Abstract
Spartium junceum L. is a typical species of Mediterranean shrubland areas, also grown in gardens and parks as an ornamental. In recent years in Europe, S. junceum has been recurrently found to be infected by different subspecies and genotypes of the quarantine regulated bacterium Xylella fastidiosa (Xf). This work presents for the first time the anatomy of S. junceum plants that we found, by means of genetic and immunochemistry analysis, to be naturally infected by Xf subsp. multiplex ST87 (XfmST87) in Monte Argentario (Grosseto, Tuscany, Italy), a new outbreak area within the EU. Our anatomical observations showed that bacteria colonized exclusively the xylem conductive elements and moved horizontally to adjacent vessels through pits. Interestingly, a pink/violet matrix was observed with Toluidine blue staining in infected conduits indicating a high content of acidic polysaccharides. In particular, when this pink-staining matrix was observed, bacterial cells were either absent or degenerated, suggesting that the matrix was produced by the host plant as a defense response against bacterial spread. In addition, a blue-staining phenolic material was found in the vessels and, at high concentration, in the pits and inter-vessels. SEM micrographs confirmed that polysaccharide and phenolic components showed different structures, which appear to be related to two different morphologies: fibrillary and granular, respectively. Moreover, our LM observations revealed bacterial infection in xylem conductive elements of green shoots and leaves only, and not in those of other plant organs such as roots and flowers.
Similar content being viewed by others
Introduction
Xylella fastidiosa (Wells et al. 1987) (Xf) is a Gram-negative bacterium, which presents six subspecies: fastidiosa, multiplex, sandyi, morus, tashke, and pauca (EFSA 2020). It is the causal agent of Pierce’s disease in grapevine, the olive quick decline syndrome in olive, the citrus variegated chlorosis in citrus plants, and leaf scorch disease in almond, coffee, oleander, elm, sycamore, pecan, pear, mulberry, maple, and oak (Hopkins and Purcell 2002; Janse and Obradovic 2010; Purcell 2013). Xf is always associated with its plant host or its insect vectors, which are sharpshooter leafhoppers (Hemiptera: Cicadellidae: Cicadellinae), spittlebugs (Hemiptera: Cercopidae) (Severin 1949; Severin 1950), and cicadas (Hemiptera: Cicadidae) (Paião et al. 2002); no evidence has been reported of Xf free-living outside its hosts. In plants, Xf lives only in xylem vessels while in the hemipteran insects, it colonizes the cuticular surface of the oral cavity or rather the anterior foregut (Backus and Morgan 2011). The similarity between the two exclusive niches of Xf (i.e., host plant and insect vector) is that in both cases, the colonized tissues are composed of non-living cells with which the bacterium mainly interacts during its life cycle. This phenomenon is probably due to the Xf genome adaptation. The genome of Xf shares some similarities with that of xanthomonads, typical bacterial pathogens, but in contrast, it has a much-reduced genome and lacks a Type III secretion system (T3SS) (Simpson et al. 2000). T3SS are complex bacterial structures that are delivered in living tissues to suppress the immune response of the host; the lack of this system may explain why Xf survives only when it is surrounded by dead cells (Roper et al. 2019).
It is well established that after colonizing xylem conduits, Xf leads to occlusion of the vascular system due to gums, tyloses, and bacterial biofilms (Sun et al. 2013; Pèrez-Donoso et al. 2016). Depending on the extent of the occlusion, vascular plugs cause partial or total arrest of water flow from the roots to the leaves. Thus, typical symptoms of the disease are leaf scorching at the early stages followed by withering of shoots until plant death, which happens in a few years (Saponari et al. 2013).
The presence of Xf has been documented in several plant species of commercial interest especially in Italy, where it caused the death of thousands of centenarian olive trees (Cariddi et al. 2014). However, many aspects of the disease still need to be clarified. Although the transmission of Xf through the hemiptera insects is well documented, the mechanism of pathogenesis is still under investigation due to the complexity of the infection process (Cornara et al. 2016; Novelli et al. 2019).
Among the several diseases caused by the different subspecies of Xf, the present work is focused on the infection of Spartium junceum L. (Fabaceae) (Spanish broom) by X. fastidiosa subsp. multiplex (Xfm). Although Xfm is thought to be native of North America, this subspecies was first detected in 1935 in Argentina causing plum leaf scald and then in 1978 in Paraguay and Brazil (Nunes et al. 2003). More recently, 41 species of angiosperms have been found susceptible to Xfm in Europe (EU 2019; Landa et al. 2020).
S. junceum is an indigenous species of the Mediterranean basin and it plays an important role in the coastal Tuscan landscape. Because of its marked adaptability and high resistance to drought, this species has also been successfully used to protect and colonize slopes that were subject to superficial erosion phenomena (Preti and Giadrossich 2009). Historically, the plant was cultivated and used as a raw textile material to make ropes spun for vessels and shipbuilding, as well as nets, bags, and sails (Katović et al. 2012). Recently, a medieval shipwreck excavation found galleys from the Byzantine period in which treenails were mainly constructed with Spanish broom wood (Akkemik and Kocaba 2012). S. junceum is also known for its medicinal properties due to anti-neoplastic activity (Abusamra et al. 2015), and as an ornamental and also poisonous plant due to the presence of quinolizidine alkaloids (Giménez et al. 2017).
During the execution of an early detection surveillance program for Xf, the DNA of Xfm was first detected in a plant of S. junceum growing in the municipality of Monte Argentario (Grosseto, Tuscany, Italy) in October 2018 (Marchi et al. 2018). Soon after the presence of its DNA was detected in some other hosts, the bacterium was isolated, a novel sequence type (ST87) was identified, and three genomes were deposited in the GenBank database (Saponari et al. 2019; Giampetruzzi et al. 2019).
The present research aimed to describe the anatomy of S. junceum plants naturally infected by Xfm ST87, highlighting pathological characteristics and possible diagnostic anatomical features. For this purpose, we investigated the interaction of Xfm ST87 with S. junceum tissues and the mechanism of bacterial spread through the plant xylem vessels until the formation of vascular plugs. Additionally, we investigated the nature of the gel matrix associated with the pathogen in xylem vessels, which could be linked to the complex mechanism of plant defense response to this parasite.
Materials and methods
Plant sampling and indexing for Xf
After the first detection of Xfm ST87 in October 2018, demarcated areas were established according to EU Decision 2015/789, and monitoring, plant sampling, whole nucleic acid extractions, and molecular indexing for Xf were carried out according to PM 7/24-3 unless otherwise stated (EPPO 2018). Briefly, during inspection procedures, specimens of S. junceum plants were collected and selected mostly based on the presence of scorching of green shoots, a nonspecific symptom putatively ascribable also to Xf. Samples were immediately stored at 5 °C for up to 2 days prior to processing. Whole nucleic acids were extracted from green shoots using the CTAB-based protocol and stored at − 20 °C until further use. Initial screening for the presence of Xf was carried out by at least two independent real-time (qPCR) protocols (Harper et al. 2010; Francis et al. 2006; Ouyang et al. 2013). A selection of samples found to be infected by Xf according to qPCR results were further subject to amplification and sequencing of either the sole nuoL gene fragment or all the 7 genes (cysG, gltT, holC, malF, leuA, nuoL, and petC) required by the MLST-typing approach (Yuan et al. 2010). For qPCR experiments, the Quantinova Probe PCR kit (Qiagen, Hilden, Germany) and Quantinova SYBR Green PCR Kit (Qiagen, Hilden, Germany) master mixes were used; meanwhile for conventional PCR, the GoTaq G2 polymerase (Promega) was used. All conventional PCR products were visualized after electrophoresis in 1.5% agarose (Genaxxon) gels in 1 × Tris–acetate–EDTA buffer (Invitrogen) and staining with Midori Green (Nippongenetics; 0.05 μl mL−1). They were purified using FastAP and Exonuclease I (Thermo) and Sanger sequenced. Sense and antisense nucleotide sequences were visualized and checked for quality using CHROMAS LITE 2.01 (Technelysium), aligned using MUSCLE as implemented in MEGA6 (Tamura et al. 2013), and single consensus sequences were determined. Searches were carried out on the Xf MLST database website (http://pubmlst.org/xfastidiosa) to determine the allele and the resulting sequence type (ST).
X. fastidiosa isolation and characterization
Xf isolation was performed at different time points throughout 2019 from plants previously indexed for the presence of Xf (see above). Green shoots were cut to a length of approximately 10 cm with flame-disinfected shears, surface disinfected by immersion in 80% ethanol followed by flaming, and finally cut into fragments of approximately 3 cm in length inside a laminar flow hood. Occasionally, bacterial isolation from different plant parts was also tested. Isolations were attempted on BCYE agar as described in Bleve et al. (2016) using 40 fragments from each plant and by delivering a single blot of sap/fragment into the isolation plate (8 fragments/plate). Plates were incubated at 28 °C and checked every 2 days during the first week of incubation and once a week thereafter for a total of 8 weeks. Plates containing bacterial colonies that become visible to the unaided eye within the first 72 h of incubation were discarded; meanwhile, those that became visible thereafter were streaked twice for purity on BCYE agar, suspended in sterile distilled water, and identified as Xf based on the results of qPCR (Harper et al. 2010) performed on 1 μl of bacterial suspension after boiling for 10 min. The DNA of two Xf isolates from each plant was extracted using CTAB buffer, resuspended in Tris–HCl 10 mM pH8, and further characterized to subspecies and ST level using the MLST approach using the conditions described by Yuan et al. (2010). QuantiFast Multiplex PCR Master Mix (Qiagen) and GoTaq G2 polymerase (Promega) were used for qPCR and conventional PCR, respectively. qPCR conditions and MLST gene fragments amplification, purification, and analysis were performed as described above. All isolation procedures were carried out in the laboratory of the Phytosanitary Service of the Tuscany Region in Porto Santo Stefano (Grosseto, Italy).
Spartium junceum sampling
Primary roots, branches, apical, medial, and basal portions of green shoots, leaves, and flowers were collected from three different S. junceum plants (replicates), hereafter referred to as SJ 668, SJ 911, and SJ 796, from which Xfm ST87 had been previously isolated. SJ 668 and SJ 911 plants showed scattered desiccation of shoots while SJ 796 appeared to be asymptomatic. Roots, twigs, and green shoots were collected from SJ 668 in October 2019, before its eradication according to EU Decision 2015/789, and from SJ 911 in February 2020. True leaves, green shoots, and flowers were collected from SJ 796 in May 2020. A fourth S. junceum plant, SJ 777, that had tested negative to qPCR indexing for Xf and from which the bacterium could not be isolated on BCYE, was collected in the same area in May 2020 and used as negative control. Finally, a healthy S. junceum plant was collected in an area close to Ginestra Fiorentina in the province of Florence (Tuscany, Italy), approximately 160 km away from the infected area, where there is no evidence of Xf presence, and used as naïve control.
All plant material used in this work was homogeneous in terms of region of origin, environmental conditions of growth, and age (3–4 years).
Light, immunohistochemistry, and scanning electron microscopy
For light microscopy (LM) observations, samples were fixed with FAA (5:5:90 v/v/v 40% formaldehyde: glacial acetic acid: 70% ethanol) at 5 °C and then dehydrated in ethanol series (70%, 80%, 95%, and 100% v/v). These samples were then embedded in Technovit 7100 resin and sectioned with a Reichert-Jung Ultracut E microtome. Primary shoots (approximately 0.5 cm in diameter), older branches, leaves, flowers, and roots were sectioned with a microtome to obtain 2 μm sections. Transverse or longitudinal sections were mounted on microscope slides and stained using different protocols: 0.5% Toluidine blue O in 0.1% Na2CO3 at pH 11.1 (O’Brian and McCully 1981), which at this pH, stains acidic polysaccharides pink/violet and phenolic components blue (Dong et al. 1997; Tagne et al. 2002; Crews et al. 2003); Alcian Blue for acid mucopolysaccarides; and Periodic acid-Schiff staining (PAS) reaction for polysaccharides. A Leitz DM-RB Fluo Optic microscope (Wetzler, Germany) equipped with a digital camera Nikon DS-L1 (Tokyo, Japan) was used for microscopic observations.
Transversal and longitudinal sections 2-μm-thick samples sections, embedded in Technovit 7100 resin, as well as 20-μm-thick sections fixed in FAA and maintained in 50% ethanol, were used for immunofluorescence determination. In order to remove the resin and retrieve the antigen, the embedded sections were treated with 10% H202 or a saturated NaOH solution diluted 1:2 in ethanol for 15′ and then washed in distilled water. For the immunohistochemical assay, the Immunofluorescence kit IF 07419 by Loewe® Biochemica GmbH (Sauerlach, Germany) was used. After a wash in PBS, the primary antibody against Xf diluted 1:1000 in PBS was applied to each slide incubated for 30 min in a humid chamber at room temperature and then washed twice for 7 min with 0.1% Tween 20 in PBS. Finally, FITC-conjugated goat anti-rabbit secondary antibody diluted 1:150 in PBS was applied and incubated for 30 min in a humid chamber at room temperature, After incubation, the slides were rinsed as previously described and mounted with Fluoroshield with DAPI (F6057, Merck, Germany). Slide observations were made using a Nikon inverted microscope (Nikon Eclipse Ti) and the following epifluorescence filter sets were used: excitation 365 nm, emission 400 nm hi-pass to observe DAPI staining; excitation 485 nm, emission 524 nm for FITC fluorescence staining, and excitation 575 nm, emission 661 nm for red fluorescence (lignin fluorescence). Images were acquired with the Coolsnap HQ2 CCD camera (Princeton instruments, USA).
For scanning electron microscopy (SEM) studies, samples were fixed with FAA and then processed with the immersion for 24 h each in a series of increasing concentrations of ethanol (70%, 80%, 95%, and 100% v/v). After dehydration, samples were critical point dried and gold metalized using a sputter coater. Finally, samples were observed by SEM Zeiss EVO 40 at MEMA center of the University of Florence (Centro di Servizi di Microscopia Elettronica e Microanalisi—MEMA). The instrument was set at 15 kV and at different working distances ranging from 6.0 to 5.5 mm.
Results and discussion
X. fastidiosa incidence in S. junceum, isolation, and characterization
Between October 2018 and December 2019, a total of 669 S. junceum plants were sampled as part of the Xf monitoring program in the demarcated area of Monte Argentario (Grosseto, Tuscany, Italy). According to the results of two independent qPCR specific assays, 106 plants were found to be infected by Xf. To confirm these results, a selection of 70 Xf-positive extracts were subject to amplification of a fragment of the Xf nuoL gene. Fifty-five of them yielded an amplicon of the expected size (557 bp) and 51 out of 55 were successfully sequenced in both forward and reverse. Sequence analysis against the Xf MLST database indicated that allele 21 of the Xfm ST87 nuoL gene had been sequenced in all samples. MLST analysis was completed on 19 of the 51 S. junceum extracts, and the presence of the sole ST 87 [5 (leuA), 3 (petC), 5 (malF), 3 (cysG), 3 (holC), 21 (nuoL), 3 (gltT)] was found.
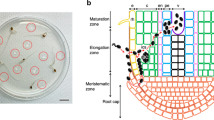
Bacterial isolation was attempted from 11 S. junceum plants arbitrarily selected among those found to be infected according to qPCR and MLST results. Fast growing, often pigmented bacterial colonies usually appeared within the sap blotted areas on BCYE agar during the first 7 days of incubation at 28 °C. Their incidence was low, ranging between 0 and 5% of the analyzed plant fragments, and their DNA could not be amplified with the Xf-specific qPCR protocol (Harper et al. 2010). White, punctiform colonies became visible to the unaided eye usually after 2–3 weeks of incubation (Fig. 1a).
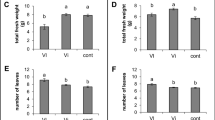
After the second round of purification, single colonies on BCYE agar were of an opalescent white color, mucoid when touched with a loop, slightly convex, and circular with entire margins with a diameter of 1 mm after 30 days of incubation at 28 °C (Fig. 1b). Nearly all isolates (110 out of 116) displaying this colony morphotype tested positive for Xf according to Harper et al. (2010). Based on these results, Xf was successfully isolated from 6 out of 11 plants of S. junceum, each sampled in a different location of Monte Argentario (Fig. 2).
The frequency of Xf isolation from fragments of these six plants ranged from 22 to 75% (Fig. 3). Two isolates from each plant (12 isolates in total) were arbitrarily selected and were confirmed to belong to ST87 [5 (leuA), 3 (petC), 5 (malF), 3 (cysG), 3 (holC), 21 (nuoL), 3 (gltT)] by MLST-typing analysis.
Absolute frequencies of isolation of Xf from 6 plants of S. junceum. From each plant, 40 green shoot fragments were chosen, squeezed with sterile pliers and the sap from one of the exposed ends blotted onto BCYE agar plates. After 8 weeks of incubation at 28 °C, the number of sap blots from which Xf was successfully isolated based on the results of qPCR (Harper et al. 2010) was counted
Based on these findings, there is circumstantial evidence that Xfm ST87 is the most recurrent if not sole X. fastidiosa phylotype in Monte Argentario.
Immunohistochemistry analysis
The presence of Xfm ST87 in infected plants was confirmed by immunohistochemical analysis. Bacterial cells were observed in several xylem vessels (Fig. 4a), close to the cambium tissue in cross and longitudinal sections where bacteria fully colonized lignified elements (Fig. 4b). In healthy plants, the presence of the bacterium in the xylem vessels of primary shoots was never detected in either the transverse (Fig. 4c) or longitudinal section (Fig. 4d).
Fluorescence images of cross (a, c) and longitudinal (b, d) sections of primary shoot of SJ 668 and control plant after immunohistochemical staining. a Bacteria localized in xylem vessels close to the cambium (white arrows); b bacteria localized in lignified elements (white arrow); c and d negative control
Anatomical observations
LM images captured from cross and longitudinal sections of green shoots of SJ 668 samples are shown in Fig. 5. Xfm ST87 was observed in a group of xylem vessels by the previous cribro-vascular bundle (Fig. 5a). In particular, Fig. 5b shows the presence of several neighbor vessels colonized by bacteria (indicated by red arrows). It seems that bacteria horizontally infected adjacent vessels moving through the pits that connect the whole vascular system as previously observed in Olea europea L. petioles colonized by X. fastidiosa subsp. pauca ST53 (Cardinale et al. 2018; Novelli et al. 2019). This diffusion mechanism was also confirmed by the SEM micrograph shown in Fig. 5c where bacteria were found crossing the pits. Another observation is that bacteria were never found in the living cells of cambium or phloem tissues, supporting the hypothesis that Xfm ST87 largely survives surrounded by dead cells throughout its life. This phenomenon was also observed in green shoots of SJ 911 (Fig. SI 1 A-B) and SJ 796 (Fig. SI 2 E).
LM images of transversal (a, b) and longitudinal (d, e) sections of young shoots of SJ 668 stained with Toluidine blue. SEM images (c, f) of longitudinal sections of young shoots of SJ 668. a Presence of a bacterial colony (surrounded by red circle) close to a vascular bundle. b Magnified detail of image (a): several cells with lignified walls are occupied by bacterial colonies (red arrows). c Bacteria crossing the pits of the adjacent vessel. d Healthy shoot tissue (negative control) where the phloem (PH), cambium (C), xylem (X), and pith (P) are visible. e Vessel infected by Xfm ST87. f SEM image of the longitudinal section of a young shoot showing bacteria filling a vessel
In S. junceum twigs, healthy tissues showed key features of the primary structure as shown in Fig. 5d, where no bacterial cells were observed in xylem conductive elements. Other examples of healthy organs (i.e., the ovary, the twig, and the leaf) of S.junceum, in particular of SJ 777, are shown in Fig. SI 2 (A-D). Interestingly, bacteria occupying the conductive element were found arranged parallel to each other following the watercourse, until they totally occluded the vessel (Fig. 5e). This was confirmed in a SEM micrograph where a vessel with spiral wall lignification was obstructed by the high density of bacteria in the lumen (Fig. 5f).
In Fig. 6a, the longitudinal section of older shoots shows intermediate infection stages up to vascular plugging. At the early stage of colonization, bacteria interacted with the xylem wall probably using adhesins, which have been reported to be produced by Xf in large amounts in comparison to other bacteria (Simpson et al. 2000; Van Sluys et al. 2003). Then, bacterial cells were joined by more cells and the diffusion process initiated, first colonizing the single vessel element through its perforation plates and, subsequently spreading in the proximity, digesting the pit membranes of adjacent xylem cells (Newman et al. 2003; Roper et al. 2019). It seems that the interaction between Xfm ST87 and the host is not univocal. Once in the xylem vessels, Xfm ST87 cells adhere to the plant cell wall as well as to each other, coordinating the exploratory movement along the conduits to enhance disease progression, similarly to what is reported by Novelli et al. (2019) for Xfp ST53 in olive. At the same time, the host plant recognizes an exogenous presence and responds to the pathogen with physical and chemical post-infection barriers. In our opinion, this plant defense response is shown in Fig. 6b where the presence of bacteria is associated with a pink/red stained matrix which is probably related to gels (gum) secretion by the parenchyma cells (Rioux et al. 1998). The bacteria appear trapped/concentrated in the matrix, where they self-replicated filling at a later stage the entire vessels (about 450 μm long), thus occluding the water passage (Fig. 6c).
LM images obtained from longitudinal (a, c, d) and cross (b) sections of a twig of SJ 668 stained with Toluidine blue. SEM images (e, f) of longitudinal sections of young shoots of SJ 668. a Vessels of young shoot infected by Xfm ST87 at an early stage. Bacterial colonies (asterisks) occupy part of the vessel lumen crossing perforation plates (black arrows). b Cross-section showing bacteria in the lumen of several vessels colonized by bacteria (black arrows), the presence of pink/violet stained mucilage in tracheary elements (black stars) in one of which degenerated bacteria are observed, and a blue-stained matrix (white star). c Vessels infected by Xfm ST87 at a later stage. Bacterial colonies (asterisks) completely occupy some of the vessels’ lumen. d A conductive element with two stainings: blue and pink. A darker blue staining was observed in the pits (red arrows) compared to the pits of unaffected vessels (black arrows). Moreover, white arrows indicate damaged or dead bacterial cells in the pink matrix. e Negative control of pits. f SEM micrograph showing a granular matrix corresponding to the blue gel occluding the pits as shown in Fig. 5d
The chemical nature of the gels inside the xylem vessels was heterogeneous. The pink/violet Toluidine blue staining (identified in Fig. 6b by black stars) would indicate the presence of gels (mucilage) in the vessels, as previously observed using specific dyes and thorough extractions (Dong et al. 1997; Tagne et al. 2002; Crews et al. 2003), indicating a high content of acidic polysaccharides. The pink-staining matrix was observed when bacteria were absent or dead, suggesting that it was produced by S. junceum as a defense response against their spreading (Fig. 6b). Amounts of pink-staining material with Toluidine blue were found in cotyledons of Brassica napus infected by Leptosphaeria maculans (Roussel et al. 1999). In addition to the pink-staining matrix, blue-staining phenolic materials were also observed in maize vessels following mechanical wounding, as reported by Crews et al. (2003). The two different color reactions were also observed in SJ 911 green shoot sections (Fig. SI 1 C and D). Furthermore, in Fig. 6d, a particular vessel element showed both colors, blue and pink/violet. The dark blue-stained mucilage was also found at higher concentrations in the pits and inter-vessels compared to those of conduits noninfected conduits (black arrows). We observed that bacterial cells found in the presence of the pink-stained region appeared degenerated compared to bacteria shown in Fig. 5c. For this reason, we hypothesized that they might be damaged or dead (shown by white arrows). In comparison with the negative control shown in Fig. 6e, the blue gel can be associated with a granular matrix that apparently caused the formation of pit plugs, as shown in Fig. 6f. Since the xylem vessel is not able to produce mucilage, the surrounding parenchyma cells must be involved in the task. The blockade of the pits by granular plugs may reduce the capability of the bacteria to move from a xylem column to the adjacent ones.
The nature of the gel composition was clarified using two different stainings, Alcian Blue (Fig. 7a and b) and PAS reaction (Fig. 7c and d), which are specific for acidic mucopolysaccharides and polysaccharides, respectively.
LM images of SJ 668 samples. a Transverse sections of a twig region stained with Alcian Blue for acidic mucopolysaccharides in the pits (black arrows) and in the vessels lumen (asterisks). b Detail of (a) in which the presence of mucopolysaccharides (asterisks) is found. c Transverse section of a twig stained with PAS reaction for polysaccharides. d Detail of (c) in which bacteria are found in the presence of polysaccharides. e Transverse section of the older branch stained with Toluidine blue. Bacteria are indicated by red arrows, polysaccharide and phenolic components by black arrows. f Longitudinal section of a green shoot stained with Toluidine blue. Bacteria are apparently covered by a blue layer (arrows)
Then, the presence of Xfm ST87 was investigated in older branches (Fig. 7e) in order to localize the bacteria in the different growth rings. From the transverse section, it was possible to date the age of the shoot, which was approximately 3 years. Bacteria were found in the cell lumen of the younger growth ring but not observed in the oldest. The vessel elements in the oldest growth ring were characterized mainly by the presence of polysaccharide and phenolic matrices without any bacteria, once again confirming that the presence of polysaccharides and phenolic gels in the lumen are not always associated with bacteria.
In some vessels, bacteria were coated by a thin blue layer and the blue matrix corresponding to the phenolic component was observed around few bacteria (Fig. 7f).
Bacteria were observed in SJ 796’s leaf sections (Fig. SI 1 E) while flower sections of SJ 796 and root sections of SJ 668 did not show any bacterial colonization (Fig. SI 1 F, 1 G, and 1 H).
The results presented in this paper indicate that a single phylotype of Xf subsp. multiplex (Xfm), i.e., ST87, is infecting S. junceum plants growing in the demarcated area of Monte Argentario (Grosseto, Tuscany, Italy). This finding, along with the fact that ST87 has been previously reported from other hosts growing in the same area and that draft genomes of three ST87 strains have an average nucleotide identity of 99.99%, brings compelling evidence of a widely spread phylotype, if not sole, of Xf at Monte Argentario (Marchi et al. 2018; Saponari et al. 2019; Giampetruzzi et al. 2019).
After the first record in the European Union territory on olive (Saponari et al. 2013), the presence of Xf subspecies pauca, multiplex, and fastidiosa was recorded in cultivated as well as in spontaneous Mediterranean plant species growing in natural and urban landscapes of Italy, France, Spain, and Portugal. Some of these plants were previously unreported hosts of these Xf subspecies, and many of them were found to be infected by different STs of different Xf subspecies in different countries and locations (EPPO 2020). Apart from underlining once more the ability of Xf to become rapidly adapted to new environments, the European scenario also indicates the existence of hosts whose potential role as inoculum sources in current and future Xf outbreaks needs to be rapidly addressed. Some of them are common plants of commerce, as in the case of the ornamental plants Polygala myrtifolia and Lavandula dentata, or of fruit trees like Prunus dulcis (EC, 2019). Others, as for example Calicotome spinosa, are present in shrubland areas throughout the Mediterranean basin. Finally, plants like Rosmarinus officinalis and Cistus albidus as well as S. junceum itself not only grow spontaneously but are also cultivated as ornamentals in gardens and parks. Despite these findings, to the best of our knowledge, the anatomy of olive plants naturally infected by Xf subsp. pauca ST 53 is the only one that has been studied and characterized in Europe to date (De Benedictis et al. 2017; Cardinale et al. 2018; Sabella et al. 2018; Sabella et al. 2019).
As a typical Mediterranean species, S. junceum presents morphological features, such as ephemeral leaves and green-photosynthetic stems that make it perfectly adapted to a xerophytic environment (Bezić et al. 2003; Nilsen et al. 1993). Hence, the anatomy of young shoots is characterized by photosynthetic cortical parenchyma with crystals, intercalated by groups of sclerenchyma fibers below a multi-layered epidermis with a thick cuticle. The secondary xylem presents distinct growth ring boundaries with a porous or semi-porous wood, vessels in a dendritic pattern, and the axial parenchyma represented by paratracheal parenchyma. It is well known that the possible plant defense responses against pathogens are at the anatomical level, where some specific features such as gels or tyloses allow to compartmentalize the spread of the pathogen. Studies on anatomical plant responses induced by pathogens have been published since the 1980s. In particular, when studying interactions between tomato and Verticillum albo-atrum, Street et al. (1986) concluded that the pathogen could induce the production of two distinct coating forms that infused pit membranes and primary walls or lined the secondary walls of xylem vessels. Working with woody plants native to Switzerland, Bonsen and Bonsen and Kučera (1990) observed a correlation between the minimum pit aperture diameter (3 μm) of a vessel-parenchyma pit pair and the type of vessel occlusion by tyloses or gums: below that minimum, gum production occurred, while above 3 μm, tyloses were observed. In our LM observations, we found a narrower pit aperture diameter (< 3 μm) which may justify the presence of gels (gums) in infected tracheary elements. Despite the presence of a paratracheal parenchyma, tyloses such as those observed in naturally infected plants of olive and Vitis vinifera (Cardinale et al. 2018; Roper et al. 2019) were never found in any of the infected S. junceum samples. Artificial infection with Xf induced tyloses in V. vinifera and V. smalliana but not in V. arizonica/candican, which produced fibrillary networks and gum occlusions (Fritschi et al. 2008). In our LM observations, the polysaccharides nets and the phenolic components were recognized through Toluidine blue staining, resulting in pink/violet and blue coloration, respectively (Fig. 8a). Thus, it seems that these polysaccharidic and phenolic components appear to be related to two different morphologies: fibrillary and granular, respectively, as observed in SEM micrographs (Fig. 8b). Furthermore, by observing Fig. 8b, we hypothesized that the thin layer coating the bacteria in Fig. 7f showed a granular matrix. Instead, filamentous networks were observed in the lumen of xylem vessels of twigs’ transverse sections (Fig. 8c), often associated with the presence of bacteria and a granular matrix, and which were absent in the negative control (Fig. 8d). Similar to our observations in S. junceum, Alves et al. (2009) found a sequential process of xylem vessels occlusion in Citrus sinensis leaf petiole and blade naturally infected by Xf, where, following the first steps of colonization, bacterial cells became covered by a fibrillary material until completely occluding the vessel lumen. Moreover, the granular matrix was observed filling the vascular lumen of the xylem vessel (Fig. 8e) in contrast to the negative control (Fig. 8f).
LM image (a) and SEM micrographs (b, c, d, e, and f) of SJ 668. a Cross-section of a green shoot with polysaccharide (pink/violet indicated by asterisk) and phenolic (blue indicated by triangle) matrices stained with Toluidine blue. Vessels colonized by bacteria (arrows). b Cross-section (corresponding to a) of a twig where xylem vessels were colonized by Xfm ST87 in association with filamentous (red arrow) and granular (yellow arrow) matrices. c Cross-section of a xylem vessel showing a filamentous matrix (asterisk). No bacteria are visible. d Cross-section of the negative control green shoot showing xylem vessels without any bacteria. e Longitudinal section where granular matrices (triangle) were identified in the vessel lumen. f Longitudinal section of the negative control green shoot without any bacteria
Conclusions
In this study, the presence of Xfm ST87 was detected by PCR and immunohistochemistry, in the xylem cells of naturally infected S. junceum plants in the outbreak area of Monte Argentario (Grosseto, Tuscany, Italy).
LM and SEM observations showed that bacteria were able to completely fill the xylem vessels after some intermediate steps of infection and spread through the pits in the adjacent conductive elements.
The host was apparently able to react to the infection by producing gums/mucilage and phenols. Xylem vessels containing a high amount of mucilage did not show presence of bacteria, while SEM images showed granular material adhering to bacterial cells, which was probably produced by the surrounding xylem parenchyma. These mechanisms may be considered as an example of constitutive defense systems of the plant against xylem pathogens. Interestingly, bacteria were also found in the conductive elements of the leaves of one of the naturally infected S. junceum plants subject of this study (SJ 796), but they were not found in either the root system or in the flowers.
Artificial inoculations will allow to depict the time course of Xfm ST87 host colonization as well as that of S. junceum responses to the invading pathogen.
References
Abusamra YA, Scuruchi M, Habibatni S, Maammeri Z, Benayache S, D’Ascola A, Avenoso A, Campo GM, Spina E (2015) Evaluation of putative cytotoxic activity of crude extracts from Onopordum acanthium leaves and Spartium junceum flowers against the U-373 glioblastoma cell line. Pak J Pharm Sci 28:1225–1232
Akkemik Ṻ, Kocaba U (2012) Woods of the old galley of Yenikapi, Istanbul. Mediter Archaeol Archaeom 13(2):31–41
Alves E, Leite B, Pascholati SF, Ishida ML, Andersen PC (2009) Citrus sinensis leaf petiole and blade colonization by Xylella fastidiosa: details of xylem vessel occlusion. Sci Agric (Piracicaba, Braz) 6(2):218–224
Backus EA, Morgan DJW (2011) Spatiotemporal colonization of Xylella fastidiosa in its vector supports the role of egestion in the inoculation mechanism of foregut-borne plant pathogens. Phytopathology 101:912–922
Bezić N, Dunkić V, Radonić A (2003) Anatomical and chemical adaptation of Spartium junceum L. in arid habitat. Acta Biol Cracov Ser Bot 45(2):43–47
Bleve G, Marchi G, Ranaldi F, Gallo A, Cimaglia F, Logrieco AF et al (2016) Molecular characteristics of a strain Salento-1 of Xylella fastidiosa isolated in Apulia (Italy) from an olive plant with the quick decline syndrome. Phytopathol Mediterr 55:139–146
Bonsen KJM, Kučera LJ (1990) Vessel occlusions in plants: morphological, functional and evolutionary aspects. IAWA Bulletinn 11(4):393–399
Cardinale M, Luvisi A, Meyer JB, Sabella E, De Bellis L, Cruz AC, Ampatzidis Y, Cherubini P et al (2018) Specific fluorescence in situ hybridization (FISH) test to highlight colonization of xylem vessels by Xylella fastidiosa in naturally infected olive trees (Olea europea L.). Front Plant Sci 9:431
Cariddi C, Saponari M, Boscia D, De Stradis A, Loconsole G, Nigro F, Porcelli F, Potere O, Martelli GP (2014) Isolation of a Xylella fastidiosa strain infecting olive and oleander in Apulia, Italy. J Plant Pathol 96:1–5
Cornara D, Sicard A, Zeilinger AR, Porcelli F, Purcell AH, Almeida RPP (2016) Transmission of Xylella fastidiosa to grapevine by the meadow spittlebug. Phytopathology 106:1285–1290
Crews LJ, McCully ME, Canny MJ (2003) Mucilage production by wounded xylem tissue of maize roots—time course and stimulus Functional. Plant Biol 30:755–766
De Benedictis M, De Caroli M, Baccelli I, Marchi G, Bleve G, Gallo A, Ranaldi F, Falco V, Vittorio P, Piro G, Di Sansebastiano MG, G. P. (2017) Vessel occlusion in three cultivars of Olea europaea naturally exposed to Xylella fastidiosa in open field. J Phytopathol 165:589–594
Dong Z, McCully ME, Canny MJ (1997) Does Acetobacter diazothrophicus live and move in the xylem of sugarcane stems? Anatomical and physiological data. Ann Bot 71:327–341
EFSA, (2020). Update of the Xylella spp. host plant database—systematic literature search up to 30 June 2019. EFSA Journal 2020, 18, 4, 6114
EPPO (2018) PM 7/24 (3). Xylella fastidiosa 48(2):175–248
EPPO (2020). Xylella fastidiosa. EPPO datasheets on pests recommended for regulation. Available online. https://gd.eppo.int.
EU, (2019). Commission database of host plants found to be susceptible to Xylella fastidiosa in the Union territory. https://ec.europa.eu/food/plant/plant_health_biosecurity/legislation/emergency_measures/xylella-fastidiosa/susceptible_en
Francis M, Lin H, Cabrera-La RJ, Doddapaneni H, Civerolo EL (2006) Genome-based PCR primers for specific and sensitive detection and quantification of Xylella fastidiosa. Eur J Plant Pathol 115:203–213
Fritschi FB, Lin H, Walker MA (2008) Scanning electron microscopy reveals different response pattern of four Vitis genotypes to Xylella fastidiosa infection. Plant Dis 92:276–286
Giampetruzzi A, D’Attoma G, Zicca S, Abou KR, Rizzo D, Boscia D et al (2019) Draft genome sequence resources of three strains (TOS4, TOS5, and TOS14) of Xylella fastidiosa infecting different host plants in the newly discovered outbreak in Tuscany, Italy. Phytopathology 109(9):1516–1518
Giménez N, Magro N, Cortés N, Guitart R (2017) Poisoning after ingestion of Spartium junceum seeds: dose-dependent effects in three boys. J Emerg Med 53:e41–e44
Harper SJ, Ward LI, Clover GRG (2010) Development of LAMP and real-time PCR methods for the rapid detection of Xylella fastidiosa for quarantine and field applications. Phytopathology 100:1282–1288
Hopkins DL, Purcell AH (2002) Xylella fastidiosa: cause of Pierce’s disease of grapevine and other emergent diseases. Plant Dis 86:1056–1066
Janse JD, Obradovic A (2010) Xylella fastidiosa: its biology, diagnosis, control and risks. J Plant Pathol 92:35–48
Katović D, Katović A, Krnčević M (2012). Annual 2010/2011 of the Croatian Academy of Engineering 23-37
Landa BB, Castillo AI, Giampetruzzi A, Kahn A, Román-Écija M, Velasco-Amo MP, Navas-Cortés JA, Marco-Noales E, Barbé S, Moralejo E, Coletta-Filho HD, Saldarelli P, Saponari M, Almeida RPP (2020) Emergence of a plant pathogen in Europe associated with multiple intercontinental introductions. Appl Environ Microbiol 86:e01521–e01519. https://doi.org/10.1128/AEM.01521-19
Marchi G, Rizzo D, Ranaldi F, Ghelardini L, Ricciolini M, Scarpelli I, Drosera L, Goti E, Capretti P, Surico G (2018) First detection of Xylella fastidiosa subsp. multiplex DNA in Tuscany (Italy). Phytopathol Mediterr 57(3):363–364
Newman KL, Almeida RPP, Purcell AH, Lindow SE (2003) Use of green fluorescent strain for analysis of Xylella fastidiosa colonization of Vitis vinifera. Appl Environ Microbiol 69:7319–7327
Nilsen ET, Karpa D, Field C (1993) Patterns of stem photosynthesis in two invasive Legumes (Spartium junceum, Cytisus scoparius) of the California coastal region. Am J Bot 80(10):1126–1136
Novelli S, Gismondi A, Di Marco G, Canuti L, Nanni V, Canini A (2019) Plant defense factors involved in Olea europaea resistance against Xylella fastidiosa infection. J Plant Res 132:439–455
Nunes LR, Rosato YB, Muto NH, Yanai GM, da Silva VS, de Leite DB, Gonçalves ER, de Souza AA, Coletta-Filho HD, Machado MA, Lopes SA, de Oliveira RC (2003) Microarray analyses of Xylella fastidiosa provide evidence of coordinated transcription control of laterally transferred elements. Genome Res 13:570–578. https://doi.org/10.1101/gr.930803
O’Brian TP, McCully ME (1981) The study of plant structure: principles and selected methods. Termarcarphi PtyLtd, Australia
Ouyang P, Arif M, Fletcher J, Melcher U, Ochoa Corona FM (2013) Enhanced reliability and accuracy for field deployable bioforensic detection and discrimination of Xylella fastidiosa subsp. pauca, causal agent of citrus variegated chlorosis using Razor Ex technology and TaqMan quantitative PCR. PLoS ONE 8:e81647
Paião FG, Meneguim AM, Casagrande EC, Leite RP (2002) Envolvimento de cigarras (Homoptera, Cicadidae) na transmissão de Xylella fastidiosa em cafeeiro. Fitopatol Bras 27:67
Pèrez-Donoso AG, Lenhof JJ, Pinney K, Labavitch JM (2016) Vessel embolism and tyloses in early stages of Pierce’ disease. Aust J Grape Wine Res 22:81–86
Preti F, Giadrossich F (2009) Root reinforcement and slope bioengineering stabilization by Spanish Broom (Spartium junceum L.). Hydrol Earth Syst Sci 13:1713–1726
Purcell AH (2013) Paradigms: examples from the bacterium Xylella fastidiosa. Annu Rev Phytopathol 51:229–356
Rioux D, Nicole M, Simard M, Ouellette GB (1998) Immunocytochemical evidence that secretion of pectin occurs during gel (gum) and tylosis formation in trees. Phytopathology 88:494–505
Roper C, Castro C, Ingel B (2019) Xylella fastidiosa: bacterial parasitism with hallmarks of commensalism. Curr Opin Plant Biol 50:140–147
Roussel S, Nicole M, Lopez F, Ricci P, Geiger J-P, Renard M, Bruna H (1999) Leptosphaeria maculans and cryptogein induce similar vascular responses in tissues undergoing the hypersensitive reaction in Brassica napus. Plant Sci 144(1):17–28
Sabella E, Luvisi A, Aprile A, Negro C, Vergine M, Nicolì F, Miceli A, De Bellis L (2018) Xylella fastidiosa induces differential expression of lignification related-genes and lignin accumulation in tolerant olive trees cv. Leccino. J Plant Physiol 220:60–68
Sabella E, Aprile A, Genga A, Siciliano T, Nutricati E, Nicolì F, Vergine M, De Bellis L, Luvisi A (2019) Xylem cavitation susceptibility and refilling mechanisms in olive trees infected by Xylella fastidiosa. Sci Rep 9:9602. https://doi.org/10.1038/s41598-019-46092-0
Saponari M, Boscia D, Nigro F, Martelli GP (2013) Identification of DNA sequences related to Xylella fastidiosa in oleander, almond and olive trees exhibiting leaf scorch symptoms in Apulia (Southern Italy). J Plant Pathol 95:659–668
Saponari M, D’Attoma G, Abou KR, Loconsole G, Altamura G, Zicca S et al (2019) A new variant of Xylella fastidiosa subspecies multiplex detected in different host plants in the recently emerged outbreak in the region of Tuscany, Italy. Eur J Plant Pathol 154(4):1195–1200
Severin H (1949) Transmission of the virus of Pierce’s disease of grapevines by leafhoppers. Hilgardia 19:190–206
Severin HHP (1950) Spittle-insect vectors of Pierce’s disease virus II. Life history and virus transmission. Hilgardia 19:357–381
Simpson AJ, Reinach FC, Arruda P, Abreu FA, Acencio M, Alvarenga R, Alves LM, Araya JE, Baia GS, Baptista CS et al (2000) The genome sequence of the plant pathogen Xylella fastidiosa. The Xylella fastidiosa consortium of the organization for nucleotide sequencing and analysis. Nature 406:151–159
Street PFS, Robb J, Ellis BE (1986) Secretion of vascular coating components by xylem parenchyma cells of tomatoes infected with Verticillium albo-atrum. Protoplasma 132:1–11
Sun Q, Sun Y, Walker MA, Labavitch JM (2013) Vascular occlusions in grapevines with Pierce’s disease make disease symptom development worse. Plant Physiol 161:1529–1541
Tagne A, Neergaard E, Hansen HJ (2002) Studies of host-pathogen interaction between maize and Acremomium strictum from Cameroon. Eur J Plant Pathol 108:93–102
Tamura K, Stecher G, Peterson D, Filipski A, Kumar S (2013) MEGA6: molecular evolutionary genetics analysis version 6.0. Mol Biol Evol 30:2725–2729
Van Sluys MA, de Oliveira MC, Monteiro-Vitorello CB, Miyaki CY, Furlan LR, Camargo LEA, da Silva ACR, Moon DH, Takita MA, Lemos EGM, El-Dorry H, Tsai SM, Carrer H, Carraro DM, de Oliveira RC, Nunes LR, Siqueira WJ, Coutinho LL, Kimura ET, Ferro ES, Harakava R, Kuramae EE, Marino CL, Giglioti E, Abreu IL, Alves LMC, do Amaral AM, Baia GS, Blanco SR, Brito MS, Cannavan FS, Celestino AV, da Cunha AF, Fenille RC, Ferro JA, Formighieri EF, Kishi LT, Leoni SG, Oliveira AR, Rosa VE Jr, Sassaki FT, Sena JAD, de Souza AA, Truffi D, Tsukumo F, Yanai GM, Zaros LG, Civerolo EL, Simpson AJG, Almeida NF Jr, Setubal JC, Kitajima JP (2003) Comparative analyses of the complete genome sequences of Pierce’s disease and citrus variegated chlorosis strains of Xylella fastidiosa. J Bacteriol 185:1018–1026
Wells JM, Raju BC, Hung H-Y, Weisburg WG, Mandelco-Paul L, Brenner DJ (1987) Xylella fastidiosa gen. nov., sp. nov: Gram-negative, xylem-limited, fastidious plant bacteria related to Xanthomonas spp. Int J Syst Evol Microbiol 37(2):136–143
Yuan X, Morano L, Bromley R, Spring-Pearson S, Stouthamer R, Nunney L (2010) Multilocus sequence typing of Xylella fastidiosa causing Pierce’s disease and oleander leaf scorch in the United States. Ecology and Epidemiology Phytopathology 100(6):601–611
Funding
Open access funding provided by Università degli Studi di Firenze within the CRUI-CARE Agreement.
Author information
Authors and Affiliations
Corresponding authors
Ethics declarations
Conflict of interest
The authors declare no competing interests.
Additional information
Handling Editor: Alexander Schulz
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
ESM 1
(PDF 1291 kb)
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Falsini, S., Tani, C., Sambuco, G. et al. Anatomical and biochemical studies of Spartium junceum infected by Xylella fastidiosa subsp. multiplex ST 87. Protoplasma 259, 103–115 (2022). https://doi.org/10.1007/s00709-021-01640-2
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00709-021-01640-2