Abstract
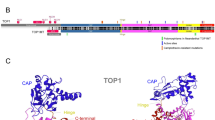
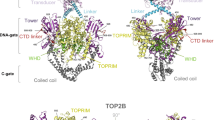
DNA topoisomerase III beta (TOP3B) is unique by operating on both DNA and RNA substrates to regulate gene expression and genomic stability. Mutations in human TOP3B are linked to neurodevelopmental and cognitive disorders, highlighting its relevance for human health. Despite the emerging importance of TOP3B, its precise cellular functions and evolutionary history remain poorly understood. Here, we show that TOP3B is conserved across main metazoan groups and evolved under strong purifying selection. Subdomain IV was identified as the most conserved TOP3B region, in agreement with its role in providing the structural foundation of the protein. On the contrary, subdomain II is the less conserved, possibly because it is the most structurally flexible region of all TOP3B regions. Interestingly, TOP3B residue at position 472, previously associated with schizophrenia, is highly variable across animals, suggesting a more specific role in humans and related species. Finally, we show that all TOP3B CXXC zinc finger motifs previously identified at the protein C-terminal region are retained across metazoans. We also found that the two major methylation sites known to regulate TOP3B activity are located in the most conserved region of the C-terminal arginine-glycine-glycine (RGG) box, suggesting that a similar regulatory mechanism may operate throughout animals. Overall, our results provide a better understanding of the evolution and functional roles of TOP3B.





Similar content being viewed by others
References
Abascal F, Zardoya R, Telford MJ (2010) TranslatorX: multiple alignment of nucleotide sequences guided by amino acid translations. Nucleic Acids Res 38:W7–W13
Aguinaldo AMA, Turbeville JM, Linford LS, Rivera MC, Garey JR, Raff RA, Lake JA (1997) Evidence for a clade of nematodes, arthropods and other moulting animals. Nature 387:489–493
Ahmad M, Shen W, Li W, Xue Y, Zou S, Xu D, Wang W (2017) Topoisomerase 3β is the major topoisomerase for mRNAs and linked to neurodevelopment and mental dysfunction. Nucleic Acids Res 45:2704–2713
Alemany S et al (2015) New suggestive genetic loci and biological pathways for attention function in adult attention-deficit/hyperactivity disorder. Am J Med Genet B Neuropsychiatr Genet 168:459–470
Amemiya CT et al (2013) The African coelacanth genome provides insights into tetrapod evolution. Nature 496:311–316
Berman HM et al (2000) The protein data bank. Nucleic Acids Res 28:235–242
Bizard AH, Hickson ID (2020) The many lives of type IA topoisomerases. J Biol Chem 295:7138–7153
Capranico G, Marinello J, Chillemi G (2017) Type I DNA topoisomerases. J Med Chem 60:2169–2192
Castresana J (2000) Selection of conserved blocks from multiple alignments for their use in phylogenetic analysis. Mol Biol Evol 17:540–552
Champoux JJ (2001) DNA topoisomerases: structure, function, and mechanism. Annu Rev Biochem 70:369–413
Collins AG, Cartwright P, McFadden CS, Schierwater B (2005) Phylogenetic context and basal metazoan model systems. Integr Comp Biol 45:585–594
Daghsni M et al (2018) TOP3B: a novel candidate gene in juvenile myoclonic epilepsy? Cytogenet Genome Res 154:1–5
Darriba D, Taboada GL, Doallo R, Posada D (2012) jModelTest 2: more models, new heuristics and parallel computing. Nat Methods 9:772–772
Del Amparo R, Branco C, Arenas J, Vicens A, Arenas M (2021) Analysis of selection in protein-coding sequences accounting for common biases. Brief Bioinform. https://doi.org/10.1093/bib/bbaa431
Duguet M, Serre M-C, de La Tour CB (2006) A universal type IA topoisomerase fold. J Mol Biol 359:805–812
Edgar RC (2004) MUSCLE: a multiple sequence alignment method with reduced time and space complexity. BMC Bioinform 5:113
Forterre P, Gribaldo S, Gadelle D, Serre M-C (2007) Origin and evolution of DNA topoisomerases. Biochimie 89:427–446
Frauer C et al (2011) Different binding properties and function of CXXC zinc finger domains in Dnmt1 and Tet1. PLoS ONE 6:e16627
Garnier F, Debat H, Nadal M (2018) Type IA DNA topoisomerases: a universal core and multiple activities. DNA Topoisomerases. Springer, New York, pp 1–20
Goto-Ito S, Yamagata A, Takahashi TS, Sato Y, Fukai S (2017) Structural basis of the interaction between Topoisomerase IIIβ and the TDRD3 auxiliary factor. Sci Rep 7:42123
Goulet I, Boisvenue S, Mokas S, Mazroui R, Côté J (2008) TDRD3, a novel Tudor domain-containing protein, localizes to cytoplasmic stress granules. Hum Mol Genet 17:3055–3074
Hansen G, Harrenga A, Wieland B, Schomburg D, Reinemer P (2006) Crystal structure of full length topoisomerase I from Thermotoga maritima. J Mol Biol 358:1328–1340
Huang L, Wang Z, Narayanan N, Yang Y (2018) Arginine methylation of the C-terminus RGG motif promotes TOP3B topoisomerase activity and stress granule localization. Nucleic Acids Res 46:3061–3074
Huelsenbeck JP, Ronquist F (2001) MRBAYES: Bayesian inference of phylogenetic trees. Bioinformatics 17:754–755
Hunt SE et al. (2018) Ensembl variation resources. Database. https://doi.org/10.1093/database/bay119
Jeffares DC, Tomiczek B, Sojo V, dos Reis M (2015) A beginners guide to estimating the non-synonymous to synonymous rate ratio of all protein-coding genes in a genome. Parasite genomics protocols. Springer, New york, pp 65–90
Kaufman CS, Genovese A, Butler MG (2016) Deletion of TOP3B is associated with cognitive impairment and facial dysmorphism. Cytogenet Genome Res 150:106–111
Kent WJ, Sugnet CW, Furey TS, Roskin KM, Pringle TH, Zahler AM, Haussler D (2002) The human genome browser at UCSC. Genome Res 12:996–1006
Kosakovsky Pond SL, Frost SD (2005) Not so different after all: a comparison of methods for detecting amino acid sites under selection. Mol Biol Evol 22:1208–1222
Kosakovsky Pond SL et al (2020) HyPhy 2.5—a customizable platform for evolutionary hypothesis testing using phylogenies. Mol Biol Evol 37:295–299
Kozlov AM, Darriba D, Flouri T, Morel B, Stamatakis A (2019) RAxML-NG: a fast, scalable and user-friendly tool for maximum likelihood phylogenetic inference. Bioinformatics 35:4453–4455
Kriventseva EV, Kuznetsov D, Tegenfeldt F, Manni M, Dias R, Simão FA, Zdobnov EM (2019) OrthoDB v10: sampling the diversity of animal, plant, fungal, protist, bacterial and viral genomes for evolutionary and functional annotations of orthologs. Nucleic Acids Res 47:D807–D811
Kwan KY, Wang JC (2001) Mice lacking DNA topoisomerase IIIβ develop to maturity but show a reduced mean lifespan. Proc Natl Acad Sci 98:5717–5721
Kwan KY, Greenwald RJ, Mohanty S, Sharpe AH, Shaw AC, Wang JC (2007) Development of autoimmunity in mice lacking DNA topoisomerase 3β. Proc Natl Acad Sci 104:9242–9247
Lee J-H, Voo KS, Skalnik DG (2001) Identification and characterization of the DNA binding domain of CpG-binding protein. J Biol Chem 276:44669–44676
Lima CD, Wang JC, Mondragón A (1994) Three-dimensional structure of the 67K N-terminal fragment of E. coli DNA topoisomerase I. Nature 367:138–146
Linder B et al (2008) Tdrd3 is a novel stress granule-associated protein interacting with the Fragile-X syndrome protein FMRP. Hum Mol Genet 17:3236–3246
Luo A et al (2010) Performance of criteria for selecting evolutionary models in phylogenetics: a comprehensive study based on simulated datasets. BMC Evol Biol 10:1–13
Miller MP, Kumar S (2001) Understanding human disease mutations through the use of interspecific genetic variation. Hum Mol Genet 10:2319–2328
Miller MA, Pfeiffer W, Schwartz T Creating the CIPRES Science Gateway for inference of large phylogenetic trees. In: 2010 gateway computing environments workshop (GCE), 2010. IEEE, pp 1–8
Mondragón A, DiGate R (1999) The structure of Escherichia coli DNA topoisomerase III. Structure 7:1373–1383
Nosenko T et al (2013) Deep metazoan phylogeny: when different genes tell different stories. Mol Phylogenet Evol 67:223–233
Pires R et al (2014) Screening of copy number variants in the 22q11. 2 region of congenital heart disease patients from the São Miguel Island, Azores, revealed the second patient with a triplication. BMC Genet 15:1–11
Rodríguez AC, Stock D (2002) Crystal structure of reverse gyrase: insights into the positive supercoiling of DNA. EMBO J 21:418–426
Ronquist F, Huelsenbeck JP (2003) MrBayes 3: Bayesian phylogenetic inference under mixed models. Bioinformatics 19:1572–1574
Rosato M et al (2019) Combined cellomics and proteomics analysis reveals shared neuronal morphology and molecular pathway phenotypes for multiple schizophrenia risk genes. Mol Psychiatr. https://doi.org/10.1038/s41380-019-0436-y
Sherry ST, Ward M-H, Kholodov M, Baker J, Phan L, Smigielski EM, Sirotkin K (2001) dbSNP: the NCBI database of genetic variation. Nucleic Acids Res 29:308–311
Srivastava M et al (2008) The Trichoplax genome and the nature of placozoans. Nature 454:955–960
Stoll G et al (2013) Deletion of TOP3β, a component of FMRP-containing mRNPs, contributes to neurodevelopmental disorders. Nat Neurosci 16:1228–1237
Talavera G, Castresana J (2007) Improvement of phylogenies after removing divergent and ambiguously aligned blocks from protein sequence alignments. Syst Biol 56:564–577
Tarsitano M et al (2014) Microduplications in 22q11. 2 and 8q22. 1 associated with mild mental retardation and generalized overgrowth. Gene 536:213–216
Thomson JP et al (2010) CpG islands influence chromatin structure via the CpG-binding protein Cfp1. Nature 464:1082–1086
Wang JC (1985) DNA topoisomerases. Annu Rev Biochem 54:665–697
Waterhouse A et al (2018) SWISS-MODEL: homology modelling of protein structures and complexes. Nucleic Acids Res 46:W296–W303
Wiegmann BM, Trautwein MD, Kim J-W, Cassel BK, Bertone MA, Winterton SL, Yeates DK (2009) Single-copy nuclear genes resolve the phylogeny of the holometabolous insects. BMC Biol 7:1–16
Wilson TM, Chen AD, Hsieh T-s (2000) Cloning and Characterization of DrosophilaTopoisomerase IIIβ relaxation of hypernegatively supercoiled DNA. J Biol Chem 275:1533–1540
Xia X (2018) DAMBE7: New and improved tools for data analysis in molecular biology and evolution. Mol Biol Evol 35:1550–1552
Xia X, Lemey P (2009) Assessing substitution saturation with DAMBE. In: Vandamme A-M, Salemi M, Lemey P (eds) The phylogenetic handbook: a practical approach to phylogenetic analysis and hypothesis testing, 2nd edn. Cambridge University Press, Cambridge, pp 615–630
Xia X, Xie Z, Salemi M, Chen L, Wang Y (2003) An index of substitution saturation and its application. Mol Phylogenet Evol 26:1–7
Xu B et al (2012) De novo gene mutations highlight patterns of genetic and neural complexity in schizophrenia. Nat Genet 44:1365–1369
Xu D et al (2013) Top3β is an RNA topoisomerase that works with fragile X syndrome protein to promote synapse formation. Nat Neurosci 16:1238–1247
Yang Y, Lu Y, Espejo A, Wu J, Xu W, Liang S, Bedford MT (2010) TDRD3 is an effector molecule for arginine-methylated histone marks. Mol Cell 40:1016–1023
Yang Y, McBride KM, Hensley S, Lu Y, Chedin F, Bedford MT (2014) Arginine methylation facilitates the recruitment of TOP3B to chromatin to prevent R loop accumulation. Mol Cell 53:484–497
Funding
This work was partially supported by a research grant to FM (SFRH/BD/131584/2017) from Fundação para a Ciência e a Tecnologia. MA was supported by a research grant (RYC-2015–18241) from the “Ministerio de Ciencia, Innovación y Universidades” of the Spanish Government.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflicts of interest
None to declare.
Additional information
Handling Editor: David Alvarez-Ponce.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Moreira, F., Arenas, M., Videira, A. et al. Molecular Evolution of DNA Topoisomerase III Beta (TOP3B) in Metazoa. J Mol Evol 89, 384–395 (2021). https://doi.org/10.1007/s00239-021-10011-7
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00239-021-10011-7