Abstract
Spatial autocorrelation is inherent to remotely sensed data. Nearby pixels are more similar than distant ones. This property can help to improve the classification performance, by adding spatial or contextual features into the model. However, it can also lead to overestimation of generalisation capabilities, if the spatial dependence between training and test sets is ignored. In this paper, we review existing approaches that deal with spatial autocorrelation for image classification in remote sensing and demonstrate the importance of bias in accuracy metrics when spatial independence between the training and test sets is not respected. We compare three spatial and non-spatial cross-validation strategies at pixel and object levels and study how performances vary at different sample sizes. Experiments based on Sentinel-2 data for mapping two simple forest classes show that spatial leave-one-out cross-validation is the better strategy to provide unbiased estimates of predictive error. Its performance metrics are consistent with the real quality of the resulting map contrary to traditional non-spatial cross-validation that overestimates accuracy. This highlight the need to change practices in classification accuracy assessment. To encourage it we developped Museo ToolBox, an open-source python library that makes spatial cross-validation possible.
Similar content being viewed by others
1 Introduction
The evaluation of classification accuracy has always been considered as an important issue in the remote sensing community (Congalton 1991; Foody 2002, 2008; Ye et al. 2018). There is a large body of literature on this subject with precise recommendations for designing and implementing robust accuracy assessments based on reference data (Olofsson et al. 2014; Stehman and Foody 2019). Significant advances have been made in defining protocols that match the objective of the quality assessment. This may concern the sampling design (stratified or not, systematic or random) (Congalton 1998; Stehman 2009; Ramezan et al. 2019), the size of the sample (Foody 2009; Chen and Wei 2009), the spatial unit of reference samples (pixel, blocks, objects) (Stehman and Wickham 2011), or the accuracy parameters (with their pitfalls) to compute from error matrix (Stehman and Foody 2019; Liu et al. 2007; Pontius and Millones 2011; Foody 2020). Some authors also explored how to account for and represent the spatial distribution of classification errors in order to highlight its non-uniformity and provide additional insights for map users (McIver and Friedl 2001; Foody 2005; Comber et al. 2012; Khatami et al. 2017).
The standard approach used to assess the classification accuracy is to split the reference data into two subsets. The first set is used to train the classification model (learning step). The second set is used to test the model and estimate the prediction errors (validation step). The test set is never used to build the model. This ensures independence between the training and test sets and provides a generally accepted estimate of the predictive power of the model.
Most often, data-splitting is based on simple, possibly stratified, random selection. This selection is sometimes repeated (resampling by bootstrapping) to compute the sampling variability of the accuracy metrics (Lyons et al. 2018). A common alternative to this traditional hold-out validation is assessing the classification accuracy by cross-validation (CV). In this approach, reference data are also split into subsets but the number of subsets can vary (k-fold). The model is trained iteratively on \(k-1\) subset(s) and tested on the remaining set. A accuracy is then measured by averaging the performance values computed on each subset (k models). Leave-one-out (LOO) distribution is a special case of cross-validation where \(k=n\) (n being the size of the reference data set). In this case, each test set is equal to 1 and the model is trained n times.
Despite the importance attached to the accuracy assessment protocol, a gap, sometimes a serious one, is often found between the performance metrics of a model and the real quality of the resulting map. This tends to reduce the confidence placed in accuracy statistics and to discredit the true capacity of remote sensing in the opnion of the t end-users. Among the factors involved in this optimistic bias (Stehman and Foody 2019; Foody 2020), an important one is the spatial dependence between the training and test sets. The spatial context is often ignored in the evaluation even tought it compromises the required independence of the data. Because of spatial autocorrelation, spectral values of close pixels are often more similar than those of distant ones, producing falsely high accuracy metrics if the sampling design is not used for testing (Roberts et al. 2017; Schratz et al. 2019; Meyer et al. 2019). This dependence exists in both the spectral features (i.e. the predictor variables) and the class to predict (i.e. the response variable). A good illustration of this spatial dependence, reported by Inglada (2018), was found in the TiSeLaC land cover classification contest held during the 2017 ECML/PKDD conference. The winner proposed a very simple classification approach to predict the nine most important land cover classes from time-series images. No individual spectral features were used to train the model, but only the pixel geographical coordinates, leading to weighted F-scores ranging from 0.90 to 0.98. The classification was based on k-NN and the validation was carried out by cross-validation Sergey (2017). The runner-up also exploit implicitly exploited the spatial autocorrelation through convolutional neural networks. The accuracy of predictions was estimated at 0.99 of the F-score (Di Mauro et al. 2017).
The existence of spatial autocorrelation is well-known in the remote sensing community (Wulder and Boots 1998). Twenty years ago, Congalton already analysed the pattern of errors found in land-cover classifications and recommended correcting the sampling scheme in the case of non-random distribution (Congalton 1998). However, despite collective awareness, spatial dependence is often ignored in classification accuracy assessment even though a number of approaches have been proposed to prevent it. This leads to systematic overestimation of generalisation capabilities due to spatial overfitting.
The objectives of this paper are thus to (i) review existing approaches that deal with spatial autocorrelation for image classification in remote sensing, (ii) evaluate the impact of spatial autocorrelation between training and test sets on classification performances and (iii) investigate how performances vary with to the data-splitting strategy used for reference samples. The main contribution of this paper is demonstrating the importance of bias in accuracy metrics when spatial independence between the training and test sets is not respected, and hence the need to change accuracy assessment practices. We compare three spatial and non-spatial strategies at pixel and object levels using different sized training samples. The experiments are conducted using a large number of reference samples composed of millions of pixels. Generalization capabilities of the models are evaluated through cross-validation at one site and hold-out validation at two other sites. Our assumption is that predictive power is inflated when the test set is used in the spatial domain of the training set.
2 Related works
One of the challenges of analysing spatial data is dealing with the interdependence between location and the value of the processes to be investigated (Anselin 1989). Spatial processes are distance-related, thus leading to spatial structures in the autocorrelated data. From a statistical point of view, two problems can arise with spatial autocorrelation: (1) spatial non-independence of the classification errors (or model residuals) and (2) spatial non-independence of the training and test sets used for accuracy assessment.
2.1 Spatial autocorrelation in model residuals
The first problem arises when the predictor variables are not able to perfectly account for the effect of the spatial structure to estimate the response variable (Roberts et al. 2017). This is a major concern in certain disciplines like ecology, particularly in biogeographical analyses and species distribution modeling (Roberts et al. 2017; Dormann et al. 2007; Miller et al. 2007; Kühn and Dormann 2012). If parametric models are used to make inferences, the spatial autocorrelation of the prediction errors may lead to erroneous conclusions (Dormann 2007; Kühn 2007). Models are built not only to predict, like in remote sensing, but also to explain the respective effect of each variable. Therefore, if the independence of the residuals (as assumed in standard regression techniques) is violated, estimation of the model parameters may be biased and the type-I error rate (i.e. incorrect rejection of the null hypothesis) may increase (Dormann et al. 2007; Kühn and Dormann 2012). This explains why in spatial ecology, testing spatial independence of residuals is becoming a standard practice.
Several approaches have been proposed to address this issue (Dormann et al. 2007; Miller et al. 2007; Beale et al. 2010). The simplest is incorporating additional predictor variables to improve the model specification. If the spatial pattern of the response variable is fully reflected by the extra autocorrelated predictors, the residuals should not be spatially autocorrelated and a non-spatial model can be used (Kühn and Dormann 2012). Otherwise, spatial models are required. In this case, the spatial dependence is explicitly incorporated in the model, from the response variable itself or from the residuals, like in autocovariate or autoregressive models (Anselin 1988). Spatial eigenvector mapping is an alternative (Dray et al. 2006) like kriging and other geostatistical techniques (see (Dormann et al. 2007; Miller et al. 2007; Beale et al. 2010) for more details).
Spatial models have also been introduced in remote sensing. A widespread approach is the Markov random field (MRF) model that incorporates local information (the neighbors class labels) in the classification process (Solberg et al. 1996; Shekhar et al. 2002; Magnussen et al. 2004). Spatial autologistic regression models have also been considered (Shekhar et al. 2002; Koutsias 2003; Mallinis and Koutsias 2008). However, the most common approach in the field remains the use of non-spatial models. The spatial information is incorporated in contextual classifiers through additional predictors (Wang et al. 2016; Ghamisi et al. 2018). A variety of techniques exist are available including morphological image analysis (Fauvel et al. 2013) and textural analysis (Franklin et al. 2000; Puissant et al. 2005; Sheeren et al. 2009), sometimes based on variograms (Atkinson and Lewis 2000; Berberoglu et al. 2007) or on a local version of principal component analysis (Comber et al. 2016). The adaptation of classifiers has also been proposed using the spatial locations of the training samples (Atkinson 2004) or local spatial patterns (Bai et al. 2020) to estimate class probabilities. Other authors suggest using local spatial statistics (Myint et al. 2007; Ghimire et al. 2010) or interpolated spectral values and their degree of similarity with actual values to improve the classification (Johnson et al. 2012). Today, deep learning has become the most powerful alternative way to incorporate spectral-spatial features for classification (Ghamisi et al. 2018; Zhao and Du 2016). A non-exhaustive summary of these representative approaches is provided in Table 1.
Despite the wide range of technical options, the primary objective of these image processing methods is not to deal with spatial autocorrelation but improve the classification performance. Spatial autocorrelation is considered as an opportunity. However, the effect is the same: the spatial dependence of classification errors is reduced since the model specification is improved. Only a few works have specifically aim to control the spatial dependence between observations in the modeling process (Rocha et al. 2019).
2.2 Spatial autocorrelation between training and test sets
The second issue, which is the focus of this paper, is spatial dependence between the training and test sets. If the classification model is calibrated using training data that are spatially correlated with the data used for testing, predictions based on testing data will be unrealistically high and will not reflect the true predictive power of the model because of spatial overfitting (Roberts et al. 2017; Schratz et al. 2019; Meyer et al. 2019). When the model is intended to classify new sites (assuming stationarity in the relationship between predictor and response variables across space), spatially independent data are required for the estimation of unbiased predictive performance and to test generalization capabilities within the image. While optimistic biases in the predictive power have already been discussed (Chen and Wei 2009; Meyer et al. 2019; Geiß et al. 2017; Hammond and Verbyla 1996; Millard and Richardson 2015), this effect is often disregarded when image classification is evaluated, probably for the sake of simplicty (Ramezan et al. 2019) and because of its importance is underestimated.
One possible way to address this issue is to spatially segregate the training and test sets during the data-splitting procedure. This can be done by imposing a spatial stratification to select the samples, either by objects (Cánovas-García et al. 2017; Inglada et al. 2017), blocks (Lyons et al. 2018; Roberts et al. 2017; Meyer et al. 2019; Valavi et al. 2019), or clusters (Schratz et al. 2019; Brenning 2012) (Table 1). Blocks can be defined arbitrarily (e.g. grid of space) or based on predefined similar characteristics. Clusters can be regular or irregular in shape (depending on the clustering techniques) and can be contiguous or disjointed, with regular or irregular spacing. Another strategy is to define distance-based buffers around the hold-out samples to be sure the learning model is only based on spatially independent data (Le Rest et al. 2014) and disjointed spectral and spatial features (Geiß et al. 2017). The distance (related to the degree of dependency) is often fixed arbitrarily but can also be defined using the correlogram or semivariogram based on the response variable or on the predictors (Valavi et al. 2019; Pohjankukka et al. 2017). The advantage of the latter approach over the other spatial partitioning methods is that the spatial autocorrelation is explicitly quantified. In the other cases, the spatial dependence between training and test sets is assumed to have been removed but this is not checked, meaning residual spatial dependence may persist.
In this paper, we use a distance-based buffer approach that relies on Moran’s I statistics to segregate spatially referenced samples and to explicitly estimate their degree of dependence. To our knowledge, this robust approach based on correlogram has not yet been evaluated on remotely sensed data using a large reference dataset. In a previous work, we applied this buffering strategy to classify tree species in satellite image time series (Karasiak et al. 2019). However, because the reference dataset was small, we were unable to disentangle the effects of spatial autocorrelation and training set size on classification performances.
3 Experimental protocol
3.1 Spatial and non-spatial data-splitting strategies
To assess the impact of spatial autocorrelation between the training and test sets, we defined a supervised classification protocol with three cross-validation (CV) sampling strategies for performance evaluation: (1) a k-fold cross-validation (k-fold-CV) based on random splitting, (2) a non-spatial leave-one-out cross-validation (LOO CV) and (3) a spatial leave-one-out cross-validation (SLOO CV) using a distance-based buffer relying on Moran’s I statistics.
We chose 2 folds (50/50; \(k=2\)) for the k-fold-CV sampling. For the LOO CV and SLOO CV, we systematically selected one sample from each class to test at each iteration (i.e. n\(-1\) per class for training) in contrast to the conventional approach that selects only one sample for testing whatever the class (i.e. no stratification per class). The number of folds is equal to \(n_{min}\), the size of the class with the fewest samples.
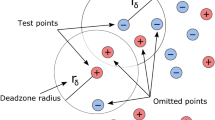
Spatial and non-spatial CV strategies to investigate the effect of spatial autocorrelation between training and test sets. Clusters of points represent objects with their related pixels. At the pixel level (a–c), data-splitting is stratified using the object as a spatial stratum. At the object level (d–f), all the pixels belonging to the objects are either used for training or for testing. Pixels and objects are sampled randomly. For the k-fold CV strategies, \(k=2\) (i.e. split of 50/50%). In the SLOO CV (c, f), a distance-based buffer related to the degree of spatial dependency is defined around the test sample(s). Pixels thar are spatially correlated with the test sample(s) are removed from the training set
Data splitting was performed at both pixel and object levels (Fig. 1). However, at the pixel level, sampling was stratified. We imposed the object as spatial stratum to sample pixels. Thus, for the pixel-based k-fold-CV strategy, after random selection of a fixed number of objects, 50% of pixels belonging to the objects were used for training and the rest for testing (Fig. 1a). Pixels were sampled randomly within the selected objects, with no constraints on the sample size per object. In the object-based k-fold-CV strategy (Fig. 1d), all the pixels belonging to the objects were used for training or testing. This object-based sampling can be viewed as a first option to account for spatial autocorrelation, without quantify it. Test samples are forced to be more spatially distant than training ones. For the remaining LOO CV and SLOO CV strategies, only one pixel (Fig. 1b–c) or object with all its related pixels (Fig. 1e–f) was used to estimate prediction error at each iteration. In SLOO CV, pixels spatially correlated with the single test sample were removed from the training set (Fig. 1c–f). The spatial dependence between nearby pixels was estimated using Moran’s Index (Moran’s I) defined as the ratio of the covariance between neighboring pixels and the variance of all pixels in the image:
where, \(x_i\) is the pixel value of x (a spectral band of the image) at location i, \(x_j\) is the pixel value of x at location j (a nearby pixel of i), \(\overline{x}\) is the average value of x, n is the number of pixels in the image, \(w_{i,j}\) is the weight equals to 1 if pixel j is within distance of d of pixel i, otherwise \(w_{i,j}=0\), and \(S_{0}\) the sum of all \(w_{i,j}\)’s:
More simply, Moran’s I expresses the correlation between the pixel value at one location and its close observations. The index ranges from -1 to +1. For positive values, the nearby pixels covary together (positive spatial autocorrelation). For negative values, the nearby pixels covary in the opposite direction (negative spatial autocorrelation). Values close to 0 indicate the absence of spatial autocorrelation (random spatial distribution).
Moran’s I was computed for each spectral band available in the images we used, from the reference pixels, and for neighborhoods containing from 1 to 2700 pixels. Then, based on correlogram (a plot of Moran’s I as a function of distance), we identified the threshold distance beyond which the spatial dependence between pixels is insignificant. Finally, this threshold was used to define the buffer radius in the SLOO CV strategy. At the pixel level, all the reference samples located in the buffer area centered on the test sample were excluded from the training set (Fig. 1c). At the object level, the exclusion of nearby pixels depends on the position of the centroids of the related objects (Fig. 1f).
Because the effect of spatial autocorrelation may vary with the size of the sample (Chen and Wei 2009), we increased the number of labeled objects (with its related pixels) progressively in the learning experiments, as follows:
-
from 3 to 10 labeled objects: incrementation per 1 object.
-
from 10 to 100 labeled objects: incrementation per 10 objects.
-
from 100 to 1000 labeled objects: incrementation per 100 objects.
The data-splitting procedure was repeated 10 times for all the strategies because we used random undersampling to investigate multiple training set sizes.
3.2 Classification algorithm
We used the nonparametric random forest (RF) learning algorithm to train the models (Breiman 2001). The ability of this algorithm to distinguish land cover classes at large scale despite the limited sensitivity of the parameter values in the classification performances has already been widely demonstrated (Rodriguez-Galiano et al. 2012; Pelletier et al. 2016). RF is also known to be robust to outliers and faster than other classifiers such as the support vector machine algorithm.
In our experiments, we set the number of trees was at 200. The number of variables used to split a node was kept at the default value (\(m = \sqrt{p}\) with p, the total number of features), as well as the stopping criteria for tree building (nodes are expanded until all leaves are pure or contain less than 2 samples). To mitigate the effect of the imbalanced distribution of the classes, we modified class weights in order to prevent bias due to the dominant class. By default, all classes have a weight equal to one. Here, weights were adjusted inversely to the proportion of the class frequency. All the models were fitted with the same hyperparameters. Spectral features were also standardized (i.e. centering and scaling to unit variance) prior to training.
To evaluate the classification performances, we deliberately abandoned the kappa index (Foody 2020). We focused our attention on producer’s accuracy per class (also known as specificity and sensitivity with two classes), in addition to overall accuracy (OA). User’s accuracy was not considered since the classifications were not intended to be used. We only report the OA as accuracy estimator in the results section. However, to control for a possible interpretation bias of OA, which is prevalence-dependent (i.e. the most prevalent class dominates the indicator value), the class-specific performances are also provided in the appendix. The average value based on 10 repetitions was computed with standard deviation as confidence interval.
We used the scikit-learn python library for implementation (Pedregosa et al. 2011). The spatial and non-spatial sampling strategies were performed using Museo ToolBox, a python library we developed to make this research reproducible (Karasiak 2020).
3.3 Image dataset
We used Sentinel-2 (S2) optical images to test the experimental protocol and in particular, the 31TDJ S2 tile (100 km x 100 km in extent). This tile is located in the South of France and partially covers four French administrative departments, and includes the city of Albi (Fig. 2). In practice, only three of departments were classified: Herault-34, Tarn-81 and Aveyron-12. The effect of spatial autocorrelation was investigated by mapping two simple forest classes (coniferous and broadleaf) that can include different tree species.
The dataset includes four S2 images acquired on July 17, August 8, September 5 and October 15, 2016. We did not select images covering all four seasons, but gave priority to images with less than 5% cloud covers. The image acquisition dates are not ideal to distinguish coniferous and broadleaf stands. However, we focused on the relative differences in accuracy between the data-splitting strategies rather than on the ability to properly distinguish the two forest classes. The S2 data were downloaded from the French national THEIA platform at level 2A (i.e. top-of-canopy reflectances corrected for atmospheric and topographic effects (Hagolle et al. 2015)). We only used a subset of the available spectral bands: Blue (B2 - 490nm), Green (B3 - 560nm), Red (B4 - 665nm) and Near Infra-Red (B8 - 842nm) at 10-m spatial resolution. The cloudy pixels in the dataset were corrected using a gap-filling approach (linear interpolation) and a mask of clouds produced using the MAJA pre-processing chain of the S2 time series (Baetens et al. 2019).
3.4 Reference dataset
The reference dataset (labeled pixels) for the two classes of forest was derived from the French National Forest Inventory spatial database (IGN BDForet®, v.2). This source provides a vector map of forest stands with a minimum area of 0.5 hectares. The composition of each stand is obtained from aerial stereo-image interpretation completed by field surveys. A stand is considered as pure if the proportion of the area covered by one species equals or exceeds 75% of the total area, otherwise, it is defined as a mixed stand. Here, we selected pure stands of coniferous and broadleaf species for the experiments. All mixed stands were excluded. Because of a temporal gap between the S2 images (2016) and the reference forest maps (dating from 2006 to 2015 depending on the department), we updated the forest maps to eliminate forest stands whose land cover had changed. This was done by masking S2 pixels with NDVI values lower than 0.4 using the image of July 17. We also eroded the vector forest layer with an inside buffer of 20 m to avoid selecting pixels of mixed classes at the boundaries of the masks. Finally, forest stands covering less than one hectare were removed to insure we kept a dense reference dataset that would enable us to explore different sampling strategies. The total number of forest stands and related pixels are given in Table 2. The forest stand is the spatial stratum used to sample pixels in pixel-based strategies and was the sample unit for the object-based ones.
3.5 Experimental setup
The six cross-validation strategies with different sample sizes were applied to the department of Herault-34. The maximum sample size was set at 1,972 labeled objects corresponding to the number of coniferous stands in this site. However, it was not possible to investigate all the sample sizes for the pixel-based LOO CV and SLOO CV strategies. The maximum number of sampled objects was limited to 10 (i.e. \(\approx\) 32,000 pixels for broadleaf spp and 5,476 for conifer sp, on average) due to long computation time. Because the number of iterations increases with the sample size in cross-validation, the time required to compute the distance matrix among pixels in Moran’s I rapidly becomes prohibitive.
We used reference samples in the neighboring departments, Tarn-81 and Aveyron-12, as additional independent datasets to validate the full model of Herault (i.e. the model with the maximum training set size of 1972 objects per class). This extra validation on spatially distant sites was carried performed after removing all reference samples in Tarn-81 and Aveyron-12 that were spatially correlated with the Herault-34 ones. We also removed some reference samples from the over-represented class of broadleaf spp through random under-sampling (30 repetitions). In this way, the accuracy metrics in the extra validation were computed with the same number of samples per class (369,164 pixels) in both departments Tarn-81 and Aveyron-12.
Combining the three CV strategies at both pixel and object levels for the different sized samples, means a total of 1240 classifications were trained and tested on Herault-34 department (i.e. 27 sets of samples from 3 to 1972 stands x 2 level of analysis x 3 CV strategies x 10 repetitions, including exceptions in sample size for the pixel-based LOO and SLOO CV). Full spatial independence between the training and test sets was ensured in two configurations of the protocol: using the SLOO CV strategy on Herault-34, and during validation on Tarn-81 and Aveyron-12. The pixel-based k-fold and LOO CV are the sampling strategies that should be most affected by spatial dependence. The object-based k-fold and LOO CV are in an intermediate position.
4 Results
4.1 Prediction errors according to wether spatial/non-spatial cross-validation strategies we used
The classification performances obtained by cross-validation on Herault-34 are given in Fig. 3. Overall, the results show that ignoring dependence between training and test sets leads to very high accuracy metrics whatever the sample size. This is particularly clear at the pixel level but also at the object level with large samples. Compared to sampling strategies that account for spatial autocorrelation, the accuracy metrics are overestimated.
Average overall accuracy based on the RF classifier for each cross-validation strategy (k-fold CV, LOO CV, SLOO CV) at pixel and object levels. Models were fitted with reference samples of Herault-34 and repeated 10 times (i.e. the y-axis provides the average OA value ± standard deviation). The premature stopping of the pixel-based LOO and SLOO CV approaches was due to excessive computational time
At the pixel level (see dotted lines with a cross in Fig. 3), the performances of k-fold CV and LOO CV sampling strategies (green and blue lines respectively) were close with very high average OA values, regardless of the size of the training set (e.g. OA of 97.86 ± 0.96% and 95.81 ± 3.71% respectively for 10 forest stands). We observed a gradual decrease in OA for k-fold CV with the increasing number of forest stands. With the pixel-based SLOO CV approach (orange line), the prediction errors were much higher than other pixel-based methods. Concerning the group-based SLOO CV the OA started with very low values (average OA of 68.72 ± 11.03%) and then increased to reach relatively stable accuracy from 50 forest stands on (average OA of 86.08 ± 5.73%).
At the object level, the general pattern of errors differed from the non-spatial pixel-based strategies (see dashed lines with the dots in Fig. 3). Starting with a very weak performance, an improvement of OA began with the increase if the size of the training set size. The accuracy of k-fold CV and LOO CV sampling strategies (green and blue lines respectively) followed the same trend. The SLOO CV approach showed a similar pattern but differed in the magnitude and variability of prediction errors. In addition, the average OA remained relatively stable from 50 forest stands on, suggesting that the maximum performance has been reached for this sied training set. In contrast, the average OA of the object-based k-fold CV and LOO CV approaches continued to grow with the increase in the size of the training set, revealing the effect of spatial dependence with large datasets (with closer objects). With the maximum sample size (1,972 forest stands), the difference in OA between SLOO CV and LOO CV was substantial (from 82.08% to 90.76% respectively) indicating an optmistic bias in the non-spatial LOO CV as well as in the k-fold CV (OA = 93.73 ± 0.13%). The differences in performance between the object-based SLOO CV and the k-fold CV and LOO CV strategies were minimal between 50 and 100 forest stands.
Compared to pixel-based CV, the performance accuracy of the object-based CV was less affected by spatial dependence (overestimation is reduced) and revealed the expected pattern with an increase in the training set. In the case of the SLOO strategy, both pixel and object levels follow the same pattern with higher OA values for the pixel-based strategy.
Object-based sampling was found to be particularly impacted by the size of the sample. Model performance was very poor when the models were trained with only a few forest stands. A higher number of pixels for training can be obtained by increasing the number of forest objects, with increased representativeness of the coniferous and broadleaf classes composed of variety of species (by reducing the sample selection bias). Since forest stands have varying extents (and are consequently composed of a diffeent number of pixels due to their irregular shapes), different training sets that are similar in size may lead to variable prediction performances, as illustrated in Fig. 4. Further, between two training sets composed of the same number of pixels, the one based on more forest stands provides higher prediction accuracy. In other words, performances increase with sample size if the number of objects also increases (together with the number of pixels). Variability is particularly apparent with sample from 10\(^3\) to 10\(^4\) pixels (Fig. 4). Beyond 50 forest stands (i.e. >10\(^4\) pixels), the variability gradually disappears with the k-fold CV and LOO CV strategies and to a lesser extent, with the SLOO CV approach.
An analysis was conducted of the number of species included in the coniferous and broadleaf classes according to the number of forest stands. The maximum number of dominant broadleavf species existing in Herault-34 (i.e. existing in pure forest stands as defined in the IGN BDForet® reference database) was systematically reached with 30 forest stands. Five dominant tree species were found in this class. For conifers, more forest stands of pure species exist with a less even distribution both in membership and space. Ten dominant conifer tree species were found. Eight of them were systematically sampled in 50 forest stands. Beyond 50, the number of average sampled species oscillated continuously between 8 and 10 up to 1000 forest stands. Thus, because of larger number and unequal frequency of species composing the conifer class, and the lower number of available conifer samples compared to broadleaf samples, the error rate for conifers was higher (see producer’s accuracy metric in Fig. 12). This was particularly clear when accuracy was assessed with the SLOO CV approach but less apparent when non-spatial strategies were used.
For the spatial CV, the distance threshold with a negligible spatial dependence between all the reference data was estimated at 19.74 km based on Moran’s I correlogram (see Fig. 9). This threshold value was used in the pixel and object SLOO CV strategies.
4.2 Prediction errors on other spatially independent and distant sites
Prediction errors were also estimated for the neighboring departments (Tarn-81 and Aveyron-12) from the Herault-34 full model, i.e. the model fitted using all the available training data (1972 forest stands with related pixels). We assumed there was no change in the dataset between Herault-34 and the neighboring departments since the dominant composition of species in the coniferous and broadleaf classes of Tarn-81 and Aveyron-12 is included in Herault-34, according to the IGN BDForet® reference database (and confirmed by the relative frequency distribution of NDVI values of each class; see Appendices 10 and 11).
Interestingly, we found OA performed similarly in predictions for Tarn-81 (80.4 ± 0.03%), Aveyron-12 (82.2 ± 0.01%) and the comparable object-based SLOO CV strategy of Herault-34 based on 1972 forest stands (82.1 ± : 0.03%). Whether by cross-validation or by validation based on distant sites, predictions made outside the spatial domain of training set produced equivalent error rates (Fig. 5) The confusion matrices are provided in the Appendices 13.
When spatial dependence between training and test sets is ignored using standard CV, it tends to overestimate the accuracy (from +8% to +14% of OA according to the strategy compared to the OA values of distant sites).
Comparison between the average predictive performances obtained with spatial and non-spatial cross-validation from the Herault-34 full model fitted with the maximum training set of 1972 forest stands (on the right) and the average predictive performances obtained with the same model applied to test sets from Tarn-81 and Aveyron-12 (in black, on the left). The number of test samples in Tarn-81 and Aveyron-12 is exactly the same with balanced class distributions. Pixel-based LOO CV and SLOO CV are not shown because of the excessive computation time required for this big training set
Additional predictions were estimated for distant sites, based on Herault-34 full models with different sized training sets. We compared the OA values of these models evaluated using the spatial and non-spatial cross-validation strategies with the OA values based on predictions made with these models in Tarn-81 (Fig. 6) and Aveyron-12 (Fig. 7). These results show that the OA values are consistent among themselves. For an OA value estimated for a distant site, we observed a marked variability in OA estimated by cross-validation, with an optimistic bias with the non-spatial CV strategy. On average, the OA estimated from SLOO CV matches the OA estimated from spatially distant sites better, especially for large samples size (more than 50 stands). The true predictive performances of the Herault-34 full models of each strategy do not differ fundamentally, for a given training set size. Apparent differences in performance are only due to differences in the way they are assessed using CV strategies. This explains why, with large training sets size (more than 500 forest stands), the predictive performance for distant sites are all equivalent (see the x-axis values of the purple dots in Figs. 6 and 7; OA is approximately to 80% with 1,972 stands).
Comparison of OA estimated by cross-validation in Herault-34 using the six spatial and non spatial data-splitting strategies for different sized training sets and OA estimated with the same Herault-34 model on test sets in Tarn-81. The number of training set sizes is given by the number of forest stands represented by the graphical color variable. Results for pixel-based LOO CV and SLOO CV are not shown for more than 10 forest stands because of excessive computation time (Color figure online)
Comparison of OA estimated by cross-validation in Herault-34 from the six spatial and non spatial data-splitting strategies for different sized training sets and OA estimated using the same Herault-34 model on test sets in Aveyron-12. The number of training set sizes is given by the number of forest stands represented through the graphical color variable. Results for pixel-based LOO CV and SLOO CV are not shown for more than 10 forest stands because of excessive computation time (Color figure online)
5 Discussion
Our results revealed notable underestimation of generalization errors when traditional non-spatial approaches were used to assess the accuracy. Pixel-based samplings were the most affected. Object-based strategies mitigate the effect of spatial dependence since the pixels used for training and testing never belong to the same forest stands. Nonetheless, non-spatial data-splitting at the object level also leads to overestimation of predictive performance.
Three distinct performance trends were observed with the increase in the size of the training set: (1) a slight gradual decrease in the non-spatial pixel-based CV, (2) a marked and continuous increase in the non-spatial object-based CV, and (3) a marked increase up to an optimal training set size in SLOO CV with no improvement beyond. To our knowledge, these contrasting patterns have never previously been demonstrated.
Learning curves can help interpret these trends. The curves describe how the model’s error rates on training and test sets vary as the training set size increases. This makes it possible to diagnose model bias and variance. Theoretically, with only a few training samples, the model error rate on the training set would be expected to be very low (if not zero). The model can fit the training samples perfectly but does not have the ability to generalize new data. Thus, the model error rate on the test set is expected to be high. With larger datasets, the error on the training set should increase because of less overfitting. Conversely, the model should perform better on the test set. The generalization capabilities should be improved, reducing the error rate on the test set with more data but keeping an irreducible error. This analysis was conducted using three contrasted CV strategies: the pixel-based k-fold CV, the object-based LOO CV, and the object-based SLOO CV.
However, the expected behavior of learning curves was not observed with the pixel-based k-fold CV most affected by spatial autocorrelation. With both small and large training sets, the error rate in training set was zero with a convergence of the learning curve on the test set from the first sample sizes onwards (Fig. 8). The narrow gap between the two curves falsely suggests a model with low variance irrespective of the size of the training set. As it also shows limited bias, the model appears to be perfect with no difficulty expected with generalization. This unrealistic behavior is mainly due to spatial dependence. The pixels in the test sets belong to the same forest stands as those in the training sets. The model consequently overfits the training samples but overfitting is masked by the overly optimistic estimates of accuracy using similar and correlated test sets. The generalization capabilities of the model are better with large samples. The spatial autocorrelation between training and test sets does not change, but the model slightly reduces overfitting on test sets, thereby increasing the error rate.
A more realistic pattern of errors was found with the object-based SLOO CV that was not affected by spatial autocorrelation. Notable model variance was observed when the training set size was small but variance tended to be reduced by adding more training samples, as expected (Fig. 8). The learning curves of the training and test converged from 50 forest stands on, suggesting that adding more samples beyond this threshold is not necessary, but there is no substantial improvement. This sample size is to be related to the number of species sampled in the coniferous class which affect the classification accuracy, as previously mentionned (see Sect. 4.1). Beyond 50 forest stands, eight out of the ten dominant coniferous species are sampled systematically, reducing confusion with broadleaf species.
The behavior of the non-spatial object-based LOO CV is a blend of the two previous trends. When the training sets are small, accuracy is less affected by spatial autocorrelation. Compared to pixel-based CV, the spatial dependence between training and test sets is lower because of object-based sampling. In addition, the generalization capabilities of the model are limited because of the high variability of species composition between the training and test sets. This leads to considerable model variance. With the addition of new samples, the model performs better on test sets but with progressive overestimation of performance. The increase in performance is not due to the improvement of generalization capabilities of the model. With reference to the SLOO CV, the predictive power of the model might be reached beyond approximately 50 forest stands. Large sample sizes tend to reduce the distance between forest stands thereby increasing the effect of spatial dependence between training and test sets and hence, estimated accuracy. This explains why the error rates between non-spatial pixel-based and object-based CV are similar with large samples.
With all these strategies, whatever the sample size, the average zero error rate in the training set was unexpected, suggesting constant model overfitting with a reduction in the error rate in the test sets (in particular, in object-based CVs). Since, to avoid overfitting, we included a large number of trees in the RF algorithm, we attribute this deceptive behavior to spatial autocorrelation too but in this case, between the samples used in the training set. And this is another fundamental point that should be taken into consideration. All spatial and non-spatial CV strategies guarantee spatial independence among training samples. Spatial data splitting removed autocorrelation between the training and test sets but this autocorrelation persisted in the training set. Thus, learning curves need to be interpreted with caution.
Our findings support evidence from previous studies that spatial CV is required to estimate unbiased predictive error (Schratz et al. 2019; Meyer et al. 2019). Our results also show that the non-spatial object-based sampling is less affected by spatial autocorrelation than pixel-based sampling, in line with Cánovas-García et al. (2017). When an optimal sample size is used for training (i.e. with no undersampling or oversampling) this strategy is a possible alternative to the SLOO CV if spatial dependence is ignored in the accuracy assessment. However, this non-spatial strategy only mitigates overfitting, it does not prevent it. Thus, we recommend to choosing a spatial CV whenever possible.
We used a distance-based buffer approach for spatial CV (Le Rest et al. 2014) instead of block or cluster-based partitioning (Roberts et al. 2017; Schratz et al. 2019; Meyer et al. 2019). Data-splitting with blocks also produces better estimates of predictive performances than random sampling. However, with no explicit quantification and no control of the degree of dependence, residual correlation may exist between training and test sets when the spatial block approach is used (i.e. where blocks consist of geographical units). Objects belonging to contiguous blocks and located close to borders may be very almost identical. Finding the ideal block size does not solve the problem, but the distance-based buffer approach avoids it. Systematic quantification of the spatial autocorrelation range should also be performed using Moran’s I or empirical variogram.
6 Conclusion
The take-home message of this paper is that we need to change practices in classification accuracy assessment using spatial imagery. A data splitting design ensuring spatial independence between the training and test sets should be the standard approach for validation. Non-spatial LOO or k-fold CV at the object level (i.e. cross-validation leaving all the pixels belonging to one object for testing and the rest for training) is a absolute minimum required to mitigate overfitting. Spatial LOO CV is a better way to provide unbiased estimates of predictive error and to reduce the gap between the accuracy statistics given to users and the real quality of the maps produced.
To facilitate this change, we assembled the Museo ToolBox, an open-source python library that makes it possible to validate classification results with a range of spatial and non-spatial CV approaches (https://museotoolbox.readthedocs.io/). We hope this library will help the community leave its traditional approach behind to instead to routinely use spatial CV. We also expect the library to open the way for new prospects on this topic.
References
Anselin, L. (1988). Spatial econometrics: Methods and models. Dordrecht: Kluwer Academic Publishers.
Anselin, L. (1989). What is special about spatial data: Alternative perspectives on spatial data analysis. Technical Paper 89-4, National Center for Geographic Information and Analysis, Santa Barbara, CA : NCGIA.
Atkinson, P., & Lewis, P. (2000). Geostatistical classification for remote sensing: An introduction. Computers & Geosciences, 26(4), 361–371.
Atkinson, P. M. (2004). Spatially weighted supervised classification for remote sensing. International Journal of Applied Earth Observation and Geoinformation, 5(4), 277–291.
Baetens, L., Desjardins, C., & Hagolle, O. (2019). Validation of copernicus sentinel-2 cloud masks obtained from maja, sen2cor, and fmask processors using reference cloud masks generated with a supervised active learning procedure. Remote Sensing, 11(4).
Bai, H., Cao, F., Atkinson, M. P., Chen, Q., Wang, J., & Ge, Y. (2020). Incorporating spatial association into statistical classifiers: local pattern-based prior tuning. International Journal of Geographical Information Science, 1–38.
Beale, C., Lennon, J., Yearsley, J., Brewer, M., & Elston, D. (2010). Regression analysis of spatial data. Ecology Letters, 13(2), 246–264.
Berberoglu, S., Curran, P., Lloyd, C., & Atkinson, P. (2007). Texture classification of Mediterranean land cover. International Journal of Applied Earth Observation and Geoinformation, 9(3), 322–334.
Breiman, L. (2001). Random forests. Machine Learning, 45(1), 5–32.
Brenning, A. (2012). Spatial cross-validation and bootstrap for the assessment of prediction rules in remote sensing: The R package sperrorest. In 2012 IEEE international geoscience and remote sensing symposium, pp 5372–5375.
Chen, D., & Wei, H. (2009). The effect of spatial autocorrelation and class proportion on the accuracy measures from different sampling designs. ISPRS Journal of Photogrammetry and Remote Sensing, 64(2), 140–150.
Comber, A., Fisher, P., Brunsdon, C., & Khmag, A. (2012). Spatial analysis of remote sensing image classification accuracy. Remote Sensing of Environment, 127, 237–246.
Comber, A. J., Harris, P., & Tsutsumida, N. (2016). Improving land cover classification using input variables derived from a geographically weighted principal components analysis. ISPRS Journal of Photogrammetry and Remote Sensing, 119, 347–360.
Congalton, R. (1991). A review of assessing the accuracy of classifications of remotely sensed data. Remote Sensing of Environment, 37(1), 35–46.
Congalton, R. (1998). A comparison of sampling schemes used in generating error matrices for assessing the accuracy of maps generated from remotely sensed data. Photogrammetric Engineering & Remote Sensing, 54(5), 593–600.
Cánovas-García, F., Alonso-Sarría, F., Gomariz-Castillo, F., & Oñate-Valdivieso, F. (2017). Modification of the random forest algorithm to avoid statistical dependence problems when classifying remote sensing imagery. Computers & Geosciences, 103, 1–11.
Di Mauro, N., Vergari, A., Basile, T., Ventola, F., and Esposito, F. (2017). End-to-end learning of deep spatio-temporal representations for satellite image time series classification. In 2017 ECML/PKDD Discovery Challenges.
Dormann, C. (2007). Effects of incorporating spatial autocorrelation into the analysis of species distribution data. Global Ecology and Biogeography, 16(2), 129–138.
Dray, S., Legendre, P., & Peres-Neto, P. (2006). Spatial modelling: A comprehensive framework for principal coordinate analysis of neighbour matrices (pcnm). Ecological Modelling, 196(3), 483–493.
F. Dormann, C., M. McPherson, J., B. Araújo, M., Bivand, R., Bolliger, J., Carl, G., G. Davies, R., Hirzel, A., Jetz, W., Daniel Kissling, W., Kühn, I., Ohlemüller, R., R. Peres-Neto, P., Reineking, B., Schröder, B., M. Schurr, F., and Wilson, R. . (2007). Methods to account for spatial autocorrelation in the analysis of species distributional data: a review. Ecography, 30(5), 609–628.
Fauvel, M., Tarabalka, Y., Benediktsson, J. A., Chanussot, J., & Tilton, J. C. (2013). Advances in spectral-spatial classification of hyperspectral images. Proceedings of the IEEE, 101(3), 652–675.
Foody, G. (2005). Local characterization of thematic classification accuracy through spatially constrained confusion matrices. International Journal of Remote Sensing, 26(6), 1217–1228.
Foody, G. (2008). Harshness in image classification accuracy assessment. International Journal of Remote Sensing, 29(11), 3137–3158.
Foody, G. (2009). Sample size determination for image classification accuracy assessment and comparison. International Journal of Remote Sensing, 30(20), 5273–5291.
Foody, G. M. (2002). Status of land cover classification accuracy assessment. Remote Sensing of Environment, 80(1), 185–201.
Foody, G. M. (2020). Explaining the unsuitability of the kappa coefficient in the assessment and comparison of the accuracy of thematic maps obtained by image classification. Remote Sensing of Environment, 239, 111630.
Franklin, S., Hall, R., Moskal, L., Maudie, A., & Lavigne, M. (2000). Incorporating texture into classification of forest species composition from airborne multispectral images. International Journal of Remote Sensing, 21(1), 61–79.
Geiß, C., Aravena Pelizari, P., Schrade, H., Brenning, A., & Taubenböck, H. (2017). On the effect of spatially non-disjoint training and test samples on estimated model generalization capabilities in supervised classification with spatial features. IEEE Geoscience and Remote Sensing Letters, 14(11), 2008–2012.
Ghamisi, P., Maggiori, E., Li, S., Souza, R., Tarablaka, Y., Moser, G., et al. (2018). New frontiers in spectral-spatial hyperspectral image classification: The latest advances based on mathematical morphology, markov random fields, segmentation, sparse representation, and deep learning. IEEE Geoscience and Remote Sensing Magazine, 6(3), 10–43.
Ghimire, B., Rogan, J., & Miller, J. (2010). Contextual land-cover classification: Incorporating spatial dependence in land-cover classification models using random forests and the getis statistic. Remote Sensing Letters, 1(1), 45–54.
Hagolle, O., Huc, M., Pascual, D., & Dedieu, G. (2015). A multi-temporal and multi-spectral method to estimate aerosol optical thickness over land, for the atmospheric correction of formosat-2, landsat, vens and sentinel-2 images. Remote Sensing, 7(3), 2668–2691.
Hammond, T. O., & Verbyla, D. L. (1996). Optimistic bias in classification accuracy assessment. International Journal of Remote Sensing, 17(6), 1261–1266.
Inglada, J. (2018). Machine learning for land cover map production - Follow-up on the TiSeLaC challenge.
Inglada, J., Vincent, A., Arias, M., Tardy, B., Morin, D., & Rodes, I. (2017). Operational high resolution land cover map production at the country scale using satellite image time series. Remote Sensing, 9(1), 95.
Johnson, B., Tateishi, R., & Xie, Z. (2012). Using geographically weighted variables for image classification. Remote Sensing Letters, 3(6), 491–499.
Karasiak, N. (2020). Museo toolbox: A python library for remote sensing including a new way to handle rasters. Journal of Open Source Software, 5(48), 1978.
Karasiak, N., Dejoux, J.-F., Fauvel, M., Willm, J., Monteil, C., & Sheeren, D. (2019). Statistical stability and spatial instability in mapping forest tree species by comparing 9 years of satellite image time series. Remote Sensing, 11(21), 2512.
Khatami, R., Mountrakis, G., & Stehman, S. (2017). Mapping per-pixel predicted accuracy of classified remote sensing images. Remote Sensing of Environment, 191, 156–167.
Koutsias, N. (2003). An autologistic regression model for increasing the accuracy of burned surface mapping using landsat thematic mapper data. International Journal of Remote Sensing, 24(10), 2199–2204.
Kühn, I. (2007). Incorporating spatial autocorrelation may invert observed patterns. Diversity and Distributions, 13(1), 66–69.
Kühn, I., & Dormann, C. (2012). Less than eight (and a half) misconceptions of spatial analysis. Journal of Biogeography, 39(5), 995–998.
Le Rest, K., Pinaud, D., Monestiez, P., Chadoeuf, J., & Bretagnolle, V. (2014). Spatial leave-one-out cross-validation for variable selection in the presence of spatial autocorrelation. Global Ecology and Biogeography, 23(7), 811–820.
Liu, C., Frazier, P., & Kumar, L. (2007). Comparative assessment of the measures of thematic classification accuracy. Remote Sensing of Environment, 107(4), 606–616.
Lyons, M. B., Keith, D. A., Phinn, S. R., Mason, T. J., & Elith, J. (2018). A comparison of resampling methods for remote sensing classification and accuracy assessment. Remote Sensing of Environment, 208, 145–153.
Magnussen, S., Boudewyn, P., & Wulder, M. (2004). Contextual classification of landsat tm images to forest inventory cover types. International Journal of Remote Sensing, 25(12), 2421–2440.
Mallinis, G., & Koutsias, N. (2008). Spectral and spatial-based classification for broad-scale land cover mapping based on logistic regression. Sensors, 8(12), 8067–8085.
McIver, D. K., & Friedl, M. A. (2001). Estimating pixel-scale land cover classification confidence using nonparametric machine learning methods. IEEE Transactions on Geoscience and Remote Sensing, 39(9), 1959–1968.
Meyer, H., Reudenbach, C., Wöllauer, S., & Nauss, T. (2019). Importance of spatial predictor variable selection in machine learning applications - moving from data reproduction to spatial prediction. Ecological Modelling, 411, 108815.
Millard, K., & Richardson, M. (2015). On the importance of training data sample selection in random forest image classification: A case study in peatland ecosystem mapping. Remote Sensing, 7(7), 8489–8515.
Miller, J., Franklin, J., & Aspinall, R. (2007). Incorporating spatial dependence in predictive vegetation models. Ecological Modelling, 202(3–4), 225–242.
Myint, S. W., Wentz, E. A., & Purkis, S. J. (2007). Employing spatial metrics in urban land-use/land-cover mapping. Photogrammetric Engineering & Remote Sensing, 73(12), 1403–1415.
Olofsson, P., Foody, G., Herold, M., Stehman, S., Woodcock, C., & Wulder, M. (2014). Good practices for estimating area and assessing accuracy of land change. Remote Sensing of Environment, 148, 42–57.
Pedregosa, F., Varoquaux, G., Gramfort, A., Michel, V., Thirion, B., Grisel, O., Blondel, M., Prettenhofer, P., Weiss, R., Dubourg, V., et al. (2011). Scikit-learn: Machine learning in python. Journal of Machine Learning Research, 12(Oct):2825–2830.
Pelletier, C., Valero, S., Inglada, J., Champion, N., & Dedieu, G. (2016). Assessing the robustness of Random Forests to map land cover with high resolution satellite image time series over large areas. Remote Sensing of Environment, 187, 156–168.
Pohjankukka, J., Pahikkala, T., Nevalainen, P., & Heikkonen, J. (2017). Estimating the prediction performance of spatial models via spatial k-fold cross validation. International Journal of Geographical Information Science, 1–19.
Pontius, R., & Millones, M. (2011). Death to kappa: birth of quantity disagreement and allocation disagreement for accuracy assessment. International Journal of Remote Sensing, 32(15), 4407–4429.
Puissant, A., Hirsch, J., & Weber, C. (2005). The utility of texture analysis to improve per-pixel classification for high to very high spatial resolution imagery. International Journal of Remote Sensing, 26(4), 733–745.
Ramezan, A., Warner, A., & Maxwell, A. (2019). Evaluation of Sampling and Cross-Validation Tuning Strategies for Regional-Scale Machine Learning Classification. Remote Sensing, 11(2).
Roberts, D. R., Bahn, V., Ciuti, S., Boyce, M. S., Elith, J., Guillera-Arroita, G., et al. (2017). Cross-validation strategies for data with temporal, spatial, hierarchical, or phylogenetic structure. Ecography, 40(8), 913–929.
Rocha, A. D., Groen, T. A., & Skidmore, A. K. (2019). Spatially-explicit modelling with support of hyperspectral data can improve prediction of plant traits. Remote Sensing of Environment, 231, 111200.
Rodriguez-Galiano, V., Ghimire, B., Rogan, J., Chica-Olmo, M., & Rigol-Sanchez, J. (2012). An assessment of the effectiveness of a random forest classifier for land-cover classification. ISPRS Journal of Photogrammetry and Remote Sensing, 67, 93–104.
Schratz, P., Muenchow, J., Iturritxa, E., Richter, J., & Brenning, A. (2019). Hyperparameter tuning and performance assessment of statistical and machine-learning algorithms using spatial data. Ecological Modelling, 406, 109–120.
Sergey, R. (2017). Temporal and spatial approaches for land cover classification. In 2017 ECML/PKDD Discovery Challenges.
Sheeren, D., Bastin, N., Ouin, A., Ladet, S., Balent, G., & Lacombe, J.-P. (2009). Discriminating small wooded elements in rural landscape from aerial photography: a hybrid pixel/object-based analysis approach. International Journal of Remote Sensing, 30(19), 4979–4990.
Shekhar, S., Schrater, P. R., Vatsavai, R. R., Weili, Wu., & Chawla, S. (2002). Spatial contextual classification and prediction models for mining geospatial data. IEEE Transactions on Multimedia, 4(2), 174–188.
Solberg, A. H. S., Taxt, T., & Jain, A. K. (1996). A markov random field model for classification of multisource satellite imagery. IEEE Transactions on Geoscience and Remote Sensing, 34(1), 100–113.
Stehman, S. (2009). Sampling designs for accuracy assessment of land cover. International Journal of Remote Sensing, 30(20), 5243–5272.
Stehman, S., & Wickham, J. (2011). Pixels, blocks of pixels, and polygons: Choosing a spatial unit for thematic accuracy assessment. Remote Sensing of Environment, 115(12), 3044–3055.
Stehman, S. V., & Foody, G. M. (2019). Key issues in rigorous accuracy assessment of land cover products. Remote Sensing of Environment, 231, 111199.
Valavi, R., Elith, J., Lahoz-Monfort, J. J., & Guillera-Arroita, G. (2019). blockcv: An r package for generating spatially or environmentally separated folds for k-fold cross-validation of species distribution models. Methods in Ecology and Evolution, 10(2), 225–232.
Wang, L., Shi, C., Diao, C., Ji, W., & Yin, D. (2016). A survey of methods incorporating spatial information in image classification and spectral unmixing. International Journal of Remote Sensing, 37(16), 3870–3910.
Wulder, M., & Boots, B. (1998). Local spatial autocorrelation characteristics of remotely sensed imagery assessed with the getis statistic. International Journal of Remote Sensing, 19(11), 2223–2231.
Ye, S., Pontius, R., & Rakshit, R. (2018). A review of accuracy assessment for object-based image analysis: From per-pixel to per-polygon approaches. ISPRS Journal of Photogrammetry and Remote Sensing, 141, 137–147.
Zhao, W., & Du, S. (2016). Spectral-spatial feature extraction for hyperspectral image classification: A dimension reduction and deep learning approach. IEEE Transactions on Geoscience and Remote Sensing, 54(8), 4544–4554.
Acknowledgements
N.K. received a PhD scholarship from the French Ministry of Higher Education and Research (University of Toulouse). The authors are grateful to the European Space Agency (ESA) and the French national THEIA infrastructure for access to Copernicus Sentinel data.
Author information
Authors and Affiliations
Contributions
Conceptualization, N.K. and D.S.; Methodology, N.K. and D.S.; Investigation, N.K. and D.S.; Software, N.K.; Visualization, N.K.; Validation, N.K. and D.S.; Funding acquisition, D.S.; Supervision, J.-F.D., C.M. and D.S.; Writing–original draft preparation, N.K. and D.S.; Writing–review and editing, J.-F.D. and C.M.
Corresponding author
Ethics declarations
Conflicts of interest
The authors declare no conflict of interest.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Editors: Dino Ienco, Thomas Corpetti, Minh-Tan Pham, Sebastien Lefevre, Roberto Interdonato.
Appendices
Moran’s I correlogram on Sentinel-2 data
See Fig. 9.
Moran’s I correlograms of each Sentinel-2 spectral band, for pixels representing forests. Each curve represents one spectral band at one date in the dataset. For a Moran’s I threshold value of 0.05, spatial independence between nearby pixels was assumed. This threshold was reached at 1,964 pixels (i.e. 19.64 km) on average. The grey dashed line represents the mean distance value (in x) where Moran’s I = 0.05 (in y)
Relative frequency distribution of NDVI values per class
Class-specific performances and class prevalence
See Fig. 12.
Confusion matrices
See Fig. 13.
Confusion matrix of the test set in Aveyron-12 (left) and in Tarn-81 (right) from the Herault-34 full model fitted with the maximum training set size of 1972 forest stands. The average accuracy metrics have been estimated from 369,164 reference pixels for each class after undersampling repeated 30 times. The values in the cells are in pixels. The units of Producer and User’s accuracy are in %
Rights and permissions
About this article
Cite this article
Karasiak, N., Dejoux, JF., Monteil, C. et al. Spatial dependence between training and test sets: another pitfall of classification accuracy assessment in remote sensing. Mach Learn 111, 2715–2740 (2022). https://doi.org/10.1007/s10994-021-05972-1
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10994-021-05972-1