Abstract
MicroRNA160 is a class of nitrogen-starvation responsive genes which governs establishment of root system architecture by down-regulating AUXIN RESPONSE FACTOR genes (ARF10, ARF16 and ARF17) in plants. The high copy number of MIR160 variants discovered by us from land plants, especially polyploid crop Brassicas, posed questions regarding genesis, duplication, evolution and function. Absence of studies on impact of whole genome and segmental duplication on retention and evolution of MIR160 homologs in descendent plant lineages prompted us to undertake the current study. Herein, we describe ancestry and fate of MIR160 homologs in Brassicaceae in context of polyploidy driven genome re-organization, copy number and differentiation. Paralogy amongst Brassicaceae MIR160a, MIR160b and MIR160c was inferred using phylogenetic analysis of 468 MIR160 homologs from land plants. The evolutionarily distinct MIR160a was found to represent ancestral form and progenitor of MIR160b and MIR160c. Chronology of evolutionary events resulting in origin and diversification of genomic loci containing MIR160 homologs was delineated using derivatives of comparative synteny. A prescient model for causality of segmental duplications in establishment of paralogy in Brassicaceae MIR160, with whole genome duplication accentuating the copy number increase, is being posited in which post-segmental duplication events viz. differential gene fractionation, gene duplications and inversions are shown to drive divergence of chromosome segments. While mutations caused the diversification of MIR160a, MIR160b and MIR160c, duplicated segments containing these diversified genes suffered gene rearrangements via gene loss, duplications and inversions. Yet the topology of phylogenetic and phenetic trees were found congruent suggesting similar evolutionary trajectory. Over 80% of Brassicaceae genomes and subgenomes showed a preferential retention of single copy each of MIR160a, MIR160b and MIR160c suggesting functional relevance. Thus, our study provides a blue-print for reconstructing ancestry and phylogeny of MIRNA gene families at genomics level and analyzing the impact of polyploidy on organismal complexity. Such studies are critical for understanding the molecular basis of agronomic traits and deploying appropriate candidates for crop improvement.








Similar content being viewed by others
References
Al-Shehbaz IA (2012) A generic and tribal synopsis of the Brassicaceae (Cruciferae). Taxon 61:931–954. https://doi.org/10.1002/tax.615002
Altschul SF, Gish W, Miller W et al (1990) Basic local alignment search tool. J Mol Biol. https://doi.org/10.1016/S0022-2836(05)80360-2
Anand S, Lal M, Das S (2019) Comparative genomics reveals origin of MIR159A–MIR159B paralogy, and complexities of PTGS interaction between miR159 and target GA-MYBs in Brassicaceae. Mol Genet Genomics 294:693–714. https://doi.org/10.1007/s00438-019-01540-4
Bailey JA, Gu Z, Clark RA et al (2002) Recent segmental duplications in the human genome. Science 297:1003–1007. https://doi.org/10.1126/science.1072047
Bailey CD, Koch MA, Mayer M et al (2006) Toward a global phylogeny of the Brassicaceae. Mol Biol Evol 23:2142–2160. https://doi.org/10.1093/molbev/msl087
Bird KA, VanBuren R, Puzey JR, Edger PP (2018) The causes and consequences of subgenome dominance in hybrids and recent polyploids. New Phytol 220:87–93. https://doi.org/10.1111/nph.15256
Bouckaert R, Heled J, Kühnert D et al (2014) BEAST 2: a software platform for Bayesian evolutionary analysis. PLoS Comput Biol 10:e1003537. https://doi.org/10.1371/journal.pcbi.1003537
Bray N, Dubchak I, Pachter L (2003) AVID: a global alignment program. Genome Res 13:97–102. https://doi.org/10.1101/gr.789803
Brosius J (1991) Retroposons—seeds of evolution. Science 251:753. https://doi.org/10.1126/science.1990437
Bustos-sanmamed P, Mao G, Deng Y et al (2013) Overexpression of miR160 affects root growth and nitrogen-fixing nodule number in Medicago truncatula. Funct Plant Biol 40:1208–1220. https://doi.org/10.1071/FP13123
Chen L, Chen L, Zhang X et al (2018) Identification of miRNAs that regulate silique development in Brassica napus. Plant Sci 269:106–117. https://doi.org/10.1016/j.plantsci.2018.01.010
Cheng F, Liu S, Wu J et al (2011) BRAD, the genetics and genomics database for Brassica plants. BMC Plant Biol 11:136. https://doi.org/10.1186/1471-2229-11-136
Cheng F, Wu J, Fang L et al (2012) Biased gene fractionation and dominant gene expression among the subgenomes of Brassica rapa. PLoS ONE 7:e36442. https://doi.org/10.1371/journal.pone.0036442
Cheng F, Wu J, Wang X (2014) Genome triplication drove the diversification of Brassica plants. Hortic Res 1:14024. https://doi.org/10.1038/hortres.2014.24
Cheng F, Wu J, Cai X et al (2018) Gene retention, fractionation and subgenome differences in polyploid plants. Nat Plants 4:258–268. https://doi.org/10.1038/s41477-018-0136-7
Conesa A, Götz S, García-Gómez JM et al (2005) Blast2GO: a universal tool for annotation, visualization and analysis in functional genomics research. Bioinformatics. https://doi.org/10.1093/bioinformatics/bti610
Damodharan S, Zhao D, Arazi T (2016) A common miRNA160-based mechanism regulates ovary patterning, floral organ abscission and lamina outgrowth in tomato. Plant J 86:458–471. https://doi.org/10.1111/tpj.13127
Dexheimer PJ, Cochella L (2020) MicroRNAs: from mechanism to organism. Front Cell Dev Biol. https://doi.org/10.3389/fcell.2020.00409
Dixon GR (2007) Vegetable brassicas and related crucifers. In: Crop production science in horticulture., vol 14. CABI, Walllingford, Oxfordshire, p 327
Drouin G, Dover GA (1990) Independent gene evolution in the potato actin gene family demonstrated by phylogenetic procedures for resolving gene conversions and the phylogeny of angiosperm actin genes. J Mol Evol 31:132–150. https://doi.org/10.1007/BF02109482
Drummond AJ, Suchard MA, Xie D, Rambaut A (2012) Bayesian phylogenetics with BEAUti and the BEAST 1.7. Mol Biol Evol 29:1969–1973. https://doi.org/10.1093/molbev/mss075
Edger PP, Poorten TJ, VanBuren R et al (2019) Origin and evolution of the octoploid strawberry genome. Nat Genet 51:541–547. https://doi.org/10.1038/s41588-019-0356-4
Fitch WM (1970) Distinguishing homologous from analogous proteins. Syst Zool 19:99–113. https://doi.org/10.2307/2412448
Franzke A, Lysak MA, Al-Shehbaz IA et al (2011) Cabbage family affairs: the evolutionary history of Brassicaceae. Trends Plant Sci 16:108–116. https://doi.org/10.1016/j.tplants.2010.11.005
Garcia-Vallvé S, Puigbo P (2002) Online software D-UPGMA developed in Biochemistry and Biotechnology Department. Undefined
Guerra-Assunção JA, Enright AJ (2012) Large-scale analysis of microRNA evolution. BMC Genomics 13:218. https://doi.org/10.1186/1471-2164-13-218
Guo Y, Liu J, Zhang J et al (2017) Selective modes determine evolutionary rates, gene compactness and expression patterns in Brassica. Plant J 91:34–44. https://doi.org/10.1111/tpj.13541
Guo Z, Kuang Z, Wang Y et al (2020) PmiREN: a comprehensive encyclopedia of plant miRNAs. Nucleic Acids Res. https://doi.org/10.1093/nar/gkz894
Hendelman A, Buxdorf K, Stav R et al (2012) Inhibition of lamina outgrowth following Solanum lycopersicum AUXIN RESPONSE FACTOR 10 (SlARF10) derepression. Plant Mol Biol 78:561–576. https://doi.org/10.1007/s11103-012-9883-4
Hohmann N, Wolf EM, Lysak MA, Koch MA (2015) A time-calibrated road map of Brassicaceae species radiation and evolutionary history. Plant Cell 27:2770–2784. https://doi.org/10.1105/tpc.15.00482
Huang C-H, Sun R, Hu Y et al (2016a) Resolution of Brassicaceae phylogeny using nuclear genes uncovers nested radiations and supports convergent morphological evolution. Mol Biol Evol 33:394–412. https://doi.org/10.1093/molbev/msv226
Huang J, Zhiyong L, Dazhong Z (2016b) Deregulation of the OsmiR160 target gene OsARF18 causes growth and developmental defects with an alteration of auxin signaling in rice. Sci Rep 6:1–14. https://doi.org/10.1038/srep29938
Jaccard P (1908) Nouvelles recherches sur la distribution florale. Bull Soc Vaud Sci Nat 44:223–270
Jain A, Das S (2016) Synteny and comparative analysis of miRNA retention, conservation, and structure across Brassicaceae reveals lineage- and sub-genome-specific changes. Funct Integr Genomics 16:253–268. https://doi.org/10.1007/s10142-016-0484-1
Jiang N, Bao Z, Zhang X et al (2004) Pack-MULE transposable elements mediate gene evolution in plants. Nature 431:569–573. https://doi.org/10.1038/nature02953
Jiao Y, Paterson AH (2014) Polyploidy-associated genome modifications during land plant evolution. Philos Trans R Soc B Biol Sci. https://doi.org/10.1098/rstb.2013.0355
Joshi G, Chauhan C, Das S (2018) Microsynteny analysis to understand evolution and impact of polyploidization on MIR319 family within Brassicaceae. Dev Genes Evol 228:227–242. https://doi.org/10.1007/s00427-018-0620-0
Kagale S, Robinson SJ, Nixon J et al (2014) Polyploid evolution of the Brassicaceae during the Cenozoic era. Plant Cell 26:2777–2791. https://doi.org/10.1105/tpc.114.126391
Katoh K, Standley DM (2013) MAFFT multiple sequence alignment software version 7: improvements in performance and usability. Mol Biol Evol 30:772–780. https://doi.org/10.1093/molbev/mst010
Katoh K, Rozewicki J, Yamada KD (2018) MAFFT online service: multiple sequence alignment, interactive sequence choice and visualization. Brief Bioinform 20:1160–1166. https://doi.org/10.1093/bib/bbx108
Kejnovsky E, Leitch IJ, Leitch AR (2009) Contrasting evolutionary dynamics between angiosperm and mammalian genomes. Trends Ecol Evol 24:572–582
Khraiwesh B, Zhu JK, Zhu J (2012) Role of miRNAs and siRNAs in biotic and abiotic stress responses of plants. Biochim Biophys Acta 1819:137–148. https://doi.org/10.1016/j.bbagrm.2011.05.001
Lallemand T, Leduc M, Landès C et al (2020) An overview of duplicated gene detection methods: why the duplication mechanism has to be accounted for in their choice. Genes (basel) 11:1046
Langham RJ, Walsh J, Dunn M et al (2004) Genomic duplication, fractionation and the origin of regulatory novelty. Genetics 166:935–945. https://doi.org/10.1534/genetics.166.2.935
Li A, Mao L (2007) Evolution of plant microRNA gene families. Cell Res 17:212–218. https://doi.org/10.1038/sj.cr.7310113
Liang G, He H, Yu D (2012) Identification of nitrogen starvation-responsive MicroRNAs in Arabidopsis thaliana. PLoS ONE. https://doi.org/10.1371/journal.pone.0048951
Lin Y, Lai Z, Tian Q et al (2015) Endogenous target mimics down-regulate miR160 mediation of ARF10, -16, and -17 cleavage during somatic embryogenesis in Dimocarpus longan Lour. Front Plant Sci. https://doi.org/10.3389/fpls.2015.00956
Liu PP, Montgomery TA, Fahlgren N et al (2007) Repression of AUXIN RESPONSE FACTOR10 by microRNA160 is critical for seed germination and post-germination stages. Plant J 52:133–146. https://doi.org/10.1111/j.1365-313X.2007.03218.x
Liu X, Huang J, Wang Y et al (2010) The role of floral organs in carpels, an Arabidopsis loss-of-function mutation in MicroRNA160a, in organogenesis and the mechanism regulating its expression. Plant J 62:416–428. https://doi.org/10.1111/j.1365-313X.2010.04164.x
Liu X, Zhang H, Zhao Y et al (2013) Auxin controls seed dormancy through stimulation of abscisic acid signaling by inducing ARF-mediated ABI3 activation in Arabidopsis. Proc Natl Acad Sci USA 110:15485–15490. https://doi.org/10.1073/pnas.1304651110
Liu X, Dong X, Liu Z et al (2016) Repression of ARF10 by microRNA160 plays an important role in the mediation of leaf water loss. Plant Mol Biol 92:313–336. https://doi.org/10.1007/s11103-016-0514-3
Lysak MA, Koch MA, Pecinka A, Schubert I (2005) Chromosome triplication found across the tribe Brassiceae. Genome Res 15:516–525. https://doi.org/10.1101/gr.3531105
Lysak MA, Cheung K, Kitschke M, Bures P (2007) Ancestral chromosomal blocks are triplicated in Brassiceae species with varying chromosome number and genome size. Plant Physiol 145:402–410. https://doi.org/10.1104/pp.107.104380
Mallory AC, Bartel DP, Bartel B (2005) MicroRNA-directed regulation of Arabidopsis Auxin Response Factor17 is essential for proper development and modulates expression of early auxin response genes. Plant Cell 17:1360–1375. https://doi.org/10.1105/tpc.105.031716
Massidon WP, Maddison DR (2018) Mesquite: a modular system for evolutionary analysis. Version 3.4. http://www.mesquiteproject.org
Masterson J (1994) Stomatal size in fossil plants: evidence for polyploidy in majority of angiosperms. Science 264:421–424. https://doi.org/10.1126/science.264.5157.421
Mauri M, Elli T, Caviglia G, et al (2017) RAWGraphs: a visualisation platform to create open outputs. In: Proceedings of the 12th biannual conference on Italian SIGCHI chapter
Mehraj H, Akter A, Miyaji N et al (2020) Genetics of clubroot and fusarium wilt disease resistance in brassica vegetables: the application of marker assisted breeding for disease resistance. Plants 9:1–15. https://doi.org/10.3390/plants9060726
Meng Y, Mao J, Tahir MM et al (2020) Mdm-miR160 participates in auxin-induced adventitious root formation of apple rootstock. Sci Hortic (amsterdam). https://doi.org/10.1016/j.scienta.2020.109442
Millar AA (2020) The function of miRNAs in plants. Plants 9:2–5. https://doi.org/10.3390/plants9020198
Murat F, Van De Peer Y, Salse J (2012) Decoding plant and animal genome plasticity from differential paleo-evolutionary patterns and processes. Genome Biol Evol 4:917–928. https://doi.org/10.1093/gbe/evs066
Navarro L, Jay F, Nomura K et al (2008) Suppression of the MicroRNA pathway by bacterial effector proteins. Science 321:964–967. https://doi.org/10.1126/science.1159505
Nikolov LA, Shushkov P, Nevado B et al (2019) Resolving the backbone of the Brassicaceae phylogeny for investigating trait diversity. New Phytol 222:1638–1651. https://doi.org/10.1111/nph.15732
Nozawa M, Miura S, Nei M (2012) Origins and evolution of microRNA genes in plant species. Genome Biol Evol 4:230–239. https://doi.org/10.1093/gbe/evs002
Oyston JW, Hughes M, Gerber S, Wills MA (2016) Why should we investigate the morphological disparity of plant clades? Ann Bot 117:859–879. https://doi.org/10.1093/aob/mcv135
Panchy N, Lehti-Shiu M, Shiu SH (2016) Evolution of gene duplication in plants. Plant Physiol 171:2294–2316. https://doi.org/10.1104/pp.16.00523
Parey E, Louis A, Cabau C et al (2020) Synteny-guided resolution of gene trees clarifies the functional impact of whole genome duplications. Mol Biol Evol. https://doi.org/10.1093/molbev/msaa149
Pinweha N, Asvarak T, Viboonjun U, Narangajavana J (2015) Involvement of miR160/miR393 and their targets in cassava responses to anthracnose disease. J Plant Physiol 174:26–35. https://doi.org/10.1016/j.jplph.2014.09.006
Prince VE, Pickett FB (2002) Splitting pairs: the diverging fates of duplicated genes. Nat Rev Genet 3:827–837
Rathore P, Geeta R, Das S (2016) Microsynteny and phylogenetic analysis of tandemly organised miRNA families across five members of Brassicaceae reveals complex retention and loss history. Plant Sci 247:35–48. https://doi.org/10.1016/j.plantsci.2016.03.002
Rojas AML, Drusin SI, Chorostecki U, et al (2020) Identification of key sequence features required for microRNA biogenesis in plants. Nat Commun 11:1–11. https://doi.org/10.1038/s41467-020-19129-6
Schnable JC, Springer NM, Freeling M (2011) Differentiation of the maize subgenomes by genome dominance and both ancient and ongoing gene loss. Proc Natl Acad Sci USA 108:4069–4074. https://doi.org/10.1073/pnas.1101368108
Schranz ME, Mitchell-Olds T (2006) Independent ancient polyploidy events in the sister families Brassicaceae and Cleomaceae. Plant Cell Online 18:1152–1165. https://doi.org/10.1105/tpc.106.041111
Shi T, Wang K, Yang P (2017) The evolution of plant microRNAs: insights from a basal eudicot sacred lotus. Plant J 89:442–457. https://doi.org/10.1111/tpj.13394
Shivaraj SM, Jain A, Singh A (2018) Highly preserved roles of Brassica MIR172 in polyploid Brassicas: ectopic expression of variants of Brassica MIR172 accelerates floral transition. Mol Genet Genomics 293:1121–1138. https://doi.org/10.1007/s00438-018-1444-3
Singh NK, Anand S, Jain A, Das S (2016) Comparative genomics and synteny analysis of KCS17-KCS18 cluster across different genomes and sub-genomes of Brassicaceae for analysis of its evolutionary history. Plant Mol Biol Rep 35:237–251. https://doi.org/10.1007/s11105-016-1019-6
Singh S, Das S, Geeta R (2018) A segmental duplication in the common ancestor of Brassicaceae is responsible for the origin of the paralogs KCS6 - KCS5, which are not shared with other angiosperms. Mol Phylogenet Evol 126:331–345. https://doi.org/10.1016/j.ympev.2018.04.018
Smith ML, Hahn MW (2021) New approaches for inferring phylogenies in the presence of paralogs. Trends Genet 37:174–187
Solovyev V, Kosarev P, Seledsov I, Vorobyev D (2006) Automatic annotation of eukaryotic genes, pseudogenes and promoters. Genome Biol 7(Suppl 1):1–12. https://doi.org/10.1186/gb-2006-7-s1-s10
Soltis DE, Albert VA, Leebens-Mack J et al (2009) Polyploidy and angiosperm diversification. Am J Bot 96:336–348. https://doi.org/10.3732/ajb.0800079
Sorin C, Bussell JD, Camus I et al (2005) Auxin and light control of adventitious rooting in Arabidopsis require Argonaute1. Plant Cell 17:1343–1359. https://doi.org/10.1105/tpc.105.031625
Sri T, Singh A (2015) Sequence and expression variation in SUPPRESSOR of OVEREXPRESSION of CONSTANS 1 (SOC1): homeolog evolution in Indian Brassicas. Dev Genes Evol 225:287–303. https://doi.org/10.1007/s00427-015-0513-4
Sun C, Wu J, Liang J et al (2015) Impacts of whole-genome triplication on miRNA evolution in Brassica rapa. Genome Biol Evol 7:3085–3096. https://doi.org/10.1093/gbe/evv206
Starega-Roslan J, Koscianska E, Kozlowski P, Krzyzosiak WJ (2011) The role of the precursor structure in the biogenesis of microRNA. Cell Mol Life Sci 68:2859–2871
Tank DC, Eastman JM, Pennell MW et al (2015) Nested radiations and the pulse of angiosperm diversification: increased diversification rates often follow whole genome duplications. New Phytol 207:454–467. https://doi.org/10.1111/nph.13491
Taylor RS, Tarver JE, Hiscock SJ, Donoghue PCJ (2014) Evolutionary history of plant microRNAs. Trends Plant Sci 19:175–182
Thomas BC, Pedersen B, Freeling M (2006) Following tetraploidy in an Arabidopsis ancestor, genes were removed preferentially from one homeolog leaving clusters enriched in dose-sensitive genes. Genome Res 16:934–946. https://doi.org/10.1101/gr.4708406
Turner M, Nizampatnam NR, Baron M et al (2013) Ectopic expression of miR160 results in auxin hypersensitivity, cytokinin hyposensitivity, and inhibition of symbiotic nodule development in soybean. Plant Physiol 162:2042–2055. https://doi.org/10.1104/pp.113.220699
VanBuren R, Man Wai C, Wang X et al (2020) Exceptional subgenome stability and functional divergence in the allotetraploid Ethiopian cereal teff. Nat Commun 11:1–11. https://doi.org/10.1038/s41467-020-14724-z
Wang J-W (2005) Control of root cap formation by microrna-targeted auxin response factors in Arabidopsis. Plant Cell Online 17:2204–2216. https://doi.org/10.1105/tpc.105.033076
Wang W, Luan Y (2015) The advance of tomato disease-related microRNAs. Plant Cell Rep 34:1089–1097
Wang X, Wang H, Wang J et al (2011) The genome of the mesopolyploid crop species Brassica rapa. Nat Genet 43:1035–1040. https://doi.org/10.1038/ng.919
Wang J, Qin J, Sun P et al (2019) Polyploidy index and its implications for the evolution of polyploids. Front Genet 10:807. https://doi.org/10.3389/fgene.2019.00807
Warwick SI, Francis A, Al-Shehbaz IA (2006) Brassicaceae: species checklist and database on CD-Rom. Plant Syst Evol 259:249–258. https://doi.org/10.1007/s00606-006-0422-0
Warwick SI, Sauder CA, Mayer MS, Al-Shehbaz IA (2009) Phylogenetic relationships in the tribes Schizopetaleae and Thelypodieae (Brassicaceae) based on nuclear ribosomal ITS region and plastid ndh F DNA sequences. Botany 87:961–985. https://doi.org/10.1139/B09-051
Warwick SI, Mummenhoff K, Sauder CA et al (2010) Closing the gaps: phylogenetic relationships in the Brassicaceae based on DNA sequence data of nuclear ribosomal ITS region. Plant Syst Evol 285:1–24. https://doi.org/10.1007/s00606-010-0271-8
Yadav D, Ahmed I, Kirti PB (2015) Genome-wide identification and expression profiling of annexins in Brassica rapa and their phylogenetic sequence comparison with B. juncea and A. thaliana annexins. Plant Gene 4:109–124. https://doi.org/10.1016/j.plgene.2015.10.001
Zhang J (2003) Evolution by gene duplication: an update. Trends Ecol Evol 18:292–298. https://doi.org/10.1016/S0169-5347(03)00033-8
Zhang B, Pan X, Cannon CH et al (2006) Conservation and divergence of plant microRNA genes. Plant J 46:243–259. https://doi.org/10.1111/j.1365-313X.2006.02697.x
Zhang K, Wang X, Cheng F (2019) Plant Polyploidy: origin, evolution, and its influence on crop domestication. Hortic Plant J 5:231–239. https://doi.org/10.1016/j.hpj.2019.11.003
Zheng Y, Jagadeeswaran G, Gowdu K et al (2013) Genome-Wide Analysis of MicroRNAs in Sacred Lotus, Nelumbo nucifera (Gaertn). Trop Plant Biol 6:117–130. https://doi.org/10.1007/s12042-013-9127-z
Acknowledgements
This work was supported by grants from Department of Biotechnology, Govt. of India (BT/PR10071/AGR/36/31/2007) and SERB-Department of Science and Technology, Govt. of India (EMR/2016/007813) received by AS. Fellowship to SS from Department of Science and Technology, Govt. of India, is gratefully acknowledged. Infrastructural support from TERI and TERI School of Advanced Studies is duly acknowledged.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Ethical approval
This article does not contain any studies with human participants or animals performed by any of the authors.
Additional information
Communicated by Stefan Hohmann.
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
438_2021_1797_MOESM1_ESM.tif
Supplementary file1 (TIF 29298 KB) Identification of 468 homologs of MIR160 from 31 families of plant kingdom. The numerical values signify number of MIR160 sequences identified within each family.
438_2021_1797_MOESM2_ESM.tif
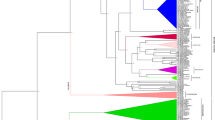
Supplementary file2 (TIF 34183 KB) Phylogenetic reconstruction of MIR160 homologs from Brassicaceae, Bryophytes and Gymnosperms. Relative to clades corresponding to MIR160b (green) and MIR160c (yellow), the clade specific to MIR160a (red) is evolutionarily closer to Bryophytes and Gymnosperms.
438_2021_1797_MOESM3_ESM.tif
Supplementary file3 (TIF 28914 KB) Phylogenetic reconstruction depicting evolutionary relationships between Brassicaceae MIR160 homologs. Clade specific to MIR160a (red) is distinct from clade containing MIR160b (green) & MIR160c (yellow). Within each clade, homologs of Brassica species derived from progenitor genomes AA, BB and CC group together in accordance with their sub-genome of origin (LF, MF1 and MF2).
438_2021_1797_MOESM4_ESM.tif
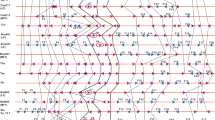
Supplementary file4 (TIF 99707 KB) Synteny analysis of 15 chromosomal segments (100 kb) containing MIR160a across 4 Brassica species including with A. thaliana as reference genome reveals gene preservation, duplication, and rearrangements. Synteny was established by mapping gene homologs of A. thaliana onto orthologous segments of Brassicaceae species. Position of MIR160a is depicted as a red triangle. Dashed connectors represent gene duplications within a genome/subgenome. Black rectangular boxes represent tandem duplicated blocks within each genome/subgenome.
438_2021_1797_MOESM5_ESM.tif
Supplementary file5 (TIF 70788 KB) Synteny analysis of 10 chromosomal segments containing MIR160b across 8 non-Brassica species with A. thaliana as reference reveals gene preservation, duplication, and rearrangements. Synteny was established by mapping gene homologs of A. thaliana onto orthologous segments (100 kb) of Brassicaceae species. Position of MIR160b is depicted as a red triangle. Dashed connectors represent gene duplications within a genome/subgenome. Black rectangular boxes represent tandem duplicated blocks within each genome/subgenome.
438_2021_1797_MOESM6_ESM.tif
Supplementary file6 (TIF 106077 KB) Synteny analysis of 16 chromosomal segments (100 kb) containing MIR160b across 4 Brassica species with A. thaliana as reference reveals gene preservation, duplication, and rearrangements. Position of MIR160b is depicted as a red triangle. Dashed connectors represent gene duplications within a genome / subgenome. Black rectangular boxes represent tandem duplicated blocks within each genome/subgenome.
438_2021_1797_MOESM7_ESM.tif
Supplementary file7 (TIF 77521 KB) Synteny analysis of 11chromosomal segments (100 kb) containing MIR160c across 8 non-Brassica species of Brassicaceae with A. thaliana as reference reveals gene preservation, duplication, and rearrangements. Position of MIR160c is depicted as a red triangle. Dashed connectors represent gene duplications within a genome/subgenome. Black rectangular boxes represent tandem duplicated blocks within each genome/subgenome.
438_2021_1797_MOESM8_ESM.tif
Supplementary file8 (TIF 99590 KB) Synteny analysis of 13 chromosomal segments (100 kb) containing MIR160c across 4 Brassica species of Brassicaceae with A. thaliana as reference reveals gene preservation, duplication, and rearrangements. Position of MIR160c is depicted as a red triangle. Dashed connectors represent gene duplications within a genome/subgenome. Black rectangular boxes represent tandem duplicated blocks within each genome/subgenome.
438_2021_1797_MOESM9_ESM.tif
Supplementary file9 (TIF 286293 KB) Comparative genome fractionation analysis across chromosomal segments (100 kb) harboring MIR160a, MIR160b and MIR160c derived from three subgenomes (LF, MF1 and MF2) of B. rapa. Comparative gene retention across subgenome specific chromosomal segments is interpreted using a hypothetical composite gene list as a reference. The reference is a non-redundant gene list comprising of both co-retained and unique genes present across the chromosomal segments derived from three subgenomes of B. rapa. The data is illustrated separately for chromosomal segments containing MIR160a (panel a), MIR160b (panel c) and MIR160c (panel e), respectively. The chromosomal segments are represented as rows of cells with cells depicting gene homologs. The extent of co-retention is depicted using mustard cells (gene retention across all the 3 subgenome), blue cells (gene retention across two subgenomes), black cells (gene homologs that are unique to a subgenome), and red cells (MIR160 homologs). Subgenome specific copy number of genes present across the three subgenomes of B. rapa is represented for each of the MIR160a (panel b), MIR160b (panel d), and MIR160c (panel f), respectively. The vertically arranged list of genes is organized based on copy number. Solid connectors join these to respective subgenome of origin shown in the right. The thickness of the connector depicts the copy number in a specific subgenome. The key for colored cells adjoining each of the listed gene correlates with the key proposed for panel a, c and e.
438_2021_1797_MOESM10_ESM.tif
Supplementary file10 (TIF 101383 KB) Multiple sequence alignment across the complete length of precursor sequences of 52 MIR160 homologs of Brassica species. The mature and star region mapping to locations of the least polymorphism in the precursor is indicated by miRNA and miRNA* respectively. SNP within miRNA and miRNA* is marked with the red arrow.
438_2021_1797_MOESM11_ESM.tif
Supplementary file11 (TIF 103378 KB) Diversity in predicted fold back structures of MIR160b precursor sequences from Brassica species (mFOLD server: RNA folding Form V2.3). The structures have been categorized into six groups based upon structural similarity.
438_2021_1797_MOESM12_ESM.tif
Supplementary file12 (TIF 96112 KB) Diversity in predicted fold back structures of MIR160c precursor sequences from Brassica species (mFOLD server: RNA folding Form V2.3). The structures have been categorized into four groups based upon structural similarity.
Rights and permissions
About this article
Cite this article
Singh, S., Singh, A. A prescient evolutionary model for genesis, duplication and differentiation of MIR160 homologs in Brassicaceae. Mol Genet Genomics 296, 985–1003 (2021). https://doi.org/10.1007/s00438-021-01797-8
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00438-021-01797-8