-
PDF
- Split View
-
Views
-
Cite
Cite
Sambuddha Sen, Zechariah Thompson, Christine Wachnowsky, Sean Cleary, Sophie R Harvey, J A Cowan, Biochemical impact of a disease-causing Ile67Asn substitution on BOLA3 protein, Metallomics, Volume 13, Issue 4, April 2021, mfab010, https://doi.org/10.1093/mtomcs/mfab010
Close - Share Icon Share
Abstract
Iron-sulfur (Fe-S) cluster biosynthesis involves the action of a variety of functionally distinct proteins, most of which are evolutionarily conserved. Mutations in these Fe-S scaffold and trafficking proteins can cause diseases such as multiple mitochondrial dysfunctions syndrome (MMDS), sideroblastic anemia, and mitochondrial encephalopathy. Herein, we investigate the effect of Ile67Asn substitution in the BOLA3 protein that results in the MMDS2 phenotype. Although the exact functional role of BOLA3 in Fe-S cluster biosynthesis is not known, the [2Fe-2S]-bridged complex of BOLA3 with GLRX5, another Fe-S protein, has been proposed as a viable intermediary cluster carrier to downstream targets. Our investigations reveal that the Ile67Asn substitution impairs the ability of BOLA3 to bind its physiological partner GLRX5, resulting in a failure to form the [2Fe-2S]-bridged complex. Although no drastic structural change in BOLA3 arises from the substitution, as evidenced by wild-type and mutant BOLA3 1H-15N HSQC and ion mobility native mass spectrometry experiments, this substitution appears to influence cluster reconstitution on downstream proteins leading to the disease phenotype. By contrast, substituted derivatives of the holo homodimeric form of BOLA3 are formed and remain active toward cluster exchange.
Ile67Asn substitution of human BOLA3 results in the disease phenotype of multiple mitochondrial dysfunctions syndrome-2. Substitution introduces no drastic structural change but impacts the ability of BOLA3 to interact with partner GLRX5 and impairs formation of a [2Fe-2S]-bridged complex of BOLA3 with GLRX5 and subsequent downstream trafficking. The homodimeric form of the BOLA3 derivative can be formed and is active in cluster exchange.
Significance to Metallomics
Herein, we investigate the impact of a disease-causing Ile67Asn substitution on the BOLA3 protein that has been implicated in iron-sulfur (Fe-S) cluster biogenesis and trafficking. Our study establishes that though this substitution introduces no drastic structural change in the protein, the ability of BOLA3 to interact with its partner GLRX5 is impacted. This in turn impairs formation of a [2Fe-2S]-bridged complex of BOLA3 with GLRX5, but not the BOLA3 holo homodimer. Our findings are suggestive of likely important physiological role of [2Fe-2S]-bridged BOLA3–GLRX5 heterocomplex in cluster trafficking.
Introduction
Mutations in several genes coding for proteins involved in iron-sulfur (Fe-S) cluster biogenesis and trafficking are known to result in multiple mitochondrial dysfunctions syndrome (MMDS). These proteins include NFU1 (MMDS1), BOLA3 (MMDS2), IBA57 (MMDS3), and ISCA2 (MMDS4).1 MMDS is characterized by metabolic acidosis with hyperglycemia, and a deficiency of respiratory complexes and lipoic acid-bound enzymes, such as pyruvate dehydrogenase and α-ketoglutarate dehydrogenase. Lipoate biogenesis is dependent on lipoate acid synthase, a [4Fe-4S]-containing mitochondrial protein, and so lipoate-dependent enzymes are affected. Two types of homozygous point mutation in the NFU1 gene, corresponding to p.Arg182Gln and p.Gly208Cys substitutions in the protein, are known to cause MMDS1. In the case of MMDS2, patients have been identified with mutations on the BOLA3 gene.2–4 In addition to truncation mutations leading to subsequent loss of the full-length protein,4 a missense mutation resulting in substitution of a highly conserved residue (p.Ile67Asn)3 has been reported.
The molecular basis for the phenotypes resulting from point substitutions is not well understood and is expected to vary with protein identity, as the proteins are structurally and functionally very different. Some of these residues lie far from their respective cluster binding site, while others are close, but the substitutions should most likely affect protein structure or function in some way. Prior studies from our laboratory with MMDS1 substitutions have demonstrated that the lethal Gly208Cys substitution in NFU1, near the Fe-S cluster binding site, severely decreases its ability to accept cluster from physiological sources, most likely due to a shift in monomer–dimer equilibrium.5 Another derivative of NFU1, Gly189Arg, is known to result in a milder phenotype of MMDS1 and also leads to a limited ability to accept cluster from physiologically relevant donors.6
In this paper, we focus on understanding the molecular basis of the MMDS2-causing Ile67Asn substitution in BOLA3. Although the exact role of BOLA3 in Fe-S cluster biogenesis is not clear, it has been implicated in mitochondrial Fe-S cluster biogenesis. BOLA3 has been shown to interact with the mitochondrial Fe-S cluster protein NFU1 that is involved in mitochondrial ISC assembly and has a strong interaction with GLRX5, a core component of the assembly machinery,7,8 forming a [2Fe-2S] cluster-bridged heterocomplex with GLRX5.9 GLRX5 is considered a core component of ISC machinery as it acts as an intermediary cluster carrier in its homodimeric form, taking up [2Fe-2S] cluster that is de novo synthesized on the scaffold protein ISCU, and then delivers it to downstream apo targets.10 Increasing evidences have suggested a close association between BOLA3 and GLRX5,9 and we have previously demonstrated that a cluster-bridged heterocomplex of BOLA3 and GLRX5 is capable of functioning in cluster trafficking with the [2Fe-2S] cluster constituted by cluster uptake from physiological donors such as ISCU. The cluster can also be readily transferred to downstream acceptors such as ferredoxin.8,11 Thus, the BOLA3–GLRX5 heterocomplex appears to be a viable candidate for Fe-S cluster trafficking. This role is in contrast with that predicted for the heterocomplex formed by GLRX5 with another related mitochondrial protein, BOLA1, as the BOLA1–GLRX5 heterocomplex is unsuitable for cluster exchange.12
Herein, we report a detailed structure–function study of the native and derivative proteins in order to understand the impact of the MMDS-causing Ile67Asn substitution on BOLA3. In addition to the disease-causing Ile67Asn mutation, we have used three other derivatives (Ile67Ala, Ile67Val, and Ile67Arg) for our study in order to better explore and define the structural and functional consequences of substitution of residue 67.
Experimental
Materials
The genes for human BOLA3 in a pET28b(+) vector (UniProt: Q53S33; residues 25–1077) and for human glutaredoxin 5 (GLRX5) located in a pET28b(+) vector between the NdeI and HindIII restriction sites, and lacking the respective mitochondrial targeting sequences (Δ1–24 for BOLA3 and Δ1–31 for GLRX5), were each purchased from GenScript.
Site-directed mutagenesis for plasmid construction
Site-directed mutagenesis was carried out using pairs of complementary primers for Ile67Asn, Ile67Val, Ile67Ala, and Ile67Arg mutations. Polymerase chain reaction (PCR) amplification was performed by use of Phusion polymerase from Fisher Scientific and pairs of complementary primers for each mutation. The protocol involved initial denaturation at 95°C for 1 min, followed by 25 cycles of initial denaturation at 95°C for 30 s, annealing at 55°C for 1 min, and 6 min of extension time at 72°C. Following amplification, Dpn1 was added to the PCR reaction mix and incubated at 37°C for 1 h to digest parental plasmid. The reaction mix was then transformed into BL21-DE3 cells by following a heat-shock method and plated on kanamycin plates. Positive clones were identified by extracting plasmid DNA from colonies using Qiagen's miniprep kit and confirmed by sequencing.
Protein expression and purification
BOLA3 and its derivatives were expressed and purified according to a previously published method.8 GLRX5, lacking the first 31 amino acids (Δ1–31) that correspond to the mitochondrial targeting sequence,13 was expressed and purified as reported previously.14 Human ISCU, human ferredoxin 1 (FDX1), and Thermotoga maritima (T. maritima NifS) were purified as previously reported.15–18 FDX1 was isolated and purified in the cluster-bound holo form and cluster was subsequently removed to prepare the apo form.19 For nuclear magnetic resonance (NMR) experiments, BOLA3 mutants were expressed in Escherichia coli BL21 cells grown in M9 minimal media at 37°C until the optical density reached 0.7. Subsequently, the cultures were induced for 4 h with 100 mg L−1 IPTG, the cells were harvested, and cell pellets resuspended in 50 mM HEPES, 100 mM NaCl, pH 7.5 buffer and lysed by sonication. The cell debris was removed by centrifugation and the supernatant passed over cobalt resin. The resin was washed with the same buffer supplemented with 5 mM imidazole before the protein was eluted with 150 mM imidazole in 50 mM HEPES, 100 mM NaCl, pH 7.5 buffer. Total protein concentration was determined by the use of the Bradford assay.
Nuclear magnetic resonance
Purified BOLA3 wild type and derivatives were buffer exchanged into 50 mM phosphate buffer with 5 mM DTT, pH 7.5, and 10% D2O. Spectra were recorded at 277 K on a Bruker Ascend 700 MHz instrument, equipped with a TXO cryoprobe, Avance III HD console, and a SampleJet autosampler. A standard water suppression [15N–1H] heteronuclear single-quantum coherence (HSQC) pulse sequence was used for data collection. All data were processed with TopSpin, version 3.6, and data were visualized using NMRFAM-SPARKY version 1.414.20
Native ion mobility mass spectrometry
Both WT and Ile67N mutant proteins were purified as described above. Proteins were then thrombin cleaved with Thrombin CleanCleave™ Kit from Sigma Aldrich for 4 h at room temperature.
All samples were buffer exchanged using a Bio-Spin 6 column (Bio-Rad) into 200 mM ammonium acetate prior to mass spectrometry analysis. Mass spectrometry analysis was performed with a Synapt G2 ion mobility mass spectrometer (Waters Corp.) using a static nanoelectrospray ionization (nanoESI) source. NanoESI emitters were prepared by pulling borosilicate capillaries (i.d. 0.78 mm Sutter Instruments) to a tip i.d. of ∼1 µm using a Flaming-Brown P-97 micropipette puller (Sutter Instruments). For each sample, ∼3–5 µl of solution was loaded into an emitter, which was placed approximately 1–2 mm from the entrance of the mass spectrometer. A platinum wire inserted into the solution was used to apply an electrical potential of 0.6–0.8 kV relative to instrumental ground to initiate electrospray. The ion source was held at a temperature of 30°C for all experiments. Mass spectra were collected in sensitivity mode for maximal signal-to-noise. Trap gas was introduced at a flow rate of 4 ml/min in all experiments. Trap/transfer collision energy settings were 4/2 V, respectively. For all analytes, mass spectra were collected continuously for ∼5 min and summed. Ion mobility settings were as follows: helium cell gas flow 100.0 ml/min, ion mobility spectrometry (IMS) gas flow 40.0 ml/min, IMS wave height 15 V, and IMS wave velocity 400 m/s. All mass spectral data were processed using MassLynx v. 4.2 (Waters Corp.).
In vitro reconstitution of apo proteins
All in vitro reconstitutions were performed in 50 mM HEPES, 100 mM NaCl, pH 7.5 buffer. Reconstitutions for the BOLA3–GLRX5 heterodimers were conducted with different BOLA3 derivatives as previously described,8,9 using a 1 : 1 molar ratio (350 µM of each protein) in the presence of 2 µM T. maritima NifS, 5 mM DTT, and 3 mM glutathione (GSH). The reaction mixture was argon purged for 30 min, and then made up to 0.6 mM in FeCl3 and 0.6 mM in l-cysteine. Following incubation for 1.5 h, the reaction mixture was passed through a PD-10 desalting column to remove excess reactants. The effective holo concentration of [2Fe-2S] BOLA3–GLRX5 protein was estimated by the use of the extinction coefficient, ε393 = 8.3 mM−1 cm−1, as reported in literature.9,21
Reconstitution of BOLA3 derivatives (Ile67Asn, Ile67Ala, Ile67Val, and Ile67Arg) (to form holo BOLA3) was performed in a reconstitution mixture containing 200 µM BOLA3, 5 mM DTT, 2 mM T. maritima NifS, 0.6 mM FeCl3, and 0.6 mM cysteine. After incubating for 45 min, the holo protein was isolated following passage through a PD-10 column. For ISCU reconstitution, 200 µM apo protein was incubated with 50 mM DTT and 8 M urea, argon purged for 30 min, and then made up to 1 mM in FeCl3 and 1 mM in Na2S. After incubation for 1 h, the reaction mixture was concentrated and passed through a PD-10 desalting column, equilibrated with 50 mM HEPES, 100 mM NaCl, and pH 7.5, to remove excess reagents.
Circular dichroism spectra of reconstituted BOLA3–GLRX5
Circular dichroism (CD) spectra of the reconstituted holo BOLA3–GLRX5 heterodimeric proteins formed from different BOLA3 derivatives were measured from 300 to 600 nm at a 200 nm/min scan rate by use of a Jasco J-815 circular dichroism spectrometer. Raw spectra were deconvoluted by use of the deconvolution function from the Spectra Manager II analysis software, which used overlapping Lorentzian curves having the same full width at the half-maximum value to accurately distinguish the peak positions for each band using Fourier self-deconvolution and finite impulse response operator methods.22,23
Cluster transfer experiments analyzed by CD spectroscopy
UV–vis CD spectroscopy was used to investigate cluster transfer from holo Fe-S cluster proteins to apo proteins using 1-cm quartz cuvettes. CD scans from 600 to 300 nm were collected under anaerobic conditions at a scan rate of 200 nm/min with 2 min interval between accumulations. A mixture of apo protein(s) in 5 mM DTT and 3 mM GSH in 50 mM HEPES, 100 mM NaCl, pH 7.5 was degassed, transferred to an anaerobic cuvette via a gas tight syringe, and reaction initiated via the addition of degassed holo protein. The concentration of [2Fe-2S] cluster for each holo protein was determined via absorbance measurements. Data were processed by use of JASCO Spectra Manager II analysis software and the deconvolution function from Spectra Manager II analysis software was used for analysis of bands in the spectra. The second-order rate constants were calculated using the program DynaFit by BioKin24 by a method similar to that developed by Johnson and colleagues.25,26 Proteins were mixed in a 1 : 1 ratio (30 µM each) and the characteristic peaks for respective holo acceptor protein (445 nm for FDX1, and 350 nm for BOLA3–GLRX5 heterocomplex) were monitored by CD. A representative figure showing the CD spectra of different [2Fe-2S]-containing species involved in the cluster transfer experiments is provided in Fig. S2. The data were converted to % cluster transfer, where the percentage of cluster transferred was determined by comparing signal intensity of holo protein after cluster transfer to that of control spectra of the holo protein with same cluster concentration upon reconstitution.
Isothermal titration calorimetry for interaction between apo BOLA3 mutants and apo GLRX5
Apo BOLA3 derivative (Ile67Asn, Ile67Ala, Ile67Val, or Ile67Arg) and GLRX5 proteins were dialyzed overnight in 50 mM HEPES, 100 mM NaCl, 5 mM GSH, pH 7.5, buffer at 4°C. GSH was added as it has been found to mediate interaction between BOLA3 and GLRX5.7 Isothermal titration calorimetry (ITC) experiments at 30°C were carried out on a MicroCal calorimeter. The sample cell contained 50 µM apo GLRX5 in abovementioned buffer used for dialysis, and the syringe contained 1 mM BOLA3 mutant in the same buffer. One millimolar of each BOLA3 mutant was titrated in 10 µl aliquots with 24 s durations and intervals of 6 min. Control titrations of buffer into protein and protein into buffer were also performed and the data subtracted from the raw experimental output. The data was analyzed using Origin software provided by MicroCal.
Results
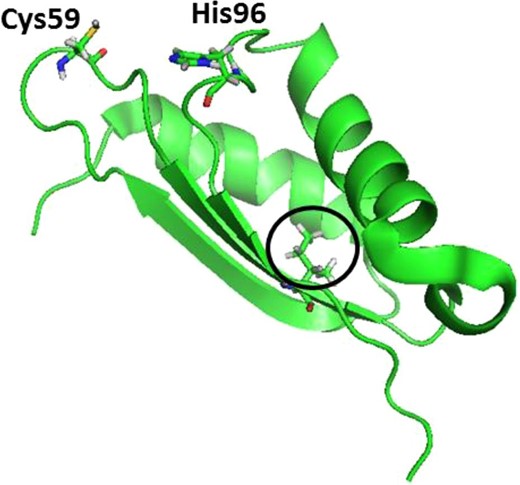
All members of the BOLA family of proteins adopt a α1β1β2α2α3β3 topology,7,27–29 with β1 and β2 antiparallel to each other and β3 parallel to β2. According to the solution structure of human BOLA3 (PDB:2NCL), the Ile67 residue lies on the β2-strand, with the hydrophobic side chain pointing to the inner core of the protein (Fig. 1), and so an outcome of the Ile67Asn substitution could be a change in structural folding of BOLA3.
Solution structure of apo BOLA3 (PDB-2NCL) with Ile67 circled. His96 and Cys59 are cluster-coordinating residues.
Ile67Asn substitution results in structural perturbations but retains overall tertiary fold
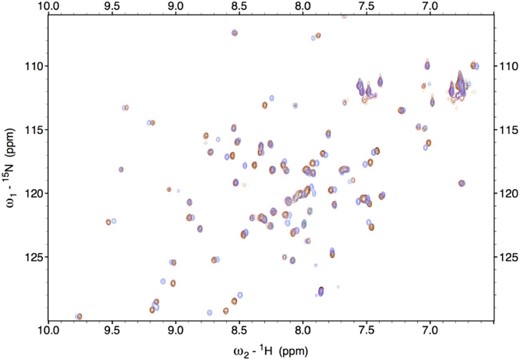
As the Ile67Asn substitution results in the MMDS2 phenotype, we were interested in exploring the structural aspects of the substituted derivative with NMR spectroscopy. 15N-labeled wild type and BOLA3 derivatives were expressed in minimal media and a series of 15N-1H-HSQC experiments were performed to examine each protein's fingerprint. This experiment can assist in determining whether or not the protein is well folded, in addition to providing a general assessment of secondary structure.30–32 If drastic changes are observed across the BOLA3 derivatives, this can be an indication that the overall protein fold is different, or whether there are structural perturbations within the protein. If the protein is unfolded, we would expect to observe the H-N chemical shifts to overlap and appear crowded between 8 and 8.5 ppm. Overall, there appears to be noticeable shifts within the spectra when comparing wild type to substituted BOLA3 protein (Fig. 2). Since the substitution is within a β-strand, we would expect that the structure for each would vary to some degree; however, this mutation does not appear to be deleterious to the overall tertiary structure as evidenced by the well-resolved and well-dispersed peaks obtained for wild-type and mutant spectra.
Overlay of 15N HSQC spectra of native (red), Ile67Asn (black), Ile67Ala (magenta), Ile67Val (purple), and Ile67Arg (orange) derivatives of BOLA3.
Ile67 substitution affects binding with apo GLRX5
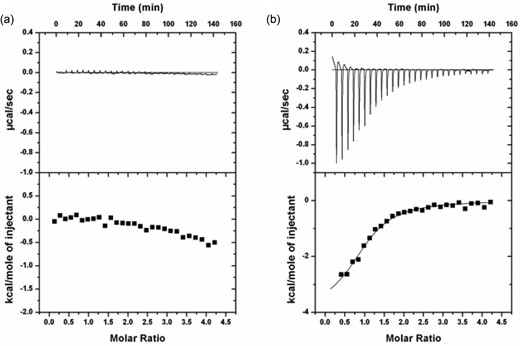
Due to the shifts observed within the protein fingerprint region for BOLA3, we examined the potential interaction between apo BOLA3 derivatives (Ile67Asn, Ile67Ala, Ile67Val, or Ile67Arg) and apo GLRX5 via ITC experiments. These revealed that only the Ile67Val derivative was able to exhibit significant binding to GLRX5 (Fig. 3b), with binding parameters very similar to those obtained with native BOLA3,8,9 and consistent with Ile67Val maintaining a native-like structure. Three other derivatives, Ile67Asn, Ile67Ala, and Ile67Arg, however, did not exhibit any meaningful binding with GLRX5 (Fig. 3a).
ITC plots for (a) Apo Ile67Asn BOLA3 titrated to apo GLRX5 that show no meaningful binding and (b) Apo Ile67Val BOLA3 titrated to apo GLRX5 (N = 1.16 ± 0.03, K = 1.5 × 105 ± 2 × 104 M−1, ΔH = −3294 ± 121 J mol−1, ΔS = 13 J mol−1 K−1). The experiments were performed in the presence of 5 mM GSH in 50 mM HEPES, 100 mM NaCl, pH 7.5. One millimolar BOLA3 was titrated into 50 µM GLRX5 in 10 µl aliquots over a period of 24 s with a 5 min interval at 30°C.
Ile67 mutation results in similar conformation and collisional cross section
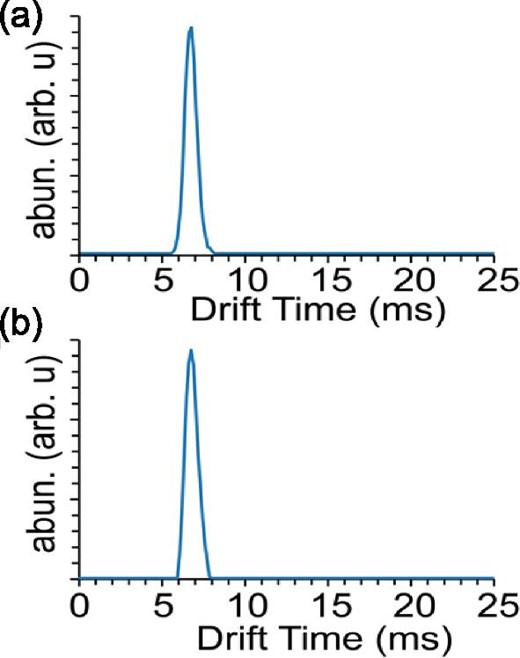
Since the Ile67Asn mutation diminishes the binding interaction between BOLA3 and GLRX5, but does not show any significant changes in the overall tertiary structure or fold of the protein, we were interested in exploring this phenomenon via alternative methods. Ion mobility native mass spectrometry (IM-MS) allows one to separate ions based upon their charge and shape and can yield insight into any structural or conformational changes that would potentially be induced as a result of the mutation.33,34 Prepared samples were ionized and data analyzed to extract the drift times of WT and Ile67Asn BOLA3 proteins. Analysis of the drift times for both proteins revealed that they show virtually identical values of 6.74 ± 0.37 and 6.75 ± 0.36 ms for both WT and Ile67Asn, respectively (Fig. 4). The similarity in drift times between the two proteins shows that they are conformationally similar, and that there are no large-scale structural differences between the wild type and Ile67Asn mutant, which further supports our NMR HSQC analysis between the two proteins.
Ion mobility chromatograms of the 5+ charge state of BOLA3 WT (a) and the 167N mutant (b). Chromatograms demonstrate that no structural change is seen between wild-type and mutant species.
Ile67 substitution affects reconstitution of BOLA3–GLRX5 heterodimer
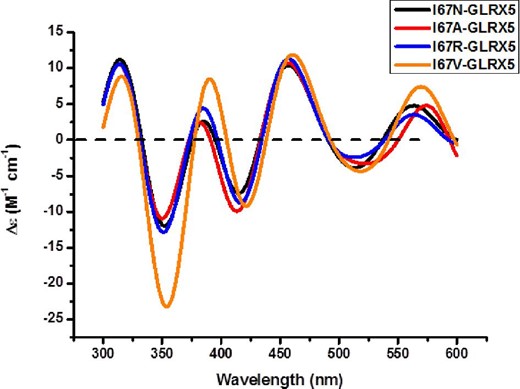
Attempts to reconstitute the BOLA3–GLRX5 heterodimer by standard Fe-S cluster reconstitution procedures, using a 1 : 1 mixture of Ile67Asn BOLA3 and GLRX5, resulted in reconstitution of the GLRX5 homodimer, instead of the anticipated BOLA3–GLRX5 heterodimer, as evidenced by CD spectra (Fig. 5). While the CD spectrum of a [2Fe-2S]-bound BOLA3–GLRX5 heterodimer has a pronounced negative peak around 350 nm,8,9 the reconstituted protein obtained using Ile67Asn, Ile67Ala, and Ile67Arg derivatives of BOLA3 lacks this feature. Instead, the CD signature resembles that of [2Fe-2S]-bound GLRX5 homodimer, with a less pronounced negative feature at 350 nm.7,8 Ile67Val, however, is able to form [2Fe-2S]-bridged heterodimer with GLRX5, as the CD signature resembles that obtained with the native BOLA3.8,9
Comparison of the CD spectra of reconstituted holo GLRX5-BOLA3 derivatives. The extinction coefficients are based on the cluster concentrations.
Ile67 mutation does not impact cluster transfer ability of holo BOLA3
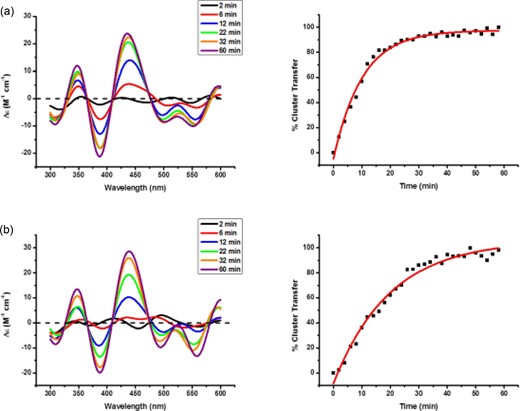
We have previously demonstrated that native BOLA3 can be reconstituted to the homodimeric form (without GLRX5 present as a heteromeric partner) by standard reconstitution procedures. The holo BOLA3 thus obtained can transfer its cluster to standard [2Fe-2S] acceptors such as FDX1 and FDX2.35 We observe that similar to the native BOLA3, all four derivatives, Ile67Asn, Ile67Ala, Ile67Val, and Ile67Arg, can be reconstituted and the holo protein thus obtained readily transfers its cluster to FDX1 at comparable rates (Fig. 6; Table 1). Thus, Ile67 substitution does not impact the cluster-binding and transfer ability of the homodimer BOLA3.
Time course measurements for cluster transfer from (a) holo Ile67Asn to apo FDX1 and (b) holo Ile67Val to apo FDX1 under anaerobic conditions. The CD signal intensity was monitored at room temperature for a 1 : 1 donor:acceptor cluster stoichiometry in the presence of 3 mM GSH. CD spectra in the 300–600 nm region were recorded every 2 min.
Second-order rate constants for the cluster transfer reactions from holo BOLA3 to apo FDX1.
| Cluster transfer . | Second-order rate constant (M−1 min−1) . |
|---|---|
| Native BOLA3 to FDX1 | 4000 ± 330 |
| Ile67Asn to FDX1 | 4200 ± 400 |
| Ile67Ala to FDX1 | 3700 ± 370 |
| Ile67Arg to FDX1 | 4600 ± 420 |
| Ile67Val to FDX1 | 2800 ± 310 |
| Cluster transfer . | Second-order rate constant (M−1 min−1) . |
|---|---|
| Native BOLA3 to FDX1 | 4000 ± 330 |
| Ile67Asn to FDX1 | 4200 ± 400 |
| Ile67Ala to FDX1 | 3700 ± 370 |
| Ile67Arg to FDX1 | 4600 ± 420 |
| Ile67Val to FDX1 | 2800 ± 310 |
Second-order rate constants for the cluster transfer reactions from holo BOLA3 to apo FDX1.
| Cluster transfer . | Second-order rate constant (M−1 min−1) . |
|---|---|
| Native BOLA3 to FDX1 | 4000 ± 330 |
| Ile67Asn to FDX1 | 4200 ± 400 |
| Ile67Ala to FDX1 | 3700 ± 370 |
| Ile67Arg to FDX1 | 4600 ± 420 |
| Ile67Val to FDX1 | 2800 ± 310 |
| Cluster transfer . | Second-order rate constant (M−1 min−1) . |
|---|---|
| Native BOLA3 to FDX1 | 4000 ± 330 |
| Ile67Asn to FDX1 | 4200 ± 400 |
| Ile67Ala to FDX1 | 3700 ± 370 |
| Ile67Arg to FDX1 | 4600 ± 420 |
| Ile67Val to FDX1 | 2800 ± 310 |
Ile67 mutation hinders formation of [2Fe-2S] BOLA3–GLRX5 heterocomplex by cluster uptake from ISCU
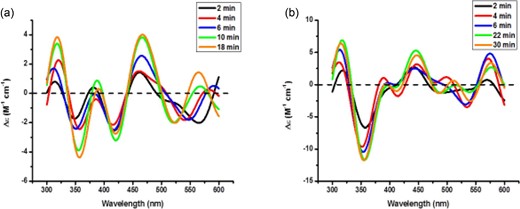
We have previously demonstrated that a 1 : 1 mixture of native BOLA3 and GLRX5 takes up [2Fe-2S] cluster from a cluster donor such as ISCU, thereby preferentially forming the holo BOLA3–GLRX5 heterodimer instead of a possible GLRX5 homodimer.8 Among the BOLA3 derivatives, only Ile67Val and GLRX5 are able to take up cluster and form the [2Fe-2S]-bridged BOLA3–GLRX5 heterodimer similar to native BOLA3. The disease-causing mutant, Ile67Asn, along with two others, Ile67Ala and Ile67Arg, is unable to form this heterodimer, as addition of holo ISCU to a 1 : 1 mixture of any one of these BOLA3 mutants and GLRX5 results in formation of holo homodimeric GLRX5, as indicated by the characteristic CD signature of the cluster-bridged holo protein (Fig. 7).
Time course measurements for the cluster transfer experiment from holo ISCU to (a) apo Ile67Asn BOLA3–GLRX5 and (b) apo Ile67Val BOLA3–GLRX5 under anaerobic conditions. The CD response was monitored at room temperature for a 1 : 1 donor:acceptor cluster stoichiometry in the presence of 3 mM GSH. CD spectra were collected in the 300–600 nm range every 2 min.
Discussion
Four different genetic alterations have been identified in BOLA3 from patients with MMDS. In addition to alterations that result in loss of full-length protein,2,4 two homozygous missense mutations have been described that result in substitution of single amino acid Ile67Asn or His96Arg in BOLA3 protein.3,36
The exact function of BOLA3 in Fe-S biosynthesis pathway is not known; however, it is believed to be involved in a late step of cluster biogenesis.4,37 It has been proposed that BOLA3 assists another protein, NFU1, to facilitate [4Fe-4S] cluster transfer to certain mitochondrial proteins.37 Another possibility is that the [2Fe-2S]-bridged heterocomplex formed by BOLA3 and GLRX5 functions as an intermediary cluster carrier, trafficking the cluster to downstream apo acceptor proteins.7,8 Biochemical investigation of proteins with the disease-causing mutation can further our understanding of BOLA3 and may help in deciphering the cellular function of the protein. With point mutations in BOLA3 causing a severe MMDS phenotype, with deficiency in essential Fe-S cluster-containing enzymes, we expected that our biochemical study would reveal a compromised structure or function of BOLA3 arising from the single amino acid substitution. The disease-causing His96Arg substitution involves the key His96 residue that is involved in [2Fe-2S] binding in the [2Fe-2S]-bridged BOLA3–GLRX5 heterocomplex,9 which would obviously result in loss of cluster-binding ability. However, the molecular basis of disease resulting from the other substitution (Ile67Asn), involving a residue situated far away from the cluster binding site, is difficult to predict. We decided to investigate the molecular basis of the disease condition by using three different derivatives, Ile67Ala, Ile67Val, and Ile67Arg, in addition to the disease-causing Ile67Asn.
Ile67 residue lies on a β-sheet, with the hydrophobic side chain pointing to the inner core of the protein, and so a possible outcome of the Ile67Asn substitution could be a drastic change in BOLA3 structure. Our 1H-15N HSQC experiments demonstrated that there are no drastic structural perturbations in any of the Ile67Asn, Ile67Ala, and Ile67Arg derivatives. The protein retains its overall fold based on the well-resolved and well-dispersed peaks observed. Most likely, the substitution results in changes within the β-sheet structure but does not result in deleterious protein unfolding. It is to be noted that the NMR experiments have been performed with apo proteins and there could be changes upon [2Fe-2S] binding and formation of heterocomplex. However, performing NMR experiments with holo proteins can often be challenging because of paramagnetic line broadening and hyperfine shifts associated with the cluster that limited our ability to perform NMR experiments with holo proteins. IM-MS experiments further confirm our NMR results as both WT and Ile67Asn BOLA3 have similar drift times that correlate to similar structure/conformation between the two proteins.
While no significant structural change is observed in the apo BOLA3 proteins, we observe significant functional change due to the substitution of Ile67. Prior studies have demonstrated that apo BOLA3 and apo GLRX5 show relatively strong binding even in the absence of the bridging cluster7,8 with a well-defined interaction surface.7 Our ITC experiments here demonstrate that the disease mutant Ile67Asn, along with Ile67Ala and Ile67Arg, does not exhibit any significant binding with its partner GLRX5, but the Ile67Val mutant exhibits native-like binding to GLRX5. Next, we investigated the effect of the substitution on Fe-S cluster-binding ability of BOLA3. Standard reconstitution methods produced Fe-S cluster-bound BOLA3, as demonstrated by the ability to transfer this cluster to acceptors such as FDX1 at rates comparable to native BOLA3. Thus, the substitution does not impact reconstitution of homodimer BOLA3. However, formation of [2Fe-2S]-bound BOLA3–GLRX5 heterodimer is impacted in Ile67Asn, Ile67Ala, and Ile67Arg because reconstitution as well as cluster uptake from cluster donor like ISCU results in formation of holo homodimeric GLRX5, and not the BOLA3–GLRX5 heterodimer, as evident from characteristic CD features.
Our functional studies thus demonstrate that the Ile67Asn disease mutation in BOLA3 severely impacts its ability to bind to its partner GLRX5. The BOLA3 and GLRX5 interaction is physiologically relevant, as evident from the detection of physical interaction between the proteins via high-throughput approaches,9 and thereby supporting a functional role for BOLA–GLRX heterodimeric complexes in vivo. While NMR and IM-MS experiments reveal no significant structural or conformational changes as a result of the substitution, we observe that the binding of BOLA3 with GLRX5 is adversely affected. According to the structural model for the BOLA3–GLRX5 heterodimer,9 the GLRX5 binding site on BOLA3 lies quite far from the Ile67 residue of BOLA3. However, NMR chemical shifts indicate that there is significant perturbation in the region around Ile67 of BOLA3 following binding of GLRX5 and the associated GSH molecule, demonstrating that a structural change is induced in the region. Moreover, the possibility of an overall disruption in the dynamics of BOLA3 cannot be ruled out. Nevertheless, such influences do not appear to disrupt BOLA3 homodimer formation in either the apo or holo states, but do perturb distal heterodimer formation with GLRX5. The Ile67Asn substitution likely leads to subtle changes in the local structure within the β-sheet that extend to the cluster binding site and contribute to abolishing the binding of GLRX5. Interestingly, two other derivatives studied here, Ile67Ala and Ile67Arg, also demonstrate similar behavior, while Ile67Val exhibits native-like binding with GLRX5. Valine, having similar size and hydrophobicity compared with isoleucine, is able to conserve wild-type-like function. Clearly, a molecular-level understanding of the structural and dynamical factors that contribute to protein–protein interaction and the selective loss of GLRX5 binding and function remains to be elucidated.
We have previously characterized the [2Fe-2S]-bridged form of holo homodimeric BOLA3 following in vitro reconstitution.35 This holo homodimer species is thermodynamically less stable than the holo BOLA3–GLRX5 heterodimer, as addition of GLRX5 in the presence of GSH immediately converts holo BOLA3 into the holo BOLA3–GLRX5 heterocomplex,8 indicating that the homodimeric BOLA3 is less likely to be physiologically important under normal conditions. However, under MMDS disease condition, our results herein show that Ile67Asn substitution impairs the ability of BOLA3 to form the holo heterodimer with GLRX5, and consequently the holo BOLA3 homodimer is likely to be a physiologically relevant species under such conditions. The substitution does not impact the ability of formation of holo BOLA3 homodimer, which is capable of cluster transfer to a subset of mitochondrial Fe-S proteins.35
In conclusion, our investigations have provided insights into the molecular understanding of the disease-causing perturbation caused by Ile67Asn substitution of BOLA3 in MMDS2. Substitution impairs the binding of BOLA3 with its partner GLRX5, most likely preventing the formation of [2Fe-2S]-bridged BOLA3–GLRX5 heterocomplex. Our conclusions are also suggestive of an important physiological role of [2Fe-2S]-bridged BOLA3–GLRX5 heterocomplex in cluster trafficking to downstream apo targets.
Acknowledgement
Z.T. acknowledges support from the Pelotonia Fellowship Program.
Funding
This work was supported by a grant from the National Institutes of Health [AI072443] and by the NIH P41 Resource for Native Mass Spectrometry Guided Structural Biology [P41GM128577].
Conflicts of interest
The authors declare no conflicts of interest.
Data availability
The data underlying this article are available in the article and in its online supplementary material.
References
Author notes
These authors contributed equally to this work.


![Ile67Asn substitution of human BOLA3 results in the disease phenotype of multiple mitochondrial dysfunctions syndrome-2. Substitution introduces no drastic structural change but impacts the ability of BOLA3 to interact with partner GLRX5 and impairs formation of a [2Fe-2S]-bridged complex of BOLA3 with GLRX5 and subsequent downstream trafficking. The homodimeric form of the BOLA3 derivative can be formed and is active in cluster exchange.](https://oup.silverchair-cdn.com/oup/backfile/Content_public/Journal/metallomics/13/4/10.1093_mtomcs_mfab010/2/m_mfab010gra.jpeg?Expires=1716411424&Signature=ozNffCb0B5RjC0BM-VKrUBltvhkJfrl6~E4E~2pkPE42iYKidAfdc0bhwWGBsIKwTuieOt~YezVIwwiC7wOhK9ptkxX~91QONpsjHxuEe2kv21m5NUctY121g1UG5CL6NFlPHzB2oqhhOt1eJTq-GTmv8GVhMPMi2P49GbDtU02WA3gdA8JkQJa7gJMOdnWHVUSc6lvnpvL1k5kqq2V7EPiVEVdVEmCbXCgVuloa2Lwclcer5Go4idEfkQv82CewH2NdQrX-YOmpoCyYIUlK~wFl3LSiShNz1OwaptIeRDmtBjJAeGcFMz-aPcoZqpl~aDJKQab67m1MPZw2Ws6Axg__&Key-Pair-Id=APKAIE5G5CRDK6RD3PGA)