Abstract
We identified a Japanese patient with Lynch syndrome with a novel large germline deletion of chromosome 2p16-21, including the EPCAM, MSH2, and KCNK12 genes. The proband was a 46-year-old man with ascending colon cancer. The clinical significance of germline KCNK12 gene deletion, which encodes one of the subfamilies of two-pore-domain potassium channels, is still unknown.
Similar content being viewed by others
Lynch syndrome (LS) is an autosomal dominant inherited disorder caused by a germline pathogenic variant in mismatch repair (MMR) genes, including the MLH1, MSH2, MSH6, and PMS2 genes. Inactivation of the MMR gene, which is caused by both germline and/or somatic variants, leads to an increased frequency of replication errors in repeated sequences in the coding regions of cancer-related genes, resulting in the development of LS-associated tumors1. Therefore, LS is characterized by the development of various cancers at a young age, including colorectal cancer, endometrial cancer, small bowel cancer, and urinary tract cancer.
The EPCAM gene, which is located 17 kb upstream of MSH2, is not an MMR gene. However, germline deletion of the 3’ end of the EPCAM gene causes LS because EPCAM deletion leads to MSH2 gene silencing by promoter hypermethylation and MSH2 inactivation2. The proportion of EPCAM gene deletions in LS is estimated to be 1.0–3.0%3,4, and to date, only a few families have been reported in Japan5.
Here, we identified a Japanese patient with LS with a novel large germline deletion of chromosome 2p16-21, including the EPCAM, MSH2, KCNK12 genes. This novel finding provides evidence of germline deletion in this region and raises questions regarding the deletion of the KCNK12 gene.
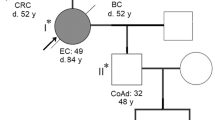
The proband was a 46-year-old man who was referred to our hospital with abdominal pain. A colonoscopy demonstrated an ascending colonic tumor, and biopsy revealed adenocarcinoma in the ascending colon. However, computed tomography showed no evidence of lymph node metastasis, distant metastasis, or other organ diseases. In addition, his mother had been treated for colon cancer, glioblastoma, gastric cancer, and endometrial cancer, and she died from glioblastoma at the age of 67 years old (Fig. 1).
The patient underwent laparoscopic right hemicolectomy, and histopathological examination revealed moderately differentiated adenocarcinoma with a mucinous component in the ascending colon tumor.
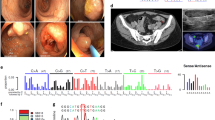
After obtaining the patient’s informed consent, we analyzed the patient’s tumor and found a high frequency of microsatellite instability. Then, after genetic counseling, we performed genetic analysis using next-generation sequencing6. The assay detected deletion of the EPCAM and MSH2 genes. We conducted further analysis using multiplex ligation-dependent probe amplification (MLPA) to confirm the deletion. Assays were performed to detect large genomic rearrangements of the EPCAM and MSH2 genes using an MLH1/MSH2 MLPA kit following the manufacturer’s protocol (SALSA MLPA KIT P-003 MLH1/MSH2, MRC-Holland, Amsterdam, The Netherlands). Then, we detected germline deletions of all MSH2 exons and exon 9 of the EPCAM gene. An additional assay was performed using an MSH6 MLPA kit (SALSA MLPA KIT P-072 MSH6), and we detected the deletion of the EPCAM gene and exon 2 of the KCNK12 gene, which was located upstream of the EPCAM gene (Fig. 2a, b).
We identified a novel germline deletion of chromosome 2p16-21, including the EPCAM, MSH2, and KCNK12 genes. Large genomic deletions and duplications, such as this one, have recently been identified using MLPA5. However, to our knowledge, there are no reports of germline deletions in this region.
The MSH2 gene is more likely to be found in genomic rearrangements at the locus of this disease because the MSH2 gene has a higher number and density of Alu repeats than other MMR genes7. At least 20% of germline MSH2 pathogenic variants are deletions of exons or multiple exons.
MSH2 variant carriers have a significantly higher risk of developing cancer of the urinary tract compared with carriers of MLH1 and MSH6 mutations8. Moreover, male MSH2 variant carriers have a high risk of developing cancer of the stomach9. However, our patient and his relatives had not developed urinary tract cancer or stomach cancer.
The difference in cancer risks between EPCAM deletion carriers and MSH2 variant carriers has been reported. The cumulative risk of endometrial cancer in EPCAM deletion carriers is lower than that in MSH2 variant carriers, while the cumulative risk of colorectal cancer in EPCAM deletion carriers is similar to that in MSH2 variant carriers10. On the other hand, the cumulative risk of colorectal cancer and endometrial cancer in carriers of a concurrent deletion of the EPCAM and MSH2 genes is reported to be similar to that in MSH2 variant carriers10. However, none of our patient’s relatives had developed endometrial cancer. In our study, to confirm the role of MMR proteins in carcinogenesis, immunohistochemical staining for MSH2 and EPCAM was conducted on colorectal cancer tissue, which showed only the loss of MSH2 expression (Fig. 3a, b). Thus, in our patient’s colorectal cancer, dysfunction of the MSH2 protein caused dysfunction of the MMR system, followed by a DNA replication error. Therefore, if the cancer risks were different between the MSH2 and EPCAM genes, the expression of MMR proteins should also be considered.
The KCNK12 gene located on 2p22-2p21 encodes one of the subfamilies of two-pore-domain potassium channels11. KCNK12 expression has been found in various organs, such as the pancreas, heart, skeletal muscle, ovary, testis, prostate, colon, peripheral blood leukocytes, small intestine, spleen, and thymus11. By setting and modulating the cellular membrane potential, potassium channels play a significant role in neuronal activity, muscular excitability, and hormone secretion12. Recent evidence supports the hypothesis that alterations in the expression and function of two-pore-domain potassium channels may contribute to cancer development and progression13, including breast cancer, colorectal cancer, leukemia, and lymphoma14,15,16. Further, a single-nucleotide polymorphism within the KCNK12 gene has been identified as a candidate genetic marker for migraine in the Finnish population17. However, our patient and his relatives did not have migraines or a neurovascular disorder.
The limitations of this report include the absence of segregation analysis and no identification of a breakpoint or rearranged genome. In conclusion, we identified a novel large germline deletion of chromosome 2p16-21, including the EPCAM, MSH2, and KCNK12 genes. However, the clinical significance of germline deletion of the KCNK12 gene is still unknown. Therefore, it is necessary to obtain and examine more data on this topic.
References
Yamaguchi, T. et al. Accumulation profile of frameshift mutations during development and progression of colorectal cancer from patients with hereditary nonpolyposis colorectal cancer. Dis. Colon Rectum 49, 399–406 (2006).
Ligtenberg, M. J. L., Kuiper, R. P., van Kessel, A. G. & Hoogerbrugge, N. EPCAM deletion carriers constitute a unique subgroup of Lynch syndrome patients. Fam. Cancer 12, 169–174 (2013).
Niessen, R. C. et al. Germline hypermethylation of MLH1 and EPCAM deletions are a frequent cause of Lynch syndrome. Genes Chromosomes Cancer 48, 737–744 (2009).
Kuiper, R. P. et al. Recurrence and variability of germline EPCAM deletions in Lynch syndrome. Hum. Mutat. 32, 407–414 (2011).
Eguchi, H. et al. Identification of a Japanese Lynch syndrome patient with large deletion in the 3’ region of the EPCAM gene. JJCO 46, 178–184 (2016).
Kohda, M. et al. Rapid detection of germline mutations for hereditary gastrointestinal polyposis/cancers using HaloPlex target enrichment and high-throughput sequencing technologies. Fam. Cancer 15, 553–562 (2016).
Van der Klift, H. et al. Molecular characterization of the spectrum of genomic deletions in the mismatch repair genes MSH2, MLH1, MSH6, and PMS2 responsible for hereditary nonpolyposis colorectal cancer (HNPCC). Genes Chromosomes Cancer 44, 123–138 (2005).
Joost, P., Therkildsen, C., Dominguez-Valentin, M., Jonsson, M. & Nilbert, M. Urinary tract cancer in Lynch syndrome; increased risk in carriers of MSH2 mutations. Urology 86, 1212–1217 (2015).
Capelle, L. G. et al. Risk and epidemiological time trends of gastric cancer in Lynch syndrome carriers in the Netherlands. Gastroenterology 138, 487–492 (2010).
Kempers, M. J. et al. Risk of colorectal and endometrial cancers in EPCAM deletion-positive Lynch syndrome: a cohort study. Lancet Oncol. 12, 49–55 (2011).
Girard, C. et al. Genomic and functional characteristics of novel human pancreatic 2P domain K+ channels. Biochem. Biophys. Res. Commun. 282, 249–256 (2001).
Jan, L. Y. & Jan, Y. N. Potassium channels and their evolving gates. Nature 371, 119–122 (1994).
Comes, N. et al. Involvement of potassium channels in the progression of cancer to a more malignant phenotype. Biochem. Biophys. Acta 1848, 2477–2492 (2015).
Keith, A. D., Zhang, W., Stayner, L. & Argos, M. Associations of two-pore domain potassium channels and triple negative breast cancer subtype in The Cancer Genome Atlas: systematic evaluation of gene expression and methylation. BMC Res. Notes 10, 475 (2017).
Kober, P., Bujko, M., Oledzki, J., Tysarowski, A. & Siedlechi, J. A. Methyl-CpG binding column-based identification of nine genes hypermethylated in colorectal cancer. Mol. Carcinog. 50, 846–856 (2011).
Gomes, A. Q. et al. Identification of a panel of ten cell surface protein antigens associated with immunotargeting of leukemias and lymphomas by peripheral blood gammadelta T cells. Haematologica 95, 1397–1404 (2010).
Nyholt, D. R. et al. A high-density association screen of 155 ion transport genes for involvement with common migraine. Hum. Mol. Genet. 17, 3318–3331 (2008).
HGV database
The relevant data from this Data Report are hosted at the Human Genome Variation Database at: https://doi.org/10.6084/m9.figshare.hgv.2990; https://doi.org/10.6084/m9.figshare.hgv.2993; https://doi.org/10.6084/m9.figshare.hgv.2996.
Author information
Authors and Affiliations
Contributions
All authors contributed to this work. S.N. prepared the description of the clinical course and wrote the manuscript; H.E., Y.O., and H.I. performed the genetic analysis; and T.Y. performed surgery, designed the study, and wrote the manuscript.
Corresponding author
Ethics declarations
Conflict of interest
The authors declare no competing interests.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Natsume, S., Yamaguchi, T., Eguchi, H. et al. Germline deletion of chromosome 2p16-21 associated with Lynch syndrome. Hum Genome Var 8, 19 (2021). https://doi.org/10.1038/s41439-021-00152-y
Received:
Revised:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41439-021-00152-y