Abstract
Background
PGPR has substituted chemical fertilizers to enhance the nutrient profile of the soil. Although gene encoding for PGP activity is present in PGPB their activity changes in response to conditions.
Objective
To study comparative genomics for three Klebsiella strains and their PGPR activity in response to in vitro and soil condition.
Methods
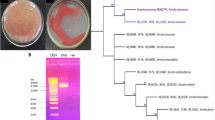
We evaluated the activity of three Klebsiella spp. in two different conditions, specific nitrogen-deficient MS media and greenhouse experiment. Applying comparative genomics, genes encoding for PGP traits were identified from the whole-genome sequencing of the three strains. With the help of the RAST tool kit and functional annotation, a total number of genes encoding for cell wall capsule, nitrogen metabolism, sulfur genes and many other functional groups were identified. With the help of blast circular genome, similarity between GC content, pseudogene and tRNA was represented. The percentage of gene similarity of SSN1 was generated against BLAST with M5a1 and SGM81. Other methods like synteny alignment and orthologous gene clusters were applied to understand the homologous present in three strains.
Results
SSN1 was actively producing the maximum amount of ammonia 10.97 ± 0.29 µmol/mL compared to the other two strains. K. oxytoca M5a1 was considered negative for all PGP traits except ammonia production. The activity of SSN1 was showing a consistent pattern both the conditions whereas M5a1 was only active in vitro condition. Gene encoding for allantoin metabolism allD, allC, allB, allA, allE, allR, allH were identified in SSN1 and M5a1 but was absent in SGM81. The highest COG was shared between SGM81 and SSN1 predicting a maximum number of similar genes. The nif gene cluster was 98 % identical to the M5a1 strain.
Conclusions
Comparatively, SSN1 expressed the additional gene for various PGP traits which suggest higher efficiency of strain in nitrogen deficiency stress.







Similar content being viewed by others
Abbreviations
- PGPB:
-
Plant growth promoting bacteria
- PGP:
-
Plant growth promoting
- WGS:
-
Whole genome sequencing
- RAST:
-
Rapid annotation using subsystem technology
- PEG:
-
Protein encoding gene
References
Agbodjato NA, Noumavo PA, Baba-Moussa F, Salami HA, Sina H, Sèzan A, Bankolé H, Adjanohoun A, Baba-Moussa L (2015) Characterization of potential plant growth promoting rhizobacteria isolated from Maize (Zea mays L.) in central and Northern Benin (West Africa). Appl Environ Soil Sci. https://doi.org/10.1155/2015/901656
Ahmad F, Ahmad I, Khan M (2008) Screening of free-living rhizospheric bacteria for their multiple plant growth promoting activities. Microbiol Res 163:173–181. https://doi.org/10.1016/j.micres.2006.04.001
Aziz RK, Bartels D, Best AA, DeJongh M, Disz T, Edwards RA, Formsma K, Gerdes S, Glass EM, Kubal M (2008) The RAST Server: rapid annotations using subsystems technology. BMC Genom 9:1–15. https://doi.org/10.1186/1471-2164-9-75
Banik A, Mukhopadhaya SK, Dangar TK (2016) Characterization of N 2-fixing plant growth promoting endophytic and epiphytic bacterial community of Indian cultivated and wild rice (Oryza spp.) genotypes. Planta 243:799–812. https://doi.org/10.1007/s00425-015-2444-8
Bhardwaj G, Shah R, Joshi B, Patel P (2017) Klebsiella pneumoniae VRE36 as a PGPR isolated from Saccharum officinarum cultivar Co99004. J App Biol Biotechnol 5:047–052. https://doi.org/10.7324/jabb.2017.50108
Bremner J (1960) Determination of nitrogen in soil by the Kjeldahl method. J Agric Sci 55:11–33. https://doi.org/10.1017/S0021859600021572
Cakmakci ML, Evans H, Seidler R (1981) Characteristics of nitrogen-fixing Klebsiella oxytoca isolated from wheat roots. Plant Soil 61:53–63. https://doi.org/10.1007/BF02277362
Canbolat MY, Bilen S, Çakmakçı R, Şahin F, Aydın A (2006) Effect of plant growth-promoting bacteria and soil compaction on barley seedling growth, nutrient uptake, soil properties and rhizosphere microflora. Biol Fertil Soils 42:350–357. https://doi.org/10.1007/s00374-005-0034-9
Coleman DC, Crossley D Jr (2003) Fundamentals of soil ecology. Academic Press
Darling AE, Mau B, Perna NT (2010) progressiveMauve: multiple genome alignment with gene gain, loss and rearrangement. PLoS One 5:e11147. https://doi.org/10.1371/journal.pone.0011147
D’Autréaux B, Tucker NP, Dixon R, Spiro S (2005) A non-haem iron centre in the transcription factor NorR senses nitric oxide. Nature 437:769–772. https://doi.org/10.1038/nature03953
Galili T, O’Callaghan A, Sidi J, Sievert C (2018) heatmaply: an R package for creating interactive cluster heatmaps for online publishing. Bioinformatics 34:1600–1602. https://doi.org/10.1093/bioinformatics/btx657
Galloway JN, Townsend AR, Erisman JW, Bekunda M, Cai Z, Freney JR, Martinelli LA, Seitzinger SP, Sutton MA (2008) Transformation of the nitrogen cycle: recent trends, questions, and potential solutions. Science 320:889–892. https://doi.org/10.1126/science.1136674
Gang S, Saraf M, Waite CJ, Buck M, Schumacher J (2018) Mutualism between Klebsiella SGM 81 and Dianthus caryophyllus in modulating root plasticity and rhizospheric bacterial density. Plant Soil 424:273–288. https://doi.org/10.1007/s11104-017-3440-5
Geisseler D, Scow KM (2014) Long-term effects of mineral fertilizers on soil microorganisms–a review. Soil Biol Biochem 75:54–63. https://doi.org/10.1016/j.soilbio.2014.03.023
Glass AD (2009) Nitrate uptake by plant roots. Botany 87:659–667. https://doi.org/10.1139/B09-014
Glick BR (2012) Plant growth-promoting bacteria: mechanisms and applications. Scientifica 2012. https://doi.org/10.6064/2012/963401
Gorgone-Barbosa E, Pivello VR, Baeza MJ, Fidelis A (2016) Disturbance as a factor in breaking dormancy and enhancing invasiveness of African grasses in a Neotropical Savanna. Acta Bot Bras 30:131–137
Gupta A, Gopal M, Thomas GV, Manikandan V, Gajewski J, Thomas G, Seshagiri S, Schuster SC, Rajesh P, Gupta R (2014) Whole genome sequencing and analysis of plant growth promoting bacteria isolated from the rhizosphere of plantation crops coconut, cocoa and arecanut. PLoS One 9:e104259. https://doi.org/10.1371/journal.pone.0104259
Iniguez AL, Dong Y, Triplett EW (2004) Nitrogen fixation in wheat provided by Klebsiella pneumoniae 342. Mol Plant Microbe Interact 17:1078–1085. https://doi.org/10.1094/mpmi.2004.17.10.1078
Iqbal A, Qiang D, Zhun W, Xiangru W, Huiping G, Hengheng Z, Nianchang P, Xiling Z, Meizhen S (2020) Growth and nitrogen metabolism are associated with nitrogen-use efficiency in cotton genotypes. Plant Physiol Biochem 149:61–74. https://doi.org/10.1016/j.plaphy.2020.02.002
Izaguirre-Mayoral ML, Lazarovits G, Baral B (2018) Ureide metabolism in plant-associated bacteria: purine plant-bacteria interactive scenarios under nitrogen deficiency. Plant Soil 428:1–34. https://doi.org/10.1007/s11104-018-3674-x
Ji SH, Gururani MA, Chun S-C (2014) Isolation and characterization of plant growth promoting endophytic diazotrophic bacteria from Korean rice cultivars. Microbiol Res 169:83–98. https://doi.org/10.1016/j.micres.2013.06.003
Johnstone TC, Nolan EM (2015) Beyond iron: non-classical biological functions of bacterial siderophores. Dalton Trans 44:6320–6339. https://doi.org/10.1039/c4dt03559c
Khan AL, Waqas M, Asaf S, Kamran M, Shahzad R, Bilal S, Khan MA, Kang S-M, Kim Y-H, Yun B-W (2017) Plant growth-promoting endophyte Sphingomonas sp. LK11 alleviates salinity stress in Solanum pimpinellifolium. Environ Exp Bot 133:58–69. https://doi.org/10.1016/j.envexpbot.2016.09.009
Körschens M, Albert E, Armbruster M, Barkusky D, Baumecker M, Behle-Schalk L, Bischoff R, Čergan Z, Ellmer F, Herbst F (2013) Effect of mineral and organic fertilization on crop yield, nitrogen uptake, carbon and nitrogen balances, as well as soil organic carbon content and dynamics: results from 20 European long-term field experiments of the twenty-first century. Arch Agron Soil Sci 59:1017–1040. https://doi.org/10.1080/03650340.2012.704548
Kuan KB, Othman R, Abdul Rahim K, Shamsuddin ZH (2016) Plant growth-promoting rhizobacteria inoculation to enhance vegetative growth, nitrogen fixation and nitrogen remobilisation of maize under greenhouse conditions. PLoS One 11:e0152478. https://doi.org/10.1371/journal.pone.0152478
Ladha JK, Pathak H, Tirol-Padre A, Dawe D, Gupta RK (2003) Productivity trends in intensive rice–wheat cropping systems in Asia. Improv Product Sustain Rice Wheat Syst Issues Impacts 65:45–76. https://doi.org/10.2134/asaspecpub65.c3
Li H, Durbin R (2010) Fast and accurate long-read alignment with Burrows–Wheeler transform. Bioinformatics 26:589–595. https://doi.org/10.1093/bioinformatics/btp698
Liu D, Chen L, Zhu X, Wang Y, Xuan Y, Liu X, Chen L, Duan Y (2018) Klebsiella pneumoniae SnebYK mediates resistance against Heterodera glycines and promotes soybean growth. Front Microbiol 9:1134. https://doi.org/10.3389/fmicb.2018.01134
McDowell N, Pockman WT, Allen CD, Breshears DD, Cobb N, Kolb T, Plaut J, Sperry J, West A, Williams DG (2008) Mechanisms of plant survival and mortality during drought: why do some plants survive while others succumb to drought? New Phytol 178:719–739. https://doi.org/10.1111/j.1469-8137.2008.02436.x
Moir J, Wood N (2001) Nitrate and nitrite transport in bacteria. Cell Mol Life Sci 58:215–224. https://doi.org/10.1007/PL00000849
Nadeem SM, Zahir ZA, Naveed M, Ashraf M (2010) Microbial ACC-deaminase: prospects and applications for inducing salt tolerance in plants. Crit Rev Plant Sci 29:360–393. https://doi.org/10.1080/07352689.2010.524518
Pace L, Pellegrini M, Palmieri S, Rocchi R, Lippa L, Del Gallo M (2020) Plant growth-promoting rhizobacteria for in vitro and ex vitro performance enhancement of Apennines’ Genepì (Artemisia umbelliformis subsp. eriantha), an endangered phytotherapeutic plant. Vitro Cell Dev Biol Plant 56:134–142. https://doi.org/10.1007/s11627-019-10035-1
Pande A, Pandey P, Mehra S, Singh M, Kaushik S (2017) Phenotypic and genotypic characterization of phosphate solubilizing bacteria and their efficiency on the growth of maize. J Genet Eng Biotechnol 15:379–391. https://doi.org/10.1016/j.jgeb.2017.06.005
Patel NP, Raju M, Haldar S, Chatterjee PB (2020) Characterization of phenazine-1-carboxylic acid by Klebsiella sp. NP-C49 from the coral environment in Gulf of Kutch, India. Arch Microbiol 202:351–359. https://doi.org/10.1007/s00203-019-01742-9
Qureshi JA, Zafar Y, Malik KA (1989) Klebsiella sp. NIAB-I: a new diazotroph, associated with roots of Kallar grass from saline sodic soils. In: Nitrogen fixation with non-legumes. Springer, pp. 115–120. https://doi.org/10.1007/978-94-009-0889-5_15
Rajput MS, Naresh Kumar G, Rajkumar S (2013) Repression of oxalic acid-mediated mineral phosphate solubilization in rhizospheric isolates of Klebsiella pneumoniae by succinate. Arch Microbiol 195:81–88. https://doi.org/10.1007/s00203-012-0850-x
Roeßler M, Sewald X, Müller V (2003) Chloride dependence of growth in bacteria. FEMS Microbiol Lett 225:161–165. https://doi.org/10.1016/S0378-1097(03)00509-3
Rostami H, Giri A, Nejad ASM, Moslem A (2013) Optimization of multiple shoot induction and plant regeneration in Indian barley (Hordeum vulgare) cultivars using mature embryos. Saudi J Biol Sci 20:251–255. https://doi.org/10.1016/j.sjbs.2013.02.008
Saitou N, Nei M (1987) The neighbor-joining method: a new method for reconstructing phylogenetic trees. Mol Biol Evol 4:406–425. https://doi.org/10.1093/oxfordjournals.molbev.a040454
Schwyn B, Neilands J (1987) Universal chemical assay for the detection and determination of siderophores. Anal Biochem 160:47–56. https://doi.org/10.1016/0003-2697(87)90612-9
Stothard P, Wishart DS (2005) Circular genome visualization and exploration using CGView. Bioinformatics 21:537–539. https://doi.org/10.1093/bioinformatics/bti054
Su S, Zhou Y, Qin JG, Yao W, Ma Z (2010) Optimization of the method for chlorophyll extraction in aquatic plants. J Freshw Ecol 25:531–538. https://doi.org/10.1080/02705060.2010.9664402
Temme K, Zhao D, Voigt CA (2012) Refactoring the nitrogen fixation gene cluster from Klebsiella oxytoca. Proc Natl Acad Sci 109:7085–7090. https://doi.org/10.1073/pnas.1120788109
Wang B, Yeun LH, Xue J-Y, Liu Y, Ané J-M, Qiu Y-L (2010) Presence of three mycorrhizal genes in the common ancestor of land plants suggests a key role of mycorrhizas in the colonization of land by plants. New Phytol 186:514–525. https://doi.org/10.1111/j.1469-8137.2009.03137.x
Weller DM, Raaijmakers JM, Gardener BBM, Thomashow LS (2002) Microbial populations responsible for specific soil suppressiveness to plant pathogens. Annu Rev Phytopathol 40:309–348. https://doi.org/10.1146/annurev.phyto.40.030402.110010
Zhou M (2009) Barley production and consumption. In: Genetics and improvement of barley malt quality. Springer, pp 1–17. https://doi.org/10.1007/978-3-642-01279-2_1
Acknowledgements
This work was supported by the Department of Science and Technology UK India Education Research (DST/INT/UK/P-156/2017). The research at the Department of Microbiology and Biotechnology, Gujarat University, School Of Sciences and Imperial College London was funded by British Council [P72898]. We thank Mr. Naman Mangukia, Research Scholar (Gujarat University) for his contribution in data submission at NCBI. DST-FIST “Fund for Improvement of S&T Infrastructure (FIST)” of the Department of Science & Technology (DST), Government of India for providing basic infrastructure grant to facilitate R&D work.
Author information
Authors and Affiliations
Corresponding authors
Additional information
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
About this article
Cite this article
Sharma, S., Gang, S., Schumacher, J. et al. Genomic appraisal of Klebsiella PGPB isolated from soil to enhance the growth of barley . Genes Genom 43, 869–883 (2021). https://doi.org/10.1007/s13258-021-01099-8
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s13258-021-01099-8