Abstract
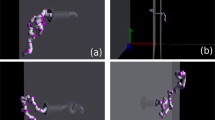
The pulling translocation of a polymer globule through a nanopore is simulated by using the Langevin dynamics method. The head of the polymer is pulled with a constant pulling force or with a constant pulling speed. The scaling relations between the polymer translocation time τ and polymer length N and between τ and the pulling force F/speed v are investigated. For translocation induced by a constant pulling force, we obtain τ ~F−β with β = 0.88 at large pulling forces and τ ~Nα for long polymer chains. However, the exponent α is dependent on the intrapolymer interaction strength. For the translocation induced by a constant pulling speed, we obtain τ ~v−0.92 at large pulling speeds and α = 1.24 for long polymer chains. The results show that the conformational changes of the polymer chain during translocation and the entropic and enthalpic forces due to the straight conformation at the trans side play important roles in the pulling translocation of polymer globules.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout









Similar content being viewed by others
References
Stewart M. Ratcheting MRNA out of the nucleus. Mol Cell. 2007;25:327.
Köhler A, Hurt E. Exporting RNA from the nucleus to the cytoplasm. Nat Rev Mol Cell Biol 2007;8:761.
Carmody SR, Wente SR. MRNA nuclear export at a glance. J Cell Sci. 2009;122:1933.
Ma J, Liu Z, Michelotti N, Pitchiaya S, Veerapaneni R, Androsavich JR, et al. High-resolution three-dimensional mapping of mRNA export through the nuclear pore. Nat Commun 2013;4:2414.
Ledwith BJ, Manam S, Troilo PJ, Barnum AB, Pauley CJ, Griffiths TG, et al. Plasmid DNA vaccines: investigation of integration into host cellular DNA following intramuscular injection in mice. Intervirology. 2000;43:258.
Wang Z, Troilo PJ, Wang X, Griffiths TG, Pacchione SJ, Barnum AB. et al. Detection of integration of plasmid DNA into host genomic DNA following intramuscular injection and electroporation. Gene Ther. 2004;11:711
Zhao H, Speir JA, Matsui T, Lin Z, Liang L, Lynn AY, et al. Structure of a bacterial virus DNA-injection protein complex reveals a decameric assembly with a constricted molecular channel. PLoS ONE. 2016;11:e0149337.
Agarraberes FA, Dice JF. Protein translocation across membranes. Biochim Biophys Acta. 2001;1513:1.
Helenius J, Ng DTW, Marolda CL, Walter P, Valvano MA, Aebi M. Translocation of lipid-linked oligosaccharides across the ER membrane requires Rft1 protein. Nature. 2002;415:447.
Schnell DJ, Hebert DN. Protein translocons: multifunctional mediators of protein translocation across membranes. Cell. 2003;112:491.
Craig EA. Hsp70 at the membrane: driving protein translocation. BMC Biol. 2018;16:11.
Gauthier MG, Slater GW. Sequence effects on the forced translocation of heteropolymers through a small channel. J Chem Phys. 2008;128:175103
Kumar R, Chaudhuri A, Kapri R. Sequencing of semiflexible polymers of varying bending rigidity using patterned pores. J Chem Phys. 2018;148:164901
Han S, Mahato RI, Sung YK, Kim SW. Development of biomaterials for gene therapy. Mol Ther. 2000;2:302.
Wong SY, Pelet JM, Putnam D. Polymer systems for gene delivery—past, present, and future. Prog Polym Sci. 2007;32:799
Sui M, Liu W, Shen Y. Nuclear drug delivery for cancer chemotherapy. J Control Release. 2011;155:227.
Parelkar SS, Letteri R, Chan-Seng D, Zolochevska O, Ellis J, Figueiredo M, et al. Polymer-peptide delivery platforms: effect of oligopeptide orientation on polymer-based DNA delivery. Biomacromolecules. 2014;15:1328.
Bezrukov SM, Vodyanoy I, Parsegian VA. Counting polymers moving through a single ion channel. Nature. 1994;370:279.
Kasianowicz JJ, Brandin E, Branton D, Deamer DW. Characterization of individual polynucleotide molecules using a membrane channel. in Proceedings of the National Academy of Sciences of the United States of America, Vol. 93. National Academy of Sciences, 1996. p. 13770–3.
Sung W, Park PJ. Polymer translocation through a pore in a membrane. Phys Rev Lett. 1996;77:783
Muthukumar M. Polymer translocation through a hole. J Chem Phys. 1999;111:10371.
Meller A, Nivon L, Branton D. Voltage-driven DNA translocations through a nanopore. Phys Rev Lett. 2001;86:3435.
Chuang J, Kantor Y, Kardar M. Anomalous dynamics of translocation. Phys Rev E. 2001;65:011802.
Slonkina E, Kolomeisky AB. Polymer translocation through a long nanopore. J Chem Phys. 2003;118:7112.
Storm AJ, Storm C, Chen J, Zandbergen H, Joanny JF, Dekker C. Fast DNA translocation through a solid-state nanopore. Nano Lett. 2005;5:1193.
Guillouzic S, Slater GW. Polymer translocation in the presence of excluded volume and explicit hydrodynamic interactions. Phys Lett A. 2006;359:261.
Luo MB. Translocation of polymer chains through interacting nanopores. Polymer. 2007;48:7679.
He YD, Qian HJ, Lu ZY, Li ZS. Polymer translocation through a nanopore in mesoscopic simulations. Polymer. 2007;48:3601.
Dekker C. Solid-state nanopores. Nat Nanotechnol 2007;2:209.
Luo K, Ala-Nissila T, Ying SC, Bhattacharya A. Dynamics of DNA translocation through an attractive nanopore. Phys Rev E. 2008;78:061911.
Wong CTA, Muthukumar M. Polymer translocation through α-hemolysin pore with tunable polymer-pore electrostatic interaction. J Chem Phys. 2010;133:045101.
Panja D, Barkema GT. Simulations of two-dimensional unbiased polymer translocation using the bond fluctuation model. J Chem Phys. 2010;132:014902.
Ying Y, Zhang J, Gao R, Long Y. Nanopore‐based sequencing and detection of nucleic acids. Angew Chem Int Ed. 2013;52:2.
Nagarajan K, Chen SB. Polyelectrolyte translocation through a tortuous nanopore. J Phys Chem B. 2019;123:9031.
Clarke J, Wu HC, Jayasinghe L, Patel A, Reid S, Bayley H. Continuous base Identification for single-molecule nanopore DNA sequencing. Nat Nanotechnol. 2009;4:265.
Postma HWC. Rapid sequencing of individual DNA molecules in graphene nanogaps. Nano Lett. 2010;10:420.
Derrington IM, Butler TZ, Collins MD, Manrao E, Pavlenok M, Niederweis M, et al. Nanopore DNA sequencing with MspA. Proc Natl Acad Sci USA 2010;107:16060.
Gu Z, Zhang Y, Luan B, Zhou R. DNA translocation through single-layer boron nitride nanopores. Soft Matter. 2016;12:817.
Anderson BN, Muthukumar M, Meller A. PH tuning of DNA translocation time through organically functionalized nanopores. ACS Nano. 2013;7:1408.
Mereuta L, Roy M, Asandei A, Lee JK, Park Y, Andricioaei I, et al. Slowing down single-molecule trafficking through a protein nanopore reveals intermediates for peptide translocation. Sci. Rep. 2014;4:3885.
Liu Y, Yobas L. Slowing DNA translocation in a nanofluidic field-effect transistor. ACS Nano. 2016;10:3985.
Liu L, Yang C, Zhao K, Li J, Wu HC. Ultrashort single-walled carbon nanotubes in a lipid bilayer as a new nanopore sensor. Nat Commun. 2013;4:2989.
Wanunu M, Sutin J, McNally B, Chow A, Meller A. DNA translocation governed by interactions with solid-state nanopores. Biophys J. 2008;95:4716.
Carson S, Wilson J, Aksimentiev A, Wanunu M. Smooth DNA transport through a narrowed pore geometry. Biophys J. 2014;107:2381.
Gopinathan A, Kim YW. Polymer translocation in crowded environments. Phys Rev Lett. 2007;99:228106.
Ying YL, Li DW, Li Y, Lee JS, Long YT. Enhanced translocation of Poly(Dt)45 through an α-hemolysin nanopore by binding with antibody. Chem Commun. 2011;47:5690.
Polson JM, McCaffrey ACM. Polymer translocation dynamics in the quasi-static limit. J Chem Phys. 2013;138:174902.
Sun LZ, Cao WP, Luo MB. Free energy landscape for the translocation of polymer through an interacting pore. J Chem Phys. 2009;131:194904.
Oberhauser AF, Marszalek PE, Carrion-Vazquez M, Fernandez JM. Single protein misfolding events captured by atomic force microscopy. Nat Struct Biol. 1999;6:1025.
Cieplak M, Filipek S, Janovjak H, Krzyśko KA. Pulling single bacteriorhodopsin out of a membrane: comparison of simulation and experiment. Biochim Biophys Acta. 2006;1758:537.
Keyser UF, Koeleman BN, Van Dorp S, Krapf D, Smeets RMM, Lemay SG, et al. Direct force measurements on DNA in a solid-state nanopore. Nat Phys. 2006;2:473.
Bulushev RD, Marion S, Petrova E, Davis SJ, Maerkl SJ, Radenovic A. Single molecule localization and discrimination of DNA-protein complexes by controlled translocation through nanocapillaries. Nano Lett. 2016;16:7882.
Menais T. Polymer translocation under a pulling force: scaling arguments and threshold forces. Phys Rev E. 2018;97:022501.
Ritort F. Single-molecule experiments in biological physics: methods and applications. J Phys Condens Matter. 2006;18:R531.
Sischka A, Spiering A, Khaksar M, Laxa M, König J, Dietz K-J, et al. Dynamic translocation of ligand-complexed DNA through solid-state nanopores with optical tweezers. J Phys Condens Matter. 2010;22:454121.
Bulushev RD, Marion S, Radenovic A. Relevance of the drag force during controlled translocation of a DNA-Protein complex through a glass nanocapillary. Nano Lett. 2015;15:7118.
Carbone P, Avendaño C. Coarse-grained methods for polymeric materials: enthalpy- and entropy-driven models. Wiley Interdiscip Rev Comput Mol Sci. 2014;4:62.
Lima NW, Gutierres LI, Gonzalez RI, Müller S, Thomaz RS, Bringa EM, et al. Molecular dynamics simulation of polymerlike thin films irradiated by fast ions: a comparison between FENE and Lennard-Jones potentials. Phys Rev B. 2016;94:195417.
Weeks JD, Chandler D, Andersen HC. Role of repulsive forces in determining the equilibrium structure of simple liquids. J Chem Phys. 1971;54:5237.
Ermann N, Hanikel N, Wang V, Chen K, Weckman NE, Keyser UF. Promoting single-file DNA translocations through nanopores using electro-osmotic flow. J Chem Phys. 2018;149:163311.
Luo K, Ala-Nissila T, Ying SC, Metzler R. Driven polymer translocation through nanopores: slow-vs.-fast dynamics. EPL. 2009;88:68006.
Wu F, Fu Y, Yang X, Sun LZ, Luo MB. Driven translocation of semiflexible polyelectrolyte through a nanopore. J Polym Sci Part B Polym Phys. 2019;57:912.
Milchev A. Single-polymer dynamics under constraints: scaling theory and computer experiment. J Phys Condens Matter. 2011;23:103101.
Zhang Y, Zhou H, Ou-Yang Z-C. Stretching single-stranded DNA: interplay of electrostatic, base-pairing, and base-pair stacking interactions. Biophys J. 2001;81:1133–43.
Chen H, Yuan G, Winardhi RS, Yao M, Popa I, Fernandez JM, et al. Dynamics of equilibrium folding and unfolding transitions of titin immunoglobulin domain under constant forces. J Am Chem Soc. 2015;137:3540–6.
Vocks H, Panja D, Barkema GT, Ball RC. Pore-blockade times for field-driven polymer translocation. J Phys Condens Matter. 2008;20:095224.
Palyulin VV, Ala-Nissila T, Metzler R. Polymer translocation: the first two decades and the recent diversification. Soft Matter. 2014;10:9016.
Luo MB, Tsehay DA, Sun LZ. Temperature dependence of the translocation time of polymer through repulsive nanopores. J Chem Phys. 2017;147:034901.
Acknowledgements
This work was supported by the National Natural Science Foundation of China under grant No. 11974305.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors declare no competing interests.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
About this article
Cite this article
Chen, X., Chen, J., Zhuo, BY. et al. Simulation study for the pulling translocation of a polymer globule. Polym J 53, 1047–1056 (2021). https://doi.org/10.1038/s41428-021-00502-1
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41428-021-00502-1
This article is cited by
-
Translocation of a Self-propelled Polymer through a Narrow Pore
Chinese Journal of Polymer Science (2022)