Abstract
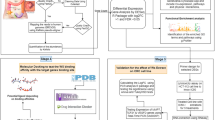
Numerous studies have demonstrated that lncRNAs participate in regulatory networks of different cancers. Dysregulation of various lncRNAs such as DUXAP8, LINC00963, and FOXD2-AS1 has been reported in the development of various cancers. The aim of this study was investigation of the importance and potential roles of DUXAP8, LINC00963, and FOXD2-AS1 in ER+ breast cancer (BC). We examined the expression levels of DUXAP8, LINC00963, and FOXD2-AS1 in 71 luminal A and B tumor tissues and two luminal A cell lines (MCF7 and T47D) compared with adjacent non-tumor tissues and MCF10A cell line by qRT-PCR assay, respectively. For identifying the relation between three lncRNAs and luminal BC, bioinformatic analyses were performed using some databases and software including GENEVESTIGATOR software, GEPIA2, DAVID, REVIGO, STRING, lncATLAS, Kaplan–Meier plotter, starBase, and miRNet tool. The results showed the significant upregulation of all three lncRNAs in luminal A and B tumor specimens and cell lines. Upregulation of DUXAP8 and FOXD2-AS1 was significantly associated with progesterone receptor-positive (PR+) and p53 protein expression in luminal BC patients, respectively. Based on bioinformatic analyses, DUXAP8 can be considered as a prognostic biomarker for patients with luminal BC. DUXAP8, LINC00963, and FOXD2-AS1 are involved in several cancer-associated signaling pathways and multiple cancer-related processes. In addition, bioinformatic analyses indicated that LINC00963/hsa-mir-130a-3p/HSPA8 axis might have potential regulatory role in BC. In conclusion, dysregulation of DUXAP8, LINC00963, and FOXD2-AS1 can play roles in the development of luminal BC. They may exert their functions through involvement in some cancer signaling pathways and processes. In addition, they may interact with miRNAs like predicted interaction of LINC00963 with miR-130a-3p.






Similar content being viewed by others
Data availability
The datasets supporting the conclusions of this article are available in: [GEPIA2 web server] at (http://www.gepia.cancer-pku.cn); [GENEVESTIGATOR software] at (https://genevestigator.com); [DAVID database] at (https://david.ncifcrf.gov); [REVIGO database] at (http://revigo.irb.hr); [Cytoscape software] at (http://www.cytoscape.org); [STRING database] at (https://string-db.org); [lncATLAS] at (https://lncatlas.crg.eu); [starBase database] at (http://starbase.sysu.edu.cn); [miRNet tool] at (http://www.mirnet.ca); [Kaplan–Meier plotter database] at (http://kmplot.com/analysis). Citation of all data is provided in the references list. The datasets supporting the conclusions of this article are also included within the article and its additional files.
Abbreviations
- BC:
-
Breast cancer
- BRCA:
-
Invasive breast carcinoma
- CeRNA:
-
Competing endogenous RNA
- DUXAP8:
-
Double homeobox A pseudogene 8
- ER:
-
Estrogen receptor
- FOXD2-AS1:
-
FOXD2 adjacent opposite strand RNA 1
- GO:
-
Gene ontology
- HER2:
-
Human epidermal growth factor receptor 2
- LncRNA:
-
Long noncoding RNAs
- LINC00963:
-
Long intergenic non-protein coding RNA 963
- PR:
-
Progesterone receptor
- BCRC-BB:
-
Breast Cancer Research Center Bio-bank
- BP:
-
Biological process
- CC:
-
Cellular component
- MF:
-
Molecular function
- qRT-PCR:
-
Real-time quantitative reverse transcription-polymerase chain reaction
References
Latgé G, Poulet C, Bours V, Josse C, Natural JG, Transcripts A. Molecular mechanisms and implications in breast cancers. Int J Mol Sci. 2018;9(1):123.
Dai X, Li T, Bai Z, Yang Y, Liu X, Zhan J, et al. Breast cancer intrinsic subtype classification, clinical use and future trends. Am J Cancer Res. 2015;5(10):2929–43.
Harbeck N, Gnant M. Lancet. Breast cancer. Lancet. 2017;389:1134–50.
Feng Y, Spezia M, Huang S, Yuan C, Zeng Z, Zhang L, et al. Breast cancer development and progression: risk factors, cancer stem cells, signaling pathways, genomics, and molecular pathogenesis. Genes Dis. 2018;5:77–106. https://doi.org/10.1016/j.gendis.2018.05.001.
Van Grembergen O, Bizet M, de Bony EJ, Calonne E, Putmans P, Brohee S, et al. Portraying breast cancers with long noncoding RNAs. Sci Adv. 2016;2:e1600220. https://doi.org/10.1126/sciadv.1600220.
Do H, Kim W. Roles of Oncogenic Long Non-coding RNAs in Cancer Development. (1598–866X (Print)).
Abolghasemi M, Tehrani SS, Yousefi T, Karimian A, Mahmoodpoor A, Ghamari A, et al. Critical roles of long noncoding RNAs in breast cancer. J Cell Physiol. 2020;235(6):5059–71. https://doi.org/10.1002/jcp.29442.
Youness RA, Gad MZ, Noncoding RNAR. Long non-coding RNAs: functional regulatory players in breast cancer. Non-coding RNA Res. 2019;4:36–44.
Sun W, Shi Y, Wang Z, Zhang J, Cai H, Zhang J, et al. Interaction of long-chain non-coding RNAs and important signaling pathways on human cancers (Review). Int J Oncol. 2018;53(6):2343–55. https://doi.org/10.3892/ijo.2018.4575.
Miao Y, Fan R, Chen L, Qian H. Clinical significance of long non-coding RNA MALAT1 expression in tissue and serum of breast cancer. Ann Clin Lab Sci. 2016;46:418–24.
Jiang B, Hailong S, Yuan J, Zhao H, Xia W, Zha Z, et al. Identification of oncogenic long noncoding RNA SNHG12 and DUXAP8 in human bladder cancer through a comprehensive profiling analysis. Biomed Pharmacother. 2018;108:500–7. https://doi.org/10.1016/j.biopha.2018.09.025.
Lin MG, Hong YK, Zhang Y, Lin BB, He XJ. Mechanism of lncRNA DUXAP8 in promoting proliferation of bladder cancer cells by regulating PTEN. Eur Rev Med Pharmacol Sci. 2018;22(11):3370–7. https://doi.org/10.26355/eurrev_201806_15158.
Xu LJ, Yu XJ, Wei B, Hui HX, Sun Y, Dai J, et al. Long non-coding RNA DUXAP8 regulates proliferation and invasion of esophageal squamous cell cancer. Eur Rev Med Pharmacol Sci. 2018;22(9):2646–52. https://doi.org/10.26355/eurrev_201805_14959.
Xu X, Xu Y, Shi C, Wang B, Yu X, Zou Y, et al. A genome-wide comprehensively analyses of long noncoding RNA profiling and metastasis associated lncRNAs in renal cell carcinoma. Oncotarget. 2017;8:87773–81. https://doi.org/10.18632/oncotarget.21206.
Chen J, Lou W, Ding B, Wang X. Aging. Overexpressed pseudogenes, DUXAP8 and DUXAP9, promote growth of renal cell carcinoma and serve as unfavorable prognostic biomarkers. Aging (Albany NY). 2019;11:5666–88.
Sun M, Nie FQ, Zang C, Wang Y, Hou J, Wei C, et al. The Pseudogene DUXAP8 promotes non-small-cell lung cancer cell proliferation and invasion by epigenetically silencing EGR1 and RHOB. Mol Ther. 2017;25:739–51.
Ma HW, Xie M, Sun M, Chen TY, Jin RR, Ma TS, et al. The pseudogene derived long noncoding RNA DUXAP8 promotes gastric cancer cell proliferation and migration via epigenetically silencing PLEKHO1 expression. Oncotarget. 2016;8:52211–24.
Yu T, Zhao Y, Hu Z, Li J, Chu D, Zhang J, et al. MetaLnc9 facilitates lung cancer metastasis via a PGK1-activated AKT/mTOR pathway. Can Res. 2017;77(21):5782–94. https://doi.org/10.1158/0008-5472.Can-17-0671.
Wu JH, Tian XY, An QM, Guan XY, Hao CY. LINC00963 promotes hepatocellular carcinoma progression by activating PI3K/AKT pathway. Eur Rev Med Pharmacol Sci. 2018;22(6):1645–52. https://doi.org/10.26355/eurrev_201803_14574.
Su F, He W, Chen C, Liu M, Liu H, Xue F, et al. The long non-coding RNA FOXD2-AS1 promotes bladder cancer progression and recurrence through a positive feedback loop with Akt and E2F1. Cell Death Dis. 2018;9:233. https://doi.org/10.1038/s41419-018-0275-9.
Chang Y, Zhang J, Zhou C, Qiu G, Wang G, Wang S, et al. Long non-coding RNA FOXD2-AS1 plays an oncogenic role in hepatocellular carcinoma by targeting miR206. Oncol Rep. 2018;40:3625–34.
Shen F, Chang H, Gao G, Zhang B, Li X, Jin BA-Ohoo, . Long noncoding RNA FOXD2-AS1 promotes glioma malignancy and tumorigenesis via targeting miR-185–5p/CCND2 axis. J Cell Biochem. 2019;120(6):9324–36.
Xu TP, Wang WY, Ma P, Shuai Y, Zhao K, Wang YF, et al. Upregulation of the long noncoding RNA FOXD2-AS1 promotes carcinogenesis by epigenetically silencing EphB3 through EZH2 and LSD1, and predicts poor prognosis in gastric cancer. Oncogene. 2018;37(36):5020–36.
Rong L, Zhao R, Lu J. Highly expressed long non-coding RNA FOXD2-AS1 promotes non-small cell lung cancer progression via Wnt/beta-catenin signaling. Biochem Biophys Res Commun. 2017;484:586–91.
Hu Q, Tai S, Wang J. Oncogenicity of lncRNA FOXD2-AS1 and its molecular mechanisms in human cancers. Pathol Res Pract. 2019;215(5):843–8.
Gao Y, Wang P, Wang Y, Ma X, Zhi H, Zhou D, et al. Lnc2Cancer v2.0: updated database of experimentally supported long non-coding RNAs in human cancers. Nucleic Acids Res. 2019;47(1):1028–33. https://doi.org/10.1093/nar/gky1096.
Majidzadeh AK, Kaviani A, Esmaeili R, Farahmand L, Shojamoradi MH, Zare AA, et al. Iranian breast cancer bio-bank: the activity and challenging issues. Cell Tissue Bank. 2013;14:11–20. https://doi.org/10.1007/s10561-012-9293-5.
Leroy B, Girard L, Hollestelle A, Minna JD, Gazdar AF, Soussi T. Analysis of TP53 mutation status in human cancer cell lines: a reassessment. Hum Mutat. 2014;35(6):756–65. https://doi.org/10.1002/humu.22556.
Livak KJ, Schmittgen TD. Analysis of relative gene expression data using real-time quantitative PCR and the 2(-Delta Delta C(T)) Method. Methods (San Diego, Calif). 2001;25(4):402–8. https://doi.org/10.1006/meth.2001.1262.
Tang Z, Kang B, Li C, Chen T, Zhang Z. GEPIA2: an enhanced web server for large-scale expression profiling and interactive analysis. Nucleic Acids Res. 2019;47:W556–60. https://doi.org/10.1093/nar/gkz430.
da Huang W, Sherman BT, Lempicki RA. Systematic and integrative analysis of large gene lists using DAVID bioinformatics resources. Nat Protoc. 2009;4:44–57. https://doi.org/10.1038/nprot.2008.211.
da Huang W, Sherman BT, Lempicki RA. Bioinformatics enrichment tools: paths toward the comprehensive functional analysis of large gene lists. Nucleic Acids Res. 2009;37:1–13. https://doi.org/10.1093/nar/gkn923.
Supek F, Bošnjak M, Škunca N, Šmuc T. REVIGO summarizes and visualizes long lists of gene ontology terms. PLoS One. 2011;6(7):e21800. https://doi.org/10.1371/journal.pone.0021800.
Szklarczyk D, Gable AL, Lyon D, Junge A, Wyder S, Huerta-Cepas J, et al. STRING v11: protein-protein association networks with increased coverage, supporting functional discovery in genome-wide experimental datasets. Nucleic Acids Res. 2019;47(D1):D607–13. https://doi.org/10.1093/nar/gky1131.
Chin CH, Chen SH, Wu HH, Ho CW, Ko MT, Lin CY. cytoHubba: identifying hub objects and sub-networks from complex interactome. BMC Syst Biol. 2014;8(Suppl 4):S11. https://doi.org/10.1186/1752-0509-8-s4-s11.
Mas-Ponte D, Carlevaro-Fita J, Palumbo E, Hermoso Pulido T, Guigo R, Johnson R. LncATLAS database for subcellular localization of long noncoding RNAs. RNA (New York, NY). 2017;23(7):1080–7. https://doi.org/10.1261/rna.060814.117.
Li JH, Liu S, Zhou H, Qu LH, Yang JH. starBase v2.0: decoding miRNA-ceRNA, miRNA-ncRNA and protein-RNA interaction networks from large-scale CLIP-Seq data. Nucleic Acids Res. 2014;42(Database issue):D92–7. https://doi.org/10.1093/nar/gkt1248.
Nagy Á, Lánczky A, Menyhárt O, Győrffy B. Validation of miRNA prognostic power in hepatocellular carcinoma using expression data of independent datasets. Sci Rep. 2018;8(1):9227. https://doi.org/10.1038/s41598-018-27521-y.
Chang L, Zhou G, Soufan O, Xia J. miRNet 2.0: network-based visual analytics for miRNA functional analysis and systems biology. Nucleic Acids Res. 2020;48(W1):W244–51. https://doi.org/10.1093/nar/gkaa467.
Nath A, Huang RS. Emerging role of long non-coding RNAs in cancer precision medicine. Mol Cell Oncol. 2020;7(1):1684130. https://doi.org/10.1080/23723556.2019.1684130.
Dong H, Cao W, Xue J. Long noncoding FOXD2-AS1 is activated by CREB1 and promotes cell proliferation and metastasis in glioma by sponging miR-185 through targeting AKT1. Biochem Biophys Res Commun. 2019;508:1074–81.
Yang X, Fau-Zhou Duan B, X, Zhou X, . Long non-coding RNA FOXD2-AS1 functions as a tumor promoter in colorectal cancer by regulating EMT and Notch signaling pathway. Euro Rev Med Pharmacol Sci. 2017;21:3586–91.
Milicevic Z, Bajic V, Zivkovic L, Kasapovic J, Andjelkovic U, Spremo-Potparevic B, et al. Identification of p53 and its isoforms in human breast carcinoma cells. Sci World J. 2014;2014:618–98.
Adler P, Kolde R, Kull M, Tkachenko A, Peterson H, Reimand J, et al. Mining for coexpression across hundreds of datasets using novel rank aggregation and visualization methods. Genome Biol. 2009;10:R139. https://doi.org/10.1186/gb-2009-10-12-r139.
Buj R, Aird KM. Deoxyribonucleotide triphosphate metabolism in cancer and metabolic disease. (1664–2392 (Print)).
Dowd J, Hendin J, Fukushiro-Lopes D, Laczynski D, Gentile S. Ion channels in breast cancer: from signaling to therapy. In: Breast cancer-from biology to medicine, Chap 13; 2017. pp 251–266.
Golubovskaya VM. FAK and Nanog cross talk with p53 in cancer stem cells. Anticancer Agents Med Chem. 2013;13(4):576–80. https://doi.org/10.2174/1871520611313040006.
Jeter CR, Yang T, Wang J, Chao HP, Tang DG. Concise review: NANOG in cancer stem cells and tumor development: an update and outstanding questions. Stem cells (Dayton, Ohio). 2015;33(8):2381–90. https://doi.org/10.1002/stem.2007.
Li H, Zhang Y, Ströse A, Tedesco D, Gurova K, Selivanova G. Integrated high-throughput analysis identifies Sp1 as a crucial determinant of p53-mediated apoptosis. Cell Death Differ. 2014;21(9):1493–502. https://doi.org/10.1038/cdd.2014.69.
Kong X, Zhang J, Li J, Shao J, Fang L. MiR-130a-3p inhibits migration and invasion by regulating RAB5B in human breast cancer stem cell-like cells. Biochem Biophys Res Commun. 2018;501(2):486–93. https://doi.org/10.1016/j.bbrc.2018.05.018.
Xian X, Tang L, Wu C, Huang L. miR-23b-3p and miR-130a-5p affect cell growth, migration and invasion by targeting CB1R via the Wnt/β-catenin signaling pathway in gastric carcinoma. Onco Targets Ther. 2018;11:7503–12. https://doi.org/10.2147/ott.s181706.
Boll K, Reiche K, Kasack K, Mörbt N, Kretzschmar AK, Tomm JM, et al. MiR-130a, miR-203 and miR-205 jointly repress key oncogenic pathways and are downregulated in prostate carcinoma. Oncogene. 2013;32(3):277–85. https://doi.org/10.1038/onc.2012.55.
Zhou YM, Liu J, Sun W. MiR-130a overcomes gefitinib resistance by targeting met in non-small cell lung cancer cell lines. Asian Pacific J Cancer Prev. 2014;15(3):1391–6. https://doi.org/10.7314/apjcp.2014.15.3.1391.
Shan N, Zhou W, Zhang S, Zhang Y. Identification of HSPA8 as a candidate biomarker for endometrial carcinoma by using iTRAQ-based proteomic analysis. Onco Targets Ther. 2016;9:2169–79. https://doi.org/10.2147/ott.s97983.
Sun G, Cao Y, Guo J, Li M, Dai Y. Heat shock cognate protein 70 enhanced integrin β1 mediated invasion in cancer cells. Cancer Manag Res. 2020;12:981–91. https://doi.org/10.2147/cmar.s235791.
Mayer MP, Bukau B. Hsp70 chaperones: cellular functions and molecular mechanism. Cell Mol Life Sci. 2005;62(6):670–84. https://doi.org/10.1007/s00018-004-4464-6.
Acknowledgements
We thank Dr. Rezvan Esmaeili (Head of Genetics Department, Breast Cancer Research Center of Motamed Cancer Institute, ACECR, Tehran, Iran) and Narges Jafarbeik Iravani for guidance of the experimental tests and other members of Genetics Department of Breast Cancer Research Center of Motamed Cancer Institute, ACECR, Tehran, Iran. In addition, the authors would like to thank Ms. Marzie Samimifar for proofreading the English language of the manuscript.
Funding
This work was supported by Faculty of Medicine, Tehran University of Medical Sciences (TUMS). The samples were obtained from Breast Cancer Research Center Bio-bank (BCRC-BB).
Author information
Authors and Affiliations
Contributions
M.A. performed experimental study and data analysis, bioinformatic analysis, literature review and wrote the manuscript. S.M. contributed to experimental study and data analysis and edited the manuscript. J.T.B. designed the research strategy, performed literature review, edited the manuscript, and reviewed the manuscript. M.M.N. performed statistical analysis and edited the manuscript. K.M. designed the research strategy, performed literature review, edited the manuscript and reviewed the manuscript. A.S. supervised the whole project and designed the research strategy, performed literature review, edited the manuscript and reviewed the manuscript. All the authors read and approved the final manuscript.
Corresponding authors
Ethics declarations
Conflict of interest
The authors declare that they have no conflicts of interest.
Ethical approval
The present study was approved by the Ethics Committee of Tehran University of Medical Sciences (TUMS) and written informed consent was obtained from all the participants (Code of Ethics: IR.TUMS.MEDICINE.REC.1398.659).
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
13577_2021_539_MOESM1_ESM.pdf
Supplementary file1 Fig. S1. The summarization of gene ontology (GO) terms related to the co-expressed genes with DUXAP8 retrieved from REVIGO. (PDF 357 KB)
13577_2021_539_MOESM2_ESM.pdf
Supplementary file2 Fig. S2. The summarization of gene ontology (GO) terms related to the co-expressed genes with LINC00963 retrieved from REVIGO. (PDF 318 KB)
13577_2021_539_MOESM3_ESM.pdf
Supplementary file3 Fig. S3. The summarization of gene ontology (GO) terms related to the co-expressed genes with FOXD2-AS1 retrieved from REVIGO. (PDF 391 KB)
13577_2021_539_MOESM4_ESM.xlsx
Supplementary file4 Data S1. The genes were co-expressed with DUXAP8, LINC00963, and FOXD2-AS1 across luminal subtypes of multiple BC datasets obtained from GENEVESTIGATOR. (XLSX 37 KB)
13577_2021_539_MOESM5_ESM.xlsx
Supplementary file5 Data S2. The potential binding miRNAs for DUXAP8, LINC00963, and FOXD2-AS1 retrieved from starbase database. (XLSX 17 KB)
13577_2021_539_MOESM7_ESM.xlsx
Supplementary file7 Data S4. 321 of 399 potential target genes have strong interactions with each other based on STRING database. (XLSX 32 KB)
Rights and permissions
About this article
Cite this article
Arabpour, M., Layeghi, S.M., Bazzaz, J.T. et al. The potential roles of lncRNAs DUXAP8, LINC00963, and FOXD2-AS1 in luminal breast cancer based on expression analysis and bioinformatic approaches. Human Cell 34, 1227–1243 (2021). https://doi.org/10.1007/s13577-021-00539-7
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s13577-021-00539-7