lateur M, Evers D, André CM, Hausman J-F (2015) Apple russeting as seen through the RNA-seq lens: strong alterations in the exocarp cell wall. Plant Mol Biol 88:21–40
Legay S, Guerriero G, André C, Guignard C, Cocco E, Charton S, Boutry M, Hausman RO, JF, (2016) MdMyb93 is a regulator of suberin deposition in russeted apple fruit skins. New Phytol 212:977–991
Li Y, Beisson F, Koo AJ, Molina I, Pollard M, Ohlrogge J (2007) Identification of acyltransferases required for cutin biosynthesis and production of cutin with suberin-like monomers. Proc Natl Acad Sci USA 104:18339–18344
Lütz-Meindl U (2016) Micrasterias as a model system in plant cell biology. Front Plant Sci 7:999
Ma Z, Wang J, Li C, Yang Y, Liu X, Zhao C, Chen D (2019) New sight on the lignin torrefaction pretreatment: Relevance between the evolution of chemical structure and the properties of torrefied gaseous, liquid, and solid products. Bioresour Technol 288:121528
Martone PT, Estevez JM, Lu F, Ruel K, Denny MW, Somerville C, Ralph J et al (2009) Discovery of lignin in seaweed reveals convergent evolution of cell-wall architecture. Curr Biol 19:169–175
Miyamoto T, Tobimatsu Y, Umezawa T (2020) MYB-mediated regulation of lignin biosynthesis in grasses. Curr Plant Biol 24:100174
Molina I, Beisson-Li Y, Beisson F, Ohlrogge J, Pollard M (2009) Identification of an Arabidopsis feruloyl-CoA transferase required for suberin synthesis. Plant Physiol 151:1317–1328
Morse AM, Whetten RW, Dubos C, Campbell MM (2009) Post-translational modification of an R2R3-MYB transcription factor by a MAP Kinase during xylem development. New Phytol 183:1001–1013
Nakano Y, Yamaguchi M, Endo H, Rejab NA, Ohtani M (2015) NAC-MYB-based transcriptional regulation of secondary cell wall biosynthesis in land plants. Front Plant Sci 6:288
Natarajan P, Akinmoju TA, Nimmakayala P, Lopez-Ortiz C, Garcia-Lozano M, Thompson BJ et al (2020) Integrated metabolomic and transcriptomic analysis to characterize cutin biosynthesis between low-and high-cutin genotypes of Capsicum chinense Jacq. Int J Mol Sci 21:1397
Oshima Y, Mitsuda N (2013) The MIXTA-like transcription factor MYB16 is a major regulator of cuticle formation in vegetative organs. Plant Signal Behav 8:e26826
Perkins ML, Schuetz M, Unda F, Smith RA, Sibout R, Hoffmann NJ (2020) Dwarfism of high-monolignol Arabidopsis plants is rescued by ectopic LACCASE overexpression. Plant direct 4:e00265
Philippe G, Gaillard C, Petit J, Geneix N, Dalgalarrondo M, Bres C et al (2016) Ester cross-link profiling of the cutin polymer of wild-type and cutin synthase tomato mutants highlights different mechanisms of polymerization. Plant Physiol 170:807–820
Poovaiah CR, Nageswara-Rao M, Soneji JR, Baxter HL, Stewart CN Jr (2014) Altered lignin biosynthesis using biotechnology to improve lignocellulosic biofuel feedstocks. Plant Biotechnol J 12:1163–1173
Popper ZA, Michel G, Hervé C, Domozych DS, Willats WGT, Tuohy MG (2011) Evolution and diversity of plant cell walls: from algae to flowering plants. Ann Rev Plant Biol 62:567–590
Quan M, Du Q, Xiao L, Lu W, Wang L, Xie J et al (2019) Genetic architecture underlying the lignin biosynthesis pathway involves noncoding RNA s and transcription factors for growth and wood properties in populus. Plant Biotechnol J 17:302–315
Ralph J, Lapierre C, Boerjan W (2019) Lignin structure and its engineering. Curr Opin Biotechnol 56:240–249
Rao X, Chen X, Shen H, Ma Q, Li G, Tang Y, Pena M, York W, Frazier TP, Lenaghan S, Xiao X, Chen F, Dixon RA (2019) Gene regulatory networks for lignin biosynthesis in switchgrass (Panicum virgatum). Plant Biotechnol J 17(3):580–593
Rautengarten C, Ebert B, Ouellet M, Nafisi M, Baidoo EE, Benke P et al (2012) Arabidopsis deficient in cutin ferulate encodes a transferase required for feruloylation of ω-hydroxy fatty acids in cutin polyester. Plant Physiol 158:654–665
Renault H, Alber A, Horst NA, Lopes AB, Fich EA, Kriegshauser L et al (2017) A phenol-enriched cuticle is ancestral to lignin evolution in land plants. Nat Commun 8:1–8
Saleme MDLS, Cesarino I, Vargas L, Kim H, Vanholme R, Goeminne G et al (2017) Silencing CAFFEOYL SHIKIMATE ESTERASE affects lignification and improves saccharification in poplar. Plant Physiol 175:1040–1057
Sørensen I, Pettolino FA, Bacic A, Ralph J, Lu F, O’Neill, et al (2011) The charophycean green algae provide insights into the early origins of plant cell walls. Plant J 68:201–211
Tamagnone L, Merida A, Stacey N, Plaskitt K, Parr A, Chang CF et al (1998) Inhibition of phenolic acid metabolism results in precocious cell death and altered cell morphology in leaves of transgenic tobacco plants. Plant Cell 10:1801–1816
To A, Joubès J, Thueux J, Kazaz S, Baud LL, S, (2020) AtMYB92 enhances fatty acid synthesis and suberin deposition in leaves of Nicotiana benthamiana. Plant J 103:660–676
Tobimatsu Y, Schuetz M (2019) Lignin polymerization: how do plants manage the chemistry so well? Curr Opin Biotechnol 56:75–81
Ursache R, Teixeira CDJV, Tendon VD, Gully K, De Bellis D, Schmid-Siegert E et al (2021) GDSL-domain proteins have key roles in suberin polymerization and degradation. Nat Plants 7:353–364
Vanholme R, De Meester B, Ralph J, Boerjan W (2019) Lignin biosynthesis and its integration into metabolism. Curr Opin Biotechnol 56:230–239
Vishwanath SJ, Delude C, Domergue F, Rowland O (2015) Suberin: biosynthesis, regulation, and polymer assembly of a protective extracellular barrier. Plant Cell Rep 34:573–586
Wallace G, Fry SC (1994) Phenolic components of the plant cell wall. Int Rev Cytol 151:229–267
Wang S, Zhang S, Xiao A, Rasmussen M, Skidmore C, Zhan J (2015) Metabolic engineering of Escherichia coli for the biosynthesis of various phenylpropanoid derivatives. Metab Eng 29:153–159
Welker CM, Balasubramanian VK, Petti C, Rai KM, DeBolt S, Mendu V (2015) Engineering Plant Biomass Lignin Content and Composition for Biofuels and Bioproducts. Energies 8:7654–7676
Withers S, Lu F, Kim H, Zhu Y, Ralph J, Wilkerson CG (2012) Identification of grass-specific enzyme that acylates monolignols with p-coumarate. J Biol Chem 287:8347–8355
Wu R, Li S, He S, Waßmann F, Yu C, Qin G et al (2011) CFL1, a WW domain protein, regulates cuticle development by modulating the function of HDG1, a class IV homeodomain transcription factor, in rice and Arabidopsis. Plant Cell 23:3392–3411
Xie M, Zhang J, Tschaplinski TJ, Tuskan GA, Chen JG, Muchero W (2018) Regulation of lignin biosynthesis and its role in growth-defense tradeoffs. Front Plant Sci 9:1427
Xin A, Herburger K (2021) Mini review: Transport of hydrophobic polymers into the plant apoplast. Front Plant Sci 11:2059
Xin A, Fei Y, Molnar A, Fry SC (2021) Cutin:cutin-acid endo-transacylase (CCT), a cuticle-remodelling enzyme activity in the plant epidermis. Biochem J 478:777–798
Yamamoto M, Tomiyama H, Koyama A, Okuizumi H, Liu S, Vanholme R et al (2020) A century-old mystery unveiled: sekizaisou is a natural lignin mutant. Plant Physiol 182:1821–1828
Yan J, Aznar A, Chalvin C, Birdseye DS, Baidoo EE, Eudes A et al (2018) Increased drought tolerance in plants engineered for low lignin and low xylan content. Biotechnol Biofuels 11:1–11
Yang S, Johnston N, Talideh E, Mitchell S, Jeffree C, Goodrich J, Ingram G (2008) The endosperm-specific ZHOUPI gene of Arabidopsis thaliana regulates endosperm breakdown and embryonic epidermal development. Development 135:3501–3509
Yeats TH, Martin LB, Viart HM, Isaacson T, He Y, Zhao L et al (2012) The identification of cutin synthase: formation of the plant polyester cutin. Nat Chem Biol 8:609–611
Zhang P, Wang R, Yang X, Ju Q, Li W, Lü S et al (2020) The R2R3-MYB transcription factor AtMYB49 modulates salt tolerance in Arabidopsis by modulating the cuticle formation and antioxidant defence. Plant, Cell Environ 43:1925–1943
Acknowledgements
We dedicate this paper to Prof. Ursula Lütz-Meindl whose outstanding research on cell wall biology truly inspired both senior and early career scientists to explore various aspects in this fascinating field—from Charophyte green algae to vascular plants.
Funding
This work was supported by the Villum Foundation project TIPorNOT (00023089) to KH.
Author information
Authors and Affiliations
Contributions
AX drafted the manuscript and KH edited it. Both authors approved the final manuscript.
Corresponding author
Ethics declarations
Conflicts of interest
Both authors declare that the work was conducted in the absence of any commercial or financial relationships that could be construed as a potential conflict of interest.
Consent to participate
Both authors agree to the preparation of this manuscript and review.
Consent for publication
Both authors agree to the publication this article if accepted.
Additional information
Handling Editor: Andreas Holzinger
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
About this article
Cite this article
Xin, A., Herburger, K. Precursor biosynthesis regulation of lignin, suberin and cutin. Protoplasma 258, 1171–1178 (2021). https://doi.org/10.1007/s00709-021-01676-4
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00709-021-01676-4