Abstract
Algal polysaccharides constitute a diverse and abundant reservoir of organic matter for marine heterotrophic bacteria, central to the oceanic carbon cycle. We investigated the uptake of alginate, a major brown macroalgal polysaccharide, by microbial communities from kelp-dominated coastal habitats. Congruent with cell growth and rapid substrate utilization, alginate amendments induced a decrease in bacterial diversity and a marked compositional shift towards copiotrophic bacteria. We traced 13C derived from alginate into specific bacterial incorporators and quantified the uptake activity at the single-cell level, using halogen in situ hybridization coupled to nanoscale secondary ion mass spectrometry (HISH-SIMS) and DNA stable isotope probing (DNA-SIP). Cell-specific alginate uptake was observed for Gammaproteobacteria and Flavobacteriales, with carbon assimilation rates ranging from 0.14 to 27.50 fg C µm−3 h−1. DNA-SIP revealed that only a few initially rare Flavobacteriaceae and Alteromonadales taxa incorporated 13C from alginate into their biomass, accounting for most of the carbon assimilation based on bulk isotopic measurements. Functional screening of metagenomic libraries gave insights into the genes of alginolytic Alteromonadales active in situ. These results highlight the high degree of niche specialization in heterotrophic communities and help constraining the quantitative role of polysaccharide-degrading bacteria in coastal ecosystems.
Similar content being viewed by others
Introduction
By recycling a large proportion of the available organic matter, heterotrophic bacteria control the oceanic fluxes of carbon and energy [1]. The dissolved fraction is the largest reservoir of oceanic organic matter, with a global estimate of 662 Pg dissolved organic carbon (DOC) [2, 3]. Marine DOC comprises diverse low- and high-molecular-weight compounds, of which carbohydrates constitute 15–50% [4], mostly derived from micro- and macro-algae in the surface ocean. With an estimated standing stock amounting hundreds of megatons in temperate and sub-polar habitats [5, 6], brown macroalgae are major primary producers in coastal regions. Brown macroalgae of the order Laminariales, collectively known as kelps, can release up to 25% of the fixed carbon as exudates, thus contributing ~1.3 kg C m−2 y−1 to the DOC pool [7, 8]. This significantly impacts coastal ecosystems, where kelp forests locally increase DOC concentrations and sustain distinct microbial communities [9]. Previous studies measured 1–1.5 mg C L−1 as DOC in seawater from kelp beds [9, 10]. Kelp exudates contain simple sugars, polysaccharides, proteins, lipids, and aromatic compounds [11, 12]. In particular, the polysaccharide alginate, which comprises ca. half of kelp dry biomass [13], accounts for 5% of the released DOC [11]. Given the density and high primary productivity of kelps in coastal regions, alginate therefore constitutes an abundant resource for planktonic bacteria, available year-round but with potential seasonal variations in quantities [7]. Alginate is a linear polysaccharide consisting of β-D-mannuronic acid and α-L-guluronic acid arranged in homo- or hetero-polymeric blocks and with varying proportions of each motif depending on algal species, tissue, age, and seasons [14, 15]. Many isolated marine bacteria can degrade alginate and use it as a carbon and energy source. Alginolytic culturable representatives belong to Proteobacteria, Flavobacteriales, Firmicutes, and Verrucomicrobiae [16,17,18,19]. In particular, several polysaccharide utilization loci (PUL) dedicated to alginate assimilation have been described, which encode alginate lyases for the breakdown of the substrate into oligosaccharides, SusC/SusD homologs for binding and uptake of degradation products, and cytoplasmic enzymes processing monomers into the central metabolism [20,21,22,23]. Alginolytic PULs are widespread in marine Flavobacteriaceae and were transferred to Gammaproteobacteria [20]. Their expression is tightly controlled by PUL-encoded regulators, allowing a rapid and massive overexpression when alginate becomes available, and might be fine-tuned by yet unknown cross-regulation, catabolic repression, or substrate prioritization mechanisms [24,25,26]. Depending on the type and cellular localization of their alginate lyases, marine bacteria adopt different ecophysiological strategies leading to niche specialization towards polymeric alginate vs. oligomeric products [27, 28]. Despite the characterization of alginate catabolism in cultivated strains, identifying the members of natural bacterial communities that use alginate remains a challenge. Previous studies showed that alginate amendments induce shifts in seawater community composition [29,30,31,32] and provided insights into the global community response and alginolytic potential. However, these approaches cannot directly link alginate utilization to individual bacterial taxa nor estimate substrate assimilation at the single-cell level, hindering our comprehension of alginate fate in marine ecosystems. To date, few studies focused on the direct detection and quantification of the utilization of a given substrate by individual bacteria in natural samples, using e.g., microautoradiography, Raman spectroscopy, fluorescently-labeled substrates, or stable isotope probing (SIP) [33, 34]. To our knowledge, SIP has seldom been applied using complex marine organic matter [35,36,37,38,39], and never to macroalgae-derived polysaccharides, despite their importance in the carbon cycle. Here, we combined DNA-SIP with halogen in situ hybridization coupled to chemical imaging via nanoscale secondary ion mass spectrometry (HISH-SIMS) to trace 13C-labeled alginate into specific incorporators within a coastal seawater community and quantify the uptake activity at the single-cell level. We show that most of the alginate is used by a few initially rare Flavobacteriaceae and Gammaproteobacteria taxa and provide the first single-cell estimates of alginate incorporation rates.
Methods
Alginate production
Natural (12C-natural) and 13C-enriched alginates were prepared as described previously [40]. Briefly, the kelp Laminaria digitata was cultivated with either natural (98.9 12C%) or 13C-labeled (99 13C%) NaHCO3. Alginate was chemically extracted as detailed in Supplementary Methods. The average molecular weight of alginate estimated using multi-angle laser light scattering (MALLS) was Mw = 1.7 105 Da. Alginates were assayed for residual proteins and DNA using the Qubit Protein Assay kit and dsDNA HS assay kit (ThermoFisher Scientific), respectively. 12C-natural alginate contained 13.7 mg protein and 284 µg DNA g−1 (i.e., 98.6% purity). 13C-enriched alginate contained 10.1 mg protein and 288 µg DNA g−1 (i.e., 99.0% purity). The absence of inhibitors was confirmed by checking the growth of the alginolytic strain Zobellia galactanivorans DsijT in minimum medium supplemented with 12C-natural or 13C-enriched alginate (1 g l−1). Alginate isotopic ratio was measured by elemental analysis coupled to isotope ratio mass spectrometry (EA-IRMS) as described previously [40]. 12C-natural and 13C-enriched alginate had 1.1337 ± 0.0004 and 3.6279 ± 0.0009 13C atom percent, respectively (mean ± s.d. n = 3 technical replicates). Before microcosm amendment, alginates were solubilized (5 g l−1) in Tris-HCl 50 mM pH 8.0 and autoclaved. The average molecular weight of autoclaved alginate determined by MALLS analysis was Mw = 1.4 105 Da with a polydispersity index of 1.364, corresponding to an average degree of polymerization of ca. 800 monomers.
Seawater microcosms
Natural seawater was collected at 12.30 pm on January 21 2019 from a kelp-dominated tidal pool at Le Bloscon, Roscoff, France (48°43'33.18“N, 3°58'7.58“W). Average surface water temperature, salinity, and pH are 10–12 °C, 35.0–35.2 g kg−1,and 7.75–7.80 in January in this area (values from long-term monitoring SOMLIT Estacade Station, 1.2 km from sampling site). Autoclaved plastic carboys were rinsed three times with water from the site before sampling. Seawater from different carboys was pooled and homogenized in a sterile container. Three aliquots (950 ml) were filtered for DNA extraction at T0. The remaining seawater was distributed in 1-liter aliquots into sterile 5-liter flasks with 0.1% marine ammonium mineral salts (DSMZ #1313). Microcosms were amended with either 12C-natural alginate (n = 3) or 13C-enriched alginate (n = 3) at 20 mg l−1, corresponding to ca. 8 mg C l−1. Unamended controls (n = 3) were prepared by adding the same volume of Tris-HCl 50 mM pH 8.0 without alginate. All flasks were incubated at 15 °C, 130 rpm in the dark. The set-up was complete within 3 h after sampling. Aliquots were sampled from all microcosms at T = 0, 18.5, 24.5, 42.5, and 47 h for flow cytometry and uronic acid measurements, and only at T = 47 h for bulk isotope, CARD-FISH and HISH-SIMS analyses, and DNA extraction.
Flow cytometry
Aliquots (250 µl) were fixed with 1.25 µl glutaraldehyde 50% and diluted with an appropriate volume of 1X TE buffer containing SybrGreen (Life Technologies, 2 µl for 15 ml TE buffer). Cells were counted in technical triplicates on BD Accuri C6 Plus Flow Cytometer.
Uronic acid measurements
Aliquots (1 ml) were centrifuged 10 min at 7000 rpm. To follow alginate concentrations, uronic acids were quantified in supernatants using the meta-hydroxy-di-phenyl (MHDP) method [41]. Samples (200 µl) were mixed with 20 µl of 4 M sulfamic acid and 1.2 ml of 75 mM sodium tetraborate. After incubation (20 min, 80 °C), 40 µl of MHDP 0.15% were added. OD525 was measured after 10 min on a spectrophotometer and compared to a standard curve of glucuronic acid from 2.5 to 100 µg ml−1.
CARD-FISH and HISH-SIMS analyses
Aliquots (1.8 ml) were preserved with paraformaldehyde (1%, 1 h at room temperature) and stored in PBS/ethanol 1:1 (v/v) at −20 °C. Fixed cells (500 µl aliquots) were filtered under moderate vacuum on 0.2 µm polycarbonate membranes (Isopore) previously sputtered with a 80/20 Au/Pd alloy, and washed twice with 5 ml PBS. Filters were air-dried and stored at −20 °C until analysis. Hybridization procedures were performed as described previously [42]. Filters were embedded in 0.1% low-melting point agarose. Cells were permeabilized with 10 mg ml−1 lysozyme in TE buffer (30 min, 37 °C). Endogenous peroxidases were inactivated in 3% H2O2 (10 min, room temperature). Hybridizations were performed at 46 °C for 3 h in 300 µl buffer (35% formamide) containing 28 nM of the following HRP-labeled probes: GAM42a (with competitor BET42a) targeting most Gammaproteobacteria [43], CF319a targeting most Flavobacteriales and other members of the CFB group [44] and NON338 used as a negative control [45]. Signal amplification was conducted at 46 °C for 45 min, using fluorine-containing tyramides synthesized from OregonGreen 488-X succinimidyl ester (Molecular Probes) as described previously [46]. Portions of hybridized filters were stained with DAPI (1 µg ml−1) before bacterial counts on an Olympus BX60 microscope with epifluorescence irradiation.
HISH-SIMS analysis was performed using a NanoSIMS-50 Ion microprobe (CAMECA, Gennevilliers, France) in scanning mode [47, 48] (details in Supplementary Methods). After Cs+ pre-implantation, five secondary ions were monitored: 12C−, 19F−, 12C14N−, 13C14N−, and 32S−. Images of 32S−, [19F−]/[32S−] and 13C atom fraction were processed using ImageJ [49]. The 13C atom fraction map was established from 12C14N− and 13C14N− images based on pixel-by-pixel calculation as follows:
13C At% = [13C14N−]/([12C14N−] + [13C14N−]) x 100%
Single-cell carbon assimilation rates were inferred from NanoSIMS data, following calculations developed in [50] and detailed in Supplementary Methods. Briefly, the fraction of carbon assimilated (KA) during incubation in microcosms containing 13C-enriched alginate was calculated for each selected cell, considering a carbon dilution of 29.38% due to the CARD-FISH treatment [51]. Volume-specific carbon assimilation rates were inferred from KA values based on the measured biovolume for each cell, the incubation time, and the partial density of carbon in bacterial cells.
Bulk isotopic analysis
Aliquots (1 ml) were filtered onto precombusted GF filters (0.45 µm pore size), which were frozen at −20 °C until EA-IRMS analysis. Carbon isotopic ratios (R = 13C/12C) were determined on filters folded into tin capsules, using a CHN analyzer (ThermoFinnigan 1112 Series) interfaced with a mass spectrometer (ThermoFinnigan MAT Delta Plus) via a Conflow III open split interface. Abundances were calculated in relation to Vienna Pee Dee Belemnite-limestone (V-PDB), using in-house casein standards calibrated against IAEA-600 and IAEA-CH-6 international standards.
Bulk carbon assimilation was calculated as atom percent excess (APE) as follows:
where Rf and Ri are the 13C isotope ratio at the final and initial sampling time, respectively.
Bulk carbon incorporation rates (Fbulk) were calculated as follows:
where KA is the fraction of assimilated carbon (calculated as described in Supplementary Methods), ρC is the amount of carbon per cell, A is the final cell abundance estimated through flow cytometry and t is the incubation time. We considered a range of ρC = 10–280 fg C cell−1 [50].
DNA extraction
Seawater (950 ml) was filtered on 0.22 µm Sterivex-GP polyethersulfone filters (Merck) using a peristaltic pump under moderate flow with a 3 µm pre-filter. DNA was extracted using the NucleoSpin PlantII kit (Macherey–Nagel) as in Ramond et al. [52], eluted in 100 µl and quantified using a Qubit fluorometer.
Isopycnic centrifugation and fractionation
DNA from 12C-natural and 13C-enriched alginate-amended microcosms was fractionated on CsCl density gradients as described previously [53]. DNA (5 µg) was mixed with gradient buffer (0.1 M Tris-HCl, 0.1 M KCl, 1 mM EDTA) and CsCl solution to a final buoyant density (BD) of 1.725 g ml−1. Ultracentrifugation was performed in a vertical rotor (VTi 65.2, Beckman), at 15 °C, 176,985 g for 64 h. Sixteen fractions of 340 µl were collected from each tube. The refractive index (RI) of each fraction was measured with a refractometer (VWR) and corrected to account for the gradient buffer RI using the equation \(RI_{corrected} = RI_{observed} - (RI_{buffer} - 1.3333)\) [54]. BD was calculated from corrected RI using the equation \(BD = a \times RI - b\) [55] where a and b are coefficients for CsCl at 20 °C (a = 10.9276; b = 13.593) [56]. The average difference in BD between successive fractions was 0.0028. DNA was recovered by overnight precipitation with 20 µg glycogen (MP Biomedicals) and 700 µl of polyethylene glycol solution (30% PEG 6000, 1.6 M NaCl) followed by centrifugation for 45 min at 13,000 g. DNA was rinsed with 70% ethanol, air-dried, and resuspended in 30 µl of molecular-biology grade water.
16S rRNA gene metabarcoding
Library preparation and sequencing were performed as described previously [57] and detailed in Supplementary Methods. A 464-bp fragment of 16S rRNA genes was amplified from non-fractionated total DNA samples [T0 (n = 3), 12C-natural alginate (T47-ALG12, n = 3), 13C-enriched alginate (T47-ALG13, n = 3) and unamended controls (T47-CTRL, n = 3)] and gradient fractions (n = 14 fractions per gradient) using primers S-D-Bact-0341-b-S-17 and S-D-Bact-0785-a-A-21 [58]. The library was prepared using the Nextera XT DNA library prep kit (Illumina). MiSeq sequencing (2 × 300 cycles, Illumina) yielded a total of 7,250,023 paired-end sequences from 96 samples (accessible at NCBI under BioProject accession PRJNA686971). Quality-filtered reads were processed using default parameters of DADA2 [59] implemented in QIIME 2 v2018.8 [60] and clustered de novo in operational taxonomic units (OTUs) at 97% using vsearch [61]. Taxonomy was assigned using SILVA ssu132 Ref NR99. OTUs representing <0.001% of total sequences or affiliated to chloroplasts, mitochondria, or archaea were discarded (2.6% of input sequences). The final dataset comprised 1074 OTUs and 2,909,992 sequences, ranging from 12,948 to 174,305 sequences per sample. Data were further analyzed in phyloseq [62]. Alpha-diversity indices for non-fractionated samples were calculated on the non-transformed dataset. Beta-diversity analyses were performed on Hellinger-transformed datasets using PCoA on a weighted Unifrac distance matrix. The effect of alginate amendments on total community structure was tested using PERMANOVA with 999 permutations. Differential abundance analysis between unamended controls (n = 3) and alginate-amended microcosms (n = 6) was performed using DESeq2 [63] based on the negative binomial distribution with Wald test and parametric fitting. Differences were considered significant when Benjamini–Hochberg corrected p were <0.05 and log2 fold-change >1.
DNA-SIP
For DNA-SIP analysis, we retained OTUs that were present at least once in each of the three gradients from 13C-enriched samples. This filtered dataset comprised 183 OTUs and 2,173,323 sequences, i.e., 98% of the data obtained from gradient fractions. Analysis was performed with HTSSIP v1.4.1 [64] using multiple-windows high-resolution SIP (MW-HR-SIP) [54, 65], with a sparsity cutoff of 0.25, a log2 fold-change null threshold of 0.25 and a significance threshold α = 0.05. Briefly, MW-HR-SIP identifies incorporators by utilizing DESeq2 to detect OTUs that have a higher relative abundance in multiple overlapping “heavy” BD windows of 13C-enriched vs. 12C-natural alginate gradients. Four heavy BD windows were tested (1.715–1.730, 1.720–1.735, 1.725–1.740, 1.730–1.745). Representative sequences of each 13C-incorporating OTU were analyzed using blastn on the NCBI 16S rRNA database.
Metagenomic fosmid library
A metagenomic fosmid library was prepared from 13.5 µg unfractionated DNA from alginate-amended microcosms following the CopyControl Fosmid Library production kit protocol (Epicentre). Briefly, end-repaired DNA was cloned into the pCC2FOS vector and transfected in Escherichia coli EPI300-T1R. The clone library was recovered on LB-agar containing chloramphenicol (12 µg ml−1) overnight at 37 °C, and stored at −80 °C in 96-well microplates. Clones were screened on LB-agarose containing chloramphenicol (12 µg ml−1), arabinose (0.2 g l−1), and sodium alginate (7 g l−1). After 4 days at 37 °C, plates were flooded for 10–30 min with 10% cetylpyridinium chloride to detect alginolytic activity as clearing zones against an opaque background. Recombinant alginate lyase AlyA1 from Zobellia galactanivorans DsijT [66] was used as positive control. Sequencing libraries were prepared from positive fosmid DNA using the Nextera XT DNA kit (Illumina) and sequenced using MiSeq v3 PE300 (Illumina). Reads were assembled using SPAdes v3.11 [67], resulting in one contig of 42,237 bp, including 34,093 bp of metagenomic insert and 8144 bp of pCC2FOS for fosmid F10, and one contig of 51,052 bp, including 42,696 bp of metagenomic insert and 8356 bp of pCC2FOS for fosmid F25. The F10 and F25 metagenomic insert sequences were deposited under Genbank accessions MW442085 and MW442086, respectively. ORFs were predicted and annotated using RAST [68] and prokka [69]. Protein sequences were analyzed using blastp on the nr database. Predicted alginate lyases were annotated using dbcan [70], CUPP [71], SignalP 5.0 [72] and Pfam [73].
Results
Dynamics of alginate utilization and effect on bacterial community
Cell abundance increased from 6.82 ± 0.06 105 to 2.25 ± 0.07 107 cell ml-1 in both 12C or 13C- alginate-amended treatments (Fig. 1). Final cell abundance was 3-fold lower in unamended controls compared to alginate-amended microcosms. Most of the added alginate (95-100%) was consumed between 18 and 25 h of incubation.
Two-days incubation strongly decreased the richness (Chao1 index) and diversity (Shannon and Simpson index) of the seawater bacterial community compared to its initial state (Figure S1A). The community was more diverse in alginate-amended microcosms compared to unamended controls (Welch’s t-test, p = 0.002 and p = 0.021 for Shannon and Simpson, respectively), although one of the microcosms amended with 13C-enriched alginate was an outlier with low diversity. Furthermore, the OTU-level community structure of alginate-amended microcosms differed strongly from that of the initial community and unamended microcosms (Figure S1B, PERMANOVA, F = 14.8, p = 0.002). There was no significant effect of the alginate type (12C-natural vs. 13C-enriched) on the final community structure (PERMANOVA, F = 0.64, p = 1).
A total of 389 genera were detected in the seawater bacterial community (Fig. 2). The initial community was dominated by the Flavobacteriaceae NS5 marine group (relative sequence abundance 13.5 ± 0.4%, mean ± s.e.m., n = 3) and the alphaproteobacterial SAR11 Clade Ia (9.6 ± 0.1%), Planktomarina (3.7 ± 0.5%) and Amylibacter (3.6 ± 0.2%). Incubation for 47 h with or without alginate amendment induced drastic shifts in taxonomic composition. The unamended controls were dominated by the Epsilonproteobacteria genus Arcobacter (33.1 ± 4.2%), followed by Colwellia (20.7 ± 1.3%) and Glaciecola (10.6 ± 1.4%) within the order Alteromonadales. Such a short-term “bottle effect” has previously been observed for seawater microcosms [74] and might be due to partial oxygen depletion, modified nutrient availability, or accumulation of metabolites in sealed containers compared to open environments. Overall, similar taxonomic composition was observed in all the alginate-amended microcosms irrespective of isotope enrichment. The most prevalent genera were Psychrobium (20.1 ± 1.2%, n = 6), Colwellia (19.2 ± 1.2%) and Psychromonas (11.6 ± 0.6%) within Alteromonadales (Gammaproteobacteria), as well as Wenyingzhuangia (11.1 ± 4.6%) and Tenacibaculum (5.4 ± 0.5%) within Flavobacteriaceae (Bacteroidia). One replicate microcosm amended with 13C-enriched alginate (T47-ALG13-1) partly differed from the other alginate-amended microcosms due to a strong enrichment in the genus Wenyingzhuangia that accounted for 34% of the sequences. The prevalence of Gammaproteobacteria and Flavobacteriales cells in microcosms amended with 13C-enriched alginate was confirmed by CARD-FISH, with final proportions reaching 45 ± 11 and 25 ± 9%, respectively (Table S1).
Individual plots are shown for triplicate incubations in each condition. Genera individually representing less than 1% relative sequence abundance have been collapsed in the “<1%” category. Abbreviations are shown on the bars for abundant groups discussed in the text. Arc Arcobacter; Col Colwellia; Gla Glaciecola; Wen Wenyingzhuangia; Ten Tenacibaculum; Psb Psychrobium; Psm Psychromonas.
A total of 64 OTUs belonging to 28 genera were found significantly more abundant in alginate-amended microcosms (n = 6) compared to unamended controls (n = 3), while 15 OTUs were significantly more abundant without amendment (Figure S2). Most of the OTUs significantly enriched in alginate-amended microcosms belonged to Gammaproteobacteria (34/64, including 24 Alteromonadales) and Bacteroidia (28/64, including 27 Flavobacteriales). The most alginate-responsive OTUs were affiliated to Psychrobium, Wenyingzhuangia, and Colwellia.
Signals of 13C-assimilation at the community and single-cell levels
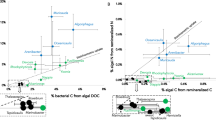
We analyzed carbon incorporation in microcosms amended with 13C-enriched alginate, by measuring bulk 13C uptake with EA-IRMS and cell-specific uptake with HISH-SIMS. Bulk samples showed a significant 13C At% excess of 0.34 ± 0.05. Using lower and upper estimates of 10–280 fg C cell−1 [50], extrapolated bulk rates of carbon incorporation from alginate ranged from 16 ± 3 to 445 ± 81 µg C L−1 d−1. Based on the strong enrichment in Flavobacteriales and Gammaproteobacteria following alginate amendment, HISH-SIMS analysis was focused on these two groups using probes CF319a and GAM42a, respectively. Cells from both groups showed significant 13C enrichment (Fig. 3, Table S2, Figure S3). 13C-enriched GAM42a-positive cells were all short and thick rods (34 analyzed cells, L = 1.56 ± 0.09 µm, W = 0.96 ± 0.05 µm). Two morphologies were detected for 13C-enriched CF319a-positive cells: (i) thin rods with length <1.2 µm (13/24 analyzed cells, L = 0.79 ± 0.05 µm, W = 0.34 ± 0.03 µm) and (ii) longer rods reaching up to 6.5 µm (11/24 analyzed cells, L = 2.42 ± 0.45 µm, W = 0.65 ± 0.11 µm, see example in Fig. 3A-C). 13C-enriched cells showed atom percent enrichment from 0.03 to 2.33 at %. Volume-specific carbon assimilation rates ranged from 0.14 to 27.50 fg C µm−3 h−1 (Fig. 4). The average carbon assimilation rate did not differ significantly between Flavobacteriales and Gammaproteobacteria (5.43 ± 0.91 and 6.34 ± 0.33 fg C µm−3 h−1, Welch’s t-test, t = −0.94, p = 0.35), but values were significantly more dispersed for Flavobacteriales (F test of variance, F = 5.43, p < 0.001).
Cells were hybridized with CF319a probe targeting Flavobacteriales (A–F) or GAM42a probe targeting Gammaproteobacteria (G–L), using fluorine-containing tyramides. Rows show parallel acquisitions of the same region. Columns display secondary ion images of 32S as total biomass indicator (A, D, G, and J; a.u: arbitrary intensity unit), the ratio 19F/32S as a marker for cell identity (B, E, H, and K) and the 13C atom fraction inferred from secondary ions (13C14N, 12C14N) as indicator of 13C incorporation from alginate (C, F, I, and L), in HSI (Hue-Saturation-Intensity) color scale. Scale bars: 2 µm.
DNA-SIP identification of alginate incorporators
To specifically identify cells deriving their carbon from alginate, we performed a DNA-SIP analysis of samples collected after 47 h of incubation. Seven OTUs were detected as incorporators of 13C from alginate, all affiliated with Flavobacteriaceae and Gammaproteobacteria (Table 1). The strongest signals (difference in relative abundance in heavy gradient fractions of 13C-enriched samples compared to corresponding fractions of 12C-natural samples) were found for OTU A (log2-fold change = 2.30) affiliated to Wenyingzhuangia and OTU E (log2-fold change = 2.31) affiliated to Colwellia (Figure S4). We further investigated the variation of relative abundance for these seven OTUs in total bacterial communities (Fig. 5). They all were initially at low relative sequence abundance in the sampled seawater (range 0–0.72%). For all but one 13C-incorporating OTU, the relative abundance increased in alginate-amended microcosms while it decreased in unamended controls (range 0–0.14%). The only exception was OTU G affiliated to Leucothrix, whose abundance stayed stable in all conditions tested (0.21–0.97%). After 47 h with alginate, two 13C-incorporating OTUs became dominant in the total bacterial community, namely OTU A (Wenyingzhuangia, final relative abundance 10.4 ± 4.6%, n = 6) and OTU D (Psychromonas, 8.4 ± 0.5%). Combining these results with final cell counts and NanoSIMS-derived cell-specific carbon assimilation rates, we estimated the carbon assimilation attributed to the 13C-incorporating OTUs identified by HTS-DNA-SIP. In total, the estimate of carbon assimilation by the seven 13C-incorporating OTUs was 438 ± 62 µg C L−1 d−1, at the upper end of the estimate obtained from bulk analysis. Collectively, the three Flavobacteriaceae OTUs A, B, and C accounted on average for 146 ± 55 µg C L−1 d−1. This estimate of carbon assimilation was significantly higher (Welch t-test, t = −2.63, p = 0.043) for the four Gammaproteobacteria OTUs D, E, F, and G, which collectively accounted for 292 ± 12 µg C L−1 d−1.
Screening and analysis of metagenomic libraries
A metagenomic library consisting of 5000 fosmid clones with ~40 kb insert was prepared using DNA obtained from alginate-amended microcosms, representing ~200 Mb of total screened DNA. Using Chao1 index as a proxy for the number of species (average value 123 in alginate-amended microcosms, see Figure S1) and an average genome size of 4 Mb, we can roughly estimate that our screening effort represents 40% of the total community. Two clones showed an alginolytic activity, corresponding to fosmids F10 and F25. Analysis of the fosmid sequences revealed that F10 and F25 contained metagenomic inserts of ~34.1 and 42.7 kb, respectively, encoding 33 and 26 open reading frames (ORFs) in both orientations (Fig. 6A). Both inserts had a homogeneous GC content of 40%. All but one ORF in the F10 insert showed high sequence similarity with proteins from Alteromonadales isolates (i.e. Gammaproteobacteria, Table S3). In particular, 31/33 ORFs in F10 had their best blastp hits with Psychromonas isolates. Similarly, 21/26 ORFs of the F25 metagenomic insert were related to Alteromonadales, in particular to Colwellia isolates. Both inserts contained two predicted alginate lyase ORFs (ORFs 1 and 2 in F10, ORFs 4 and 16 in F25) together with ORFs related to signal transduction, transport or outer membrane proteins, amino acid synthesis, and DNA replication (Fig. 6B). In addition, F10 insert encoded ORFs related to assimilatory sulfate reduction and protein translation. Modular analysis showed that F10 ORF1 encoded a predicted 22.5 kDa cytoplasmic alginate lyase of the polysaccharide lyase family PL7, subfamily 3. ORF 2 from F10 and ORF 4 from F25 both encoded secreted multimodular alginate lyases consisting of two F5/8 type C domains and a C-terminal PL18 catalytic domain. Despite this similar architecture, ORF 2 and 4 only shared 32% protein sequence identity. Finally, ORF16 from F25 encoded a predicted 83.3 kDa secreted alginate lyase comprising two PL17 domains of subfamily 2.
A Genomic map of the DNA insert of the two fosmids F10 and F25 for which positive alginolytic activity was detected. ORFs are numbered sequentially and colored according to predicted function. B. Molecular weight and modular analysis of the four predicted alginate lyases in fosmid inserts. S Sec signal peptide type 1, F5/8-C F5/8 type C domain, PL polysaccharide lyase family (CAZy classification) with subfamily when available.
Discussion
Estimating the quantitative role of specific heterotrophic bacteria for alginate degradation is essential to constrain carbon budgets in macroalgae-dominated coastal habitats. Here, alginate amendments to the DOC pool of coastal seawater decreased the bacterial diversity in a few hours and favored the growth of a limited number of Flavobacteriaceae and Gammaproteobacteria OTUs. Combined HISH-SIMS and DNA-SIP analyses provided evidence of alginate assimilation at the single-cell level, directly linking bacterial metabolic functioning to taxonomic identity. The average incorporation rate (~5.89 fg C µm−3 h−1) was 10-fold higher than that recently measured for phytoplankton-derived DOC uptake by Flavobacteriales and Rhodobacteraceae cells (~0.42 fg C µm−3 h−1) [39]. These higher rates might partly be due to the increased alginate availability in microcosms compared to natural environments. They might also highlight the adaptation of the detected 13C incorporators to efficient utilization of alginate pulses in coastal seawater. In addition, it underlines the quantitative importance of alginate utilizers for carbon cycling in coastal regions, both for bacterial secondary production and remineralization. Assuming a complete utilization of alginate during the 2-day experiment (Fig. 1), we can extrapolate that maximum 12% of the alginate carbon was assimilated into bacterial biomass, the remaining 88% being respired to CO2. This fits previous estimates of carbon flow for kelp mucilage and debris [75, 76], showing bacterial conversion efficiencies of 11–27%. Yet, the large dispersion observed for alginate incorporation rates by Flavobacteriales cells (Fig. 4) suggests that they adopt variable nutrient acquisition strategies, while it seems more homogeneous for gammaproteobacterial alginate consumers. This confirms the high degree of niche specialization within closely related clades of marine Bacteroidetes [77, 78].
DNA-SIP revealed that only a few initially rare taxa incorporated 13C from alginate into their biomass. The small number of detected alginate incorporators might partially reflect (i) the relatively low isotopic ratio of the alginate (3.6 13C At%), compared to traditional SIP experiments with highly labeled substrates and (ii) the high specificity of MW-HR-SIP analysis compared to other DNA-SIP methods [65]. We cannot exclude the possibility that other OTUs enriched in alginate-amended microcosms utilized the substrate, though not enough to be detected using DNA-SIP. However, the proportion of detected alginate incorporators (7/1074 OTUs, i.e., 0.65%) matches other MW-HR-SIP studies on the assimilation of cellulose (63/5940 OTUs, 1.06%) or xylose (49/5940 OTUs, i.e., 0.82%) by soil bacteria [54]. Furthermore, estimation of total carbon assimilation by the seven incorporating OTUs matches the upper limit of bulk measurements (~445 µg C L−1 d−1), indicating DNA-SIP likely accounted for most quantitatively relevant alginate-assimilating bacteria. Therefore, the small number of detected alginate incorporators in coastal seawater rather indicates that alginolytic systems are not phylogenetically widespread and that polysaccharide availability only selects specialized taxa with distinct ecological niches, as suggested previously [29, 32, 79]. Alginate-assimilating bacteria might be more abundant on algal surfaces, where substrates are constantly present at higher concentration [16]. The alphaproteobacterial SAR11 and flavobacterial NS5 clades dominated the initial bacterial community. Both clades are ubiquitous in open ocean and coastal waters worldwide [78, 80, 81]. They generally feature small genomes of ~2 Mb, which might minimize their metabolic requirements and allow their growth in nutrient-depleted environments [82,83,84]. The NS5 marine group responds positively to phytoplankton blooms [85] and some NS5 metagenome-assembled genomes feature alginate PULs [86]. Yet, our data suggest they were outcompeted by initially rare copiotrophic taxa that grew rapidly by exploiting the transient increase in alginate. These efficient and fast-growing alginate incorporators notably belonged to Wenyingzhuangia, Polaribacter and Tenacibaculum (Flavobacteriaceae), and Psychromonas and Colwellia (Alteromonadales). This corroborates previous findings on cultivated strains that showed alginate-degrading activities in these genera [17, 87,88,89]. Recent studies also identified alginate-degrading enzymes and PULs in metagenome-assembled genomes related to Polaribacter, Colwellia, Psychromonas, and Tenacibaculum in seawater above macroalgal forests [90] or during spring phytoplankton bloom [86]. Here, functional screening of metagenomic libraries confirmed Alteromonadales representatives were among the major alginate degraders and featured complementary alginate lyases from different CAZy families. PL7-3 and PL17-2 families are widespread in alginolytic bacteria and comprise endoguluronate lyases [66] and exolytic oligoalginate lyases [91], respectively. By contrast, PL18 alginate lyases seem specific to Proteobacteria, as suggested by the lack of reported family members in any Bacteroidetes on the CAZy database (http://www.cazy.org/PL18_bacteria.html). The N-terminal pairs of F5/8 type C domain, also known as carbohydrate-binding module of family 32, likely influence the catalytic activity, enzyme stability, or substrate binding as shown for other alginate lyases [92,93,94]. In both fosmid inserts, alginate lyase genes were close to ilv genes involved in branched amino acid biosynthesis (valine, leucine, isoleucine). Since these pathways incorporate pyruvate, an end-product of alginate degradation [95], the colocalization of alginate lyase and ilv genes might indicate an important metabolic route for carbon assimilation from alginate.
Directly linking alginate assimilation to a few Flavobacteriaceae and Alteromonadales strengthens previous studies on the effect of alginate amendments on community composition. Wietz et al. [32] showed that the addition of 0.001% soluble alginate to seawater from the Patagonian continental shelf induced a strong increase in Alteromonadaceae that could reach 80% final relative abundance. Similar alginate amendment to Arctic seawater favored a few Bacteroidia and Gammaproteobacteria, including Polaribacter and Colwellia [30]. Alginate particles added to coastal surface seawater from California [31] or Massachusetts [29] also induced the growth of Bacteroidetes and Alteromonadales, including Psychromonas. Therefore, it appears that closely related taxa respond to alginate in distant, contrasting environments. Their success for polysaccharide degradation relies on numerous PULs targeting diverse algal compounds, including alginate. The tight control of flavobacterial PUL expression that allows overexpression of alginolytic genes within minutes after alginate becomes available [24] might explain the rapid substrate exploitation in microcosms. Recently, a tripartite conceptual model of the different bacterial strategies at play during marine polysaccharide degradation was proposed [96]. Selfish bacteria break up polysaccharides and internalize oligomers with virtually no loss of low-molecular-weight products to the environment [34], contrasting with external hydrolyzers that use extracellular enzymes and release degradation products to the milieu. The liberated products are subsequently used by scavenging bacteria, which cannot or do not produce extracellular enzymes. In this model, both selfish and external hydrolyzers could compete for highly complex polysaccharides present in high abundance and often forming gels [96]. Here, the major flavobacterial alginate incorporators could represent selfish bacteria, owing to their concerted substrate degradation and uptake, archetypal to PULs involving SusC/D-like membrane proteins in Bacteroidetes but not in Gammaproteobacteria [97]. They may have varying efficiency in assimilating degradation products, as suggested by their large range of incorporation rates. Conversely, the Alteromonadales incorporators likely represent external hydrolyzers, as recently proposed in a study of substrate utilization in the Atlantic Ocean where “sharing” organisms belonged mostly to Alteromonadaceae [98]. The fact that three of the four predicted alginate lyases encoded in the fosmid inserts feature signal peptides supports this hypothesis. The other detected alginate incorporators (i.e. Marinomonas and Leucothrix) might be scavengers, relying on previous degradation by other microorganisms and therefore not showing the same rapid increase in relative abundance. Indeed, Marinomonas isolates generally do not degrade polymeric alginate [99] and metatranscriptomics of coastal seawater suggested Oceanospirillales are efficient scavengers of monosaccharides from DOM [100]. Investigation of isotope incorporation at successive time points would help disentangle the interactions between alginate consumers with different strategies, and decipher the dynamics of carbon transfer via the microbial loop. Furthermore, our work paves the way for isotopic tracing studies using purified macroalgal compounds, exudates, or intact tissues to identify (1) the substrate niche(s) of marine bacteria and (2) trophic chains based on utilization of specific substrates via diverse detrital pathways, leading thus to better characterization and quantification of their contributions to the coastal carbon cycle.
References
Azam F, Malfatti F. Microbial structuring of marine ecosystems. Nat Rev Micro. 2007;5:782–91.
Hansell DA, Carlson CA, Repeta DJ, Schlitzer R. Dissolved organic matter in the ocean a controversy stimulates new insights. Oceanography. 2009;22:202–11.
Moran MA, Kujawinski EB, Stubbins A, Fatland R, Aluwihare LI, Buchan A, et al. Deciphering ocean carbon in a changing world. Proc Natl Acad Sci. 2016;113:3143–51.
Benner R, Pakulski JD, Mccarthy M, Hedges JI, Hatcher PG, Pakulski JD, et al. Bulk chemical characteristics of dissolved organic matter in the ocean. Science. 1992;255:1561–4.
Carpenter LJ, Liss PS. On temperate sources of bromoform and other reactive organic bromine gases. J Geophys Res. 2000;105:20539–47.
Mac Monagail M, Cornish L, Morrison L, Araújo R, Critchley AT. Sustainable harvesting of wild seaweed resources. Eur J Phycol. 2017;52:371–90.
Abdullah MI, Fredriksen S. Production, respiration and exudation of dissolved organic matter by the kelp Laminaria hyperborea along the west coast of Norway. J Mar Biol Assoc UK. 2004;84:887–94.
Weigel BL, Pfister CA. The dynamics and stoichiometry of dissolved organic carbon release by kelp. Ecology. 2021;102:e03221.
Pfister CA, Altabet MA, Weigel BL. Kelp beds and their local effects on seawater chemistry, productivity, and microbial communities. Ecology. 2019;100:e02798.
Schapira M, McQuaid CD, Froneman PW. Free-living and particle-associated prokaryote metabolism in giant kelp forests: Implications for carbon flux in a sub-Antarctic coastal area. Estuar Coast Shelf Sci. 2012;106:69–79.
Newell R, Lucas M, Velirnirov B, Seiderer L. Quantitative significance of dissolved organic losses following fragmentation of kelp (Ecklonia maxima and Laminaria pallida). Mar Ecol Prog Ser. 1980;2:45–59.
Lozada M, Diéguez MC, García PE, Bigatti G, Livore JP, Gil MN, et al. Undaria pinnatifida exudates trigger shifts in seawater chemistry and microbial communities from Atlantic Patagonian coasts. bioRxiv 2020; 2020.10.21.349233.
Kloareg B, Quatrano RS. Structure of the cell walls of marine algae and ecophysiological functions of the matrix polysaccharides. Oceanogr Mar Biol An Annu Rev. 1988;26:259–315.
Gacesa P. Alginates. Carbohydr Polym. 1988;8:161–82.
Rehm BHA. Alginates: biology and applications. Microbiology Monographs. 2009. Springer.
Martin M, Barbeyron T, Martin R, Portetelle D, Michel G, Vandenbol M. The cultivable surface microbiota of the brown alga Ascophyllum nodosum is enriched in macroalgal-polysaccharide-degrading bacteria. Front Microbiol. 2015;6:1–14.
Lin JD, Lemay MA, Parfrey LW. Diverse bacteria utilize alginate within the microbiome of the giant kelp Macrocystis pyrifera. Front Microbiol. 2018;9:1–16.
Sangwan P, Chen X, Hugenholtz P, Janssen PH. Chthoniobacter flavus gen. nov., sp. nov., the first pure-culture representative of subdivision two, Spartobacteria classis nov., of the phylum Verrucomicrobia. Appl Environ Microbiol. 2004;70:5875–81.
Ji S, Wang B, Lu M, Li F. Defluviitalea phaphyphila sp. nov., a novel thermophilic bacterium that degrades brown algae. Appl Environ Microbiol. 2016;82:868–77.
Thomas F, Barbeyron T, Tonon T, Génicot S, Czjzek M, Michel G. Characterization of the first alginolytic operons in a marine bacterium: from their emergence in marine Flavobacteriia to their independent transfers to marine Proteobacteria and human gut Bacteroides. Environ Microbiol. 2012;14:2379–94.
Kabisch A, Otto A, König S, Becher D, Albrecht D, Schüler M, et al. Functional characterization of polysaccharide utilization loci in the marine Bacteroidetes ‘Gramella forsetii’ KT0803. ISME J. 2014;8:1492–502.
Koch H, Freese HM, Hahnke R, Simon M, Wietz M. Adaptations of Alteromonas sp. 76-1 to polysaccharide degradation: A CAZyme plasmid for ulvan degradation and two alginolytic systems. Front Microbiol. 2019;10:504.
Gobet A, Barbeyron T, Matard-Mann M, Magdelenat G, Vallenet D, Duchaud E, et al. Evolutionary evidence of algal polysaccharide degradation acquisition by Pseudoalteromonas carrageenovora 9T to adapt to macroalgal niches. Front Microbiol. 2018;9:1–16.
Dudek M, Dieudonné A, Jouanneau D, Rochat T, Michel G, Sarels B, et al. Regulation of alginate catabolism involves a GntR family repressor in the marine flavobacterium Zobellia galactanivorans DsijT. Nucleic Acids Res. 2020;48:7786–7800.
Koch H, Dürwald A, Schweder T, Noriega-Ortega B, Vidal-Melgosa S, Hehemann JH, et al. Biphasic cellular adaptations and ecological implications of Alteromonas macleodii degrading a mixture of algal polysaccharides. ISME J. 2019;13:92–103.
Thomas F, Bordron P, Eveillard D, Michel G. Gene expression analysis of Zobellia galactanivorans during the degradation of algal polysaccharides reveals both substrate-specific and shared transcriptome-wide responses. Front Microbiol. 2017;8:1808.
Zhu Y, Thomas F, Larocque R, Li N, Duffieux D, Cladière L, et al. Genetic analyses unravel the crucial role of a horizontally acquired alginate lyase for brown algal biomass degradation by Zobellia galactanivorans. Environ Microbiol. 2017;19:2164–81.
Hehemann J-H, Arevalo P, Datta MS, Yu X, Corzett CH, Henschel A, et al. Adaptive radiation by waves of gene transfer leads to fine-scale resource partitioning in marine microbes. Nat Commun. 2016;7:12860.
Enke TN, Datta MS, Schwartzman J, Cermak N, Schmitz D, Barrere J, et al. Modular assembly of polysaccharide-degrading marine microbial communities. Curr Biol. 2019;29:1528–35.
Jain A, Krishnan KP, Begum N, Singh A, Thomas FA, Gopinath A. Response of bacterial communities from Kongsfjorden (Svalbard, Arctic Ocean) to macroalgal polysaccharide amendments. Mar Environ Res. 2020;155:104874.
Mitulla M, Dinasquet J, Guillemette R, Simon M, Azam F, Wietz M. Response of bacterial communities from California coastal waters to alginate particles and an alginolytic Alteromonas macleodii strain. Environ Microbiol. 2016;18:4369–77.
Wietz M, Wemheuer B, Simon H, Giebel H-A, Seibt MA, Daniel R, et al. Bacterial community dynamics during polysaccharide degradation at contrasting sites in the Southern and Atlantic Oceans. Environ Microbiol. 2015;17:3822–31.
Musat N, Foster R, Vagner T, Adam B, Kuypers MMM. Detecting metabolic activities in single cells, with emphasis on nanoSIMS. FEMS Microbiol Rev. 2012;36:486–511.
Reintjes G, Arnosti C, Fuchs BM, Amann R. An alternative polysaccharide uptake mechanism of marine bacteria. ISME J. 2017;11:1640–50.
Liu Y, Fang J, Jia Z, Chen S, Zhang L, Gao W. DNA stable-isotope probing reveals potential key players for microbial decomposition and degradation of diatom-derived marine particulate matter. Microbiologyopen. 2020;9:1–24.
Orsi WD, Smith JM, Liu S, Liu Z, Sakamoto CM, Wilken S, et al. Diverse, uncultivated bacteria and archaea underlying the cycling of dissolved protein in the ocean. ISME J. 2016;10:2158–73.
Cunliffe M, Hollingsworth A, Bain C, Sharma V, Taylor JD. Algal polysaccharide utilisation by saprotrophic planktonic marine fungi. Fungal Ecol. 2017;30:135–8.
Alonso C, Musat N, Adam B, Kuypers M, Amann R. HISH-SIMS analysis of bacterial uptake of algal-derived carbon in the Río de la Plata estuary. Syst Appl Microbiol. 2012;35:541–8.
Arandia-Gorostidi N, Alonso-Sáez L, Stryhanyuk H, Richnow HH, Morán XAG, Musat N. Warming the phycosphere: differential effect of temperature on the use of diatom-derived carbon by two copiotrophic bacterial taxa. Environ Microbiol. 2020;22:1381–96.
Thomas F, Le Duff N, Leroux C, Dartevelle L, Riera P. Isotopic labeling of cultured macroalgae and isolation of 13C-labeled cell wall polysaccharides for trophic investigations. Adv Bot Res. 2020;95:1–17.
Hardouin K, Burlot AS, Umami A, Tanniou A, Stiger-Pouvreau V, Widowati I, et al. Biochemical and antiviral activities of enzymatic hydrolysates from different invasive French seaweeds. J Appl Phycol. 2014;26:1029–42.
Pernthaler A, Pernthaler J, Amann R. Fluorescence in situ hybridization and catalyzed reporter deposition for the identification of marine bacteria. Appl Environ Microbiol. 2002;68:3094–101.
Manz W, Amann R, Ludwig W, Wagner M, Schleifer KH. Phylogenetic oligodeoxynucleotide probes for the major subclasses of Proteobacteria: problems and solutions. Syst Appl Microbiol. 1992;15:593–600.
Manz W, Amann R, Ludwig W, Vancanneyt M, Schleifer KH. Application of a suite of 16S rRNA-specific oligonucleotide probes designed to investigate bacteria of the phylum cytophaga-flavobacter-bacteroides in the natural environment. Microbiology. 1996;142:1097–106.
Wallner G, Amann R, Beisker W. Optimizing fluorescent in situ hybridization with rRNA‐targeted oligonucleotide probes for flow cytometric identification of microorganisms. Cytometry. 1993;14:136–43.
Pernthaler A, Pernthaler J Fluorescene in situ hybridization for the identification of environmental microbes. In: Hilario E, Mackay J (eds). Methods in Molecular Biology. Totowa, NJ: Humana Press Inc.; 2004. pp 153–64.
Guerquin-Kern JL, Wu T Di, Quintana C, Croisy A. Progress in analytical imaging of the cell by dynamic secondary ion mass spectrometry (SIMS microscopy). Biochim Biophys Acta - Gen Subj. 2005;1724:228–38.
Slodzian G, Daigne B, Girard F, Boust F, Hillion F. Scanning secondary ion analytical microscopy with parallel detection. Biol Cell. 1992;74:43–50.
Schneider CA, Rasband WS, Eliceiri KW. NIH Image to ImageJ: 25 years of Image Analysis. Nat Methods. 2012;9:671–5.
Stryhanyuk H, Calabrese F, Kümmel S, Musat F, Richnow HH, Musat N. Calculation of single cell assimilation rates from sip-nanosims-derived isotope ratios: a comprehensive approach. Front Microbiol. 2018;9:1–15.
Woebken D, Burow LC, Behnam F, Mayali X, Schintlmeister A, Fleming ED, et al. Revisiting N2 fixation in Guerrero Negro intertidal microbial mats with a functional single-cell approach. ISME J. 2015;9:485–96.
Ramond P, Sourisseau M, Simon N, Romac S, Schmitt S, Rigaut-Jalabert F, et al. Coupling between taxonomic and functional diversity in protistan coastal communities. Environ Microbiol. 2019;21:730–49.
Thomas F, Corre E, Cébron A. Stable isotope probing and metagenomics highlight the effect of plants on uncultured phenanthrene-degrading bacterial consortium in polluted soil. ISME J. 2019;13:1814–30.
Pepe-Ranney C, Campbell AN, Koechli CN, Berthrong S, Buckley DH. Unearthing the ecology of soil microorganisms using a high resolution DNA-SIP approach to explore cellulose and xylose metabolism in soil. Front Microbiol. 2016;7:1–17.
Buckley DH, Huangyutitham V, Hsu SF, Nelson TA. Stable isotope probing with 15N achieved by disentangling the effects of genome G+C content and isotope enrichment on DNA density. Appl Environ Microbiol. 2007;73:3189–95.
Birnie G Centrifugal separations in molecular and cell biology. Boston: Butterworth & Co Publishers Ltd.; 1978.
Thomas F, Dittami SM, Brunet M, Le Duff N, Tanguy G, Leblanc C, et al. Evaluation of a new primer combination to minimize plastid contamination in 16S rDNA metabarcoding analyses of alga‐associated bacterial communities. Environ Microbiol Rep. 2020;12:30–37.
Klindworth A, Pruesse E, Schweer T, Peplies J, Quast C, Horn M, et al. Evaluation of general 16S ribosomal RNA gene PCR primers for classical and next-generation sequencing-based diversity studies. Nucleic Acids Res. 2013;41:1–11.
Callahan BJ, McMurdie PJ, Rosen MJ, Han AW, Johnson AJA, Holmes SP. DADA2: High-resolution sample inference from Illumina amplicon data. Nat Methods. 2016;13:581–3.
Bolyen E, Rideout JR, Dillon MR, Bokulich NA, Abnet CC, Al-Ghalith GA, et al. Reproducible, interactive, scalable and extensible microbiome data science using QIIME 2. Nat Biotechnol. 2019;37:852–7.
Rognes T, Flouri T, Nichols B, Quince C, Mahé F. VSEARCH: A versatile open source tool for metagenomics. PeerJ. 2016;4:e2584.
Mcmurdie PJ, Holmes S. phyloseq: an R package for reproducible interactive analysis and graphics of microbiome census data. PLoS One. 2013;8:1–11.
Love MI, Huber W, Anders S. Moderated estimation of fold change and dispersion for RNA-seq data with DESeq2. Genome Biol. 2014;15:1–21.
Youngblut ND, Barnett SE, Buckley DH. HTSSIP: an r package for analysis of high throughput sequencing data from nucleic acid stable isotope probing (sip) experiments. PLoS One. 2018;13:1–8.
Youngblut ND, Barnett SE, Buckley DH. SIPSim: a modeling toolkit to predict accuracy and aid design of DNA-SIP experiments. Front Microbiol. 2018;9:570.
Thomas F, Lundqvist LCE, Jam M, Jeudy A, Barbeyron T, Sandström C, et al. Comparative characterization of two marine alginate lyases from Zobellia galactanivorans reveals distinct modes of action and exquisite adaptation to their natural substrate. J Biol Chem. 2013;288:23021–37.
Bankevich A, Nurk S, Antipov D, Gurevich AA, Dvorkin M, Kulikov AS, et al. SPAdes: A new genome assembly algorithm and its applications to single-cell sequencing. J Comput Biol. 2012;19:455–77.
Aziz RK, Bartels D, Best A, DeJongh M, Disz T, Edwards RA, et al. The RAST Server: Rapid annotations using subsystems technology. BMC Genomics. 2008;9:1–15.
Seemann T. Prokka: Rapid prokaryotic genome annotation. Bioinformatics. 2014;30:2068–9.
Zhang H, Yohe T, Huang L, Entwistle S, Wu P, Yang Z, et al. DbCAN2: A meta server for automated carbohydrate-active enzyme annotation. Nucleic Acids Res. 2018;46:W95–W101.
Barrett K, Lange L. Peptide-based functional annotation of carbohydrate-active enzymes by conserved unique peptide patterns (CUPP). Biotechnol Biofuels. 2019;12:1–21.
Almagro Armenteros JJ, Tsirigos KD, Sønderby CK, Petersen TN, Winther O, Brunak S, et al. SignalP 5.0 improves signal peptide predictions using deep neural networks. Nat Biotechnol. 2019;37:420–3.
El-Gebali S, Mistry J, Bateman A, Eddy SR, Luciani A, Potter SC, et al. The Pfam protein families database in 2019. Nucleic Acids Res. 2019;47:D427–D432.
Ferguson RL, Buckley EN, Palumbo AV. Response of marine bacterioplankton to differential filtration and confinement. Appl Environ Microbiol. 1984;47:49–55.
Lucas M, Newell R, Velimirov B. Heterotrophic utilisation of mucilage released during fragmentation of kelp (Ecklonia maxima and Laminaria pallida) II. Differential utilisation of dissolved organic components from kelp mucilage. Mar Ecol Prog Ser. 1981;4:43–55.
Koop K, Newell R, Lucas M. Biodegradation and carbon flow based on kelp (Ecklonia maxima) debris in a sandy beach microcosm. Mar Ecol Prog Ser. 1982;7:315–26.
Barbeyron T, Thomas F, Barbe V, Teeling H, Schenowitz C, Dossat C, et al. Habitat and taxon as driving forces of carbohydrate catabolism in marine heterotrophic bacteria: Example of the model algae-associated bacterium Zobellia galactanivorans DsijT. Environ Microbiol. 2016;18:4610–27.
Díez-Vives C, Nielsen S, Sánchez P, Palenzuela O, Ferrera I, Sebastián M, et al. Delineation of ecologically distinct units of marine Bacteroidetes in the Northwestern Mediterranean Sea. Mol Ecol. 2019;28:2846–59.
Balmonte JP, Buckley A, Hoarfrost A, Ghobrial S, Ziervogel K, Teske A, et al. Community structural differences shape microbial responses to high molecular weight organic matter. Environ Microbiol. 2019;21:557–71.
Alonso-Sáez L, Díaz-Pérez L, Morán XAG. The hidden seasonality of the rare biosphere in coastal marine bacterioplankton. Environ Microbiol. 2015;17:3766–80.
Giovannoni SJ. SAR11 bacteria: the most abundant plankton in the oceans. Ann Rev Mar Sci. 2017;9:231–55.
Grote J, Cameron Thrash J, Huggett MJ, Landry ZC, Carini P, Giovannoni SJ, et al. Streamlining and core genome conservation among highly divergent members of the SAR11 clade. MBio. 2012;3:1–13.
Ngugi DK, Stingl U. High-quality draft single-cell genome sequence of the NS5 marine group from the coastal Red Sea. Genome Announc. 2018;6:5–6.
Woyke T, Xie G, Copeland A, González JM, Han C, Kiss H, et al. Assembling the marine metagenome, one cell at a time. PLoS One. 2009;4:e5299.
Teeling H, Fuchs BM, Bennke CM, Krüger K, Chafee M, Kappelmann L, et al. Recurring patterns in bacterioplankton dynamics during coastal spring algae blooms. Elife. 2016;5:1–29.
Krüger K, Chafee M, Ben Francis T, Glavina del Rio T, Becher D, Schweder T, et al. In marine Bacteroidetes the bulk of glycan degradation during algae blooms is mediated by few clades using a restricted set of genes. ISME J. 2019;13:2800–16.
Pei X, Chang Y. Shen J. Cloning, expression and characterization of an endo-acting bifunctional alginate lyase of marine bacterium Wenyingzhuangia fucanilytica. Protein Expr Purif. 2019;154:44–51.
Xing P, Hahnke RL, Unfried F, Markert S, Huang S, Barbeyron T, et al. Niches of two polysaccharide-degrading Polaribacter isolates from the North Sea during a spring diatom bloom. ISME J. 2015;9:1410–22.
Tanaka R, Shibata T, Miyake H, Mori T, Tamaru Y, Ueda M, et al. Temporal fluctuation in the abundance of alginate-degrading bacteria in the gut of abalone Haliotis gigantea over 1 year. Aquac Res. 2016;47:2899–908.
Bunse C, Koch H, Breider S, Simon M, Wietz M. Sweet and magnetic: succession and CAZyme expression of marine bacterial communities encountering a mix of alginate and pectin particles. bioRxiv 2020; 2020.12.08.416354.
Park HH, Kam N, Lee EY, Kim HS. Cloning and characterization of a novel oligoalginate lyase from a newly isolated bacterium Sphingomonas sp. MJ-3. Mar Biotechnol. 2012;14:189–202.
Sim PF, Furusawa G, Teh AH. Functional and structural studies of a multidomain alginate lyase from Persicobacter sp. CCB-QB2. Sci Rep. 2017;7:1–9.
Lyu Q, Zhang K, Zhu Q, Li Z, Liu Y, Fitzek E, et al. Structural and biochemical characterization of a multidomain alginate lyase reveals a novel role of CBM32 in CAZymes. Biochim Biophys Acta Gen Subj. 2018;1862:1862–9.
Han W, Gu J, Cheng Y, Liu H, Li Y, Li F. Novel alginate lyase (Aly5) from a polysaccharide-degrading marine bacterium, Flammeovirga sp. strain MY04: Effects of module truncation on biochemical characteristics, alginate degradation patterns, and oligosaccharide-yielding properties. Appl Environ Microbiol. 2016;82:364–74.
Kim DH, Wang D, Yun EJ, Kim S, Kim SR, Kim KH. Validation of the metabolic pathway of the alginate-derived monomer in Saccharophagus degradans 2-40T by gas chromatography–mass spectrometry. Process Biochem. 2016;51:1374–9.
Arnosti C, Wietz M, Brinkhoff T, Hehemann J-H, Probandt D, Zeugner L, et al. The biogeochemistry of marine polysaccharides: sources, inventories, and bacterial drivers of the carbohydrate cycle. Ann Rev Mar Sci. 2021;13:9.1–9.28.
Grondin JM, Tamura K, Déjean G, Abbott DW, Brumer H. Polysaccharide Utilization Loci: Fuelling microbial communities. J Bacteriol. 2017;199:e00860–16.
Reintjes G, Arnosti C, Fuchs B. Amann R. Selfish, sharing and scavenging bacteria in the Atlantic Ocean: a biogeographical study of bacterial substrate utilisation. ISME J. 2019;13:1119–32.
Arahal DR, Lucena T, Macián MC, Ruvira MA, González JM, Lekumberri I, et al. Marinomonas blandensis sp. nov., a novel marine gammaproteobacterium. Int J Syst Evol Microbiol. 2016;66:5544–9.
Pontiller B, Martínez-García S, Lundin D, Pinhassi J. Labile dissolved organic matter compound characteristics select for divergence in marine bacterial activity and transcription. Front Microbiol. 2020;11:1–19.
Acknowledgements
We thank the PICT-IBiSA imaging facility at the Institut Curie for the use of the NanoSIMS ion microprobe. We thank the Institut Français de Bioinformatique and the Roscoff Analysis and Bioinformatics for Marine Science platform ABiMS (https://abims.sb-roscoff.fr) for providing computing and data storage resources. We are grateful to Nicolas Gayet (IFREMER Plouzané) for granting access to a sputter coater, and to Diane Jouanneau for help with MALLS analysis. This work benefited from the support of the CNRS via the MITI Isotop project ALGOSIP and of the French government via the National Research Agency programs IDEALG (ANR-10-BTBR-04) and ALGAVOR (ANR-18-CE02-0001). The UMR1136 is supported by the ANR through the Labex Arbre (ANR-11-LABX-0002-01).
Author information
Authors and Affiliations
Contributions
Author contributions following the CRediT taxonomy (https://casrai.org/credit/) are as follows: Conceptualization: FT; Formal analysis: FT, AC, CL, T-DW, J-LG-K; Funding acquisition: FT; Investigation: FT, NLD, PR, AC, CL, GT, EL, SU, T-DW; Project administration: FT; Visualization: FT, T-DW; Writing – original draft: FT; Writing – review and editing: all authors.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Thomas, F., Le Duff, N., Wu, TD. et al. Isotopic tracing reveals single-cell assimilation of a macroalgal polysaccharide by a few marine Flavobacteria and Gammaproteobacteria. ISME J 15, 3062–3075 (2021). https://doi.org/10.1038/s41396-021-00987-x
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41396-021-00987-x
This article is cited by
-
Spatial organization of the kelp microbiome at micron scales
Microbiome (2022)
-
Consuming fresh macroalgae induces specific catabolic pathways, stress reactions and Type IX secretion in marine flavobacterial pioneer degraders
The ISME Journal (2022)
-
Microbial metabolites in the marine carbon cycle
Nature Microbiology (2022)