Abstract
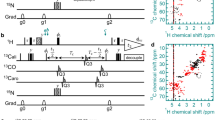
A unique aspect of NMR is its capacity to provide integrated insight into both the structure and intrinsic dynamics of biomolecules. Chemical exchange phenomena that often serve as probes of dynamic processes in biological macromolecules can be quantitatively investigated with chemical exchange saturation transfer (CEST) experiments. 2H-decoupling sidebands, however, always occur in the profiles of 13CHD2 13C-CEST experiments when using the simple CW (continuous wave) method, which may obscure the detection of minor dips of excited states. Traditionally, these sidebands are manually eliminated from the profiles before data analysis by removing experimental points in the range of 2H-decoupling field strength ±50 Hz away from the major dips of the ground state on either side of the dips. Unfortunately, this may also eliminate potential minor dips if they overlap with the decoupling sidebands. Here, we developed methods that use pseudo-continuous waves with variable RF amplitudes distributed onto ramps for 2H decoupling. The new methods were thoroughly validated on Bruker spectrometers at a range of fields (1H frequencies of 600, 700, and 850 MHz, and 1.1 GHz). By using these methods, we successfully removed the sidebands from the NMR profiles of 13CHD2 13C-CEST experiments.







Similar content being viewed by others
Data availability
The datasets generated during and/or analyzed during the current study are available from the corresponding author on reasonable request.
References
Baldwin AJ, Kay LE (2009) NMR spectroscopy brings invisible protein states into focus. Nat Chem Biol 5:808–814. https://doi.org/10.1038/nchembio.238
Bouvignies G, Kay LE (2012) A 2D 13C-CEST experiment for studying slowly exchanging protein systems using methyl probes: an application to protein folding. J Biomol NMR 53:303–310. https://doi.org/10.1007/s10858-012-9640-7
Ernst RR (1966) Nuclear magnetic double resonance with an incoherent radio-frequency field. J Chem Phys 45:3845–3861. https://doi.org/10.1063/1.1727409
Freeman R, Kupče E (1997) Decoupling: theory and practice I. Current methods and recent concepts. NMR Biomed. 10:372–380
Huang C, Kalodimos CG (2017) Structures of large protein complexes determined by nuclear magnetic resonance spectroscopy. Annu Rev Biophys 46:317–336. https://doi.org/10.1146/annurev-biophys-070816-033701
Kupče Ē, Freeman R, Wider G, Wüthrich K (1996) Suppression of cycling sidebands using bi-level adiabatic decoupling. J Magn Reson 122A:81–84. https://doi.org/10.1006/jmra.1996.0180
Levitt MH (1982) Symmetrical composite pulse sequences for NMR population inversion. II. Compensation of resonance offset. J Magn Reson 50:95–110. https://doi.org/10.1016/0022-2364(82)90035-X
Palmer III AG (2014) Chemical exchange in biomacromolecules: past, present, and future. J Magn Reson 241:3–17. https://doi.org/10.1016/j.jmr.2014.01.008
Palmer III AG, Kroenke CD, Loria JP (2001) Nuclear magnetic resonance methods for quantifying microsecond-to-millisecond motions in biological macromolecules. Methods Enzymol 339:204–238. https://doi.org/10.1016/s0076-6879(01)39315-1
Rennella E, Huang R, Velyvis A, Kay LE (2015) 13CHD2–CEST NMR spectroscopy provides an avenue for studies of conformational exchange in high molecular weight proteins. J Biomol NMR 63:187–199. https://doi.org/10.1007/s10858-015-9974-z
Rucker S, Shaka AJ (1989) Broadband homonuclear cross polarization in 2D N.M.R. using DIPSI-2. Mol Phys 68:509–517. https://doi.org/10.1080/00268978900102331
Schanda P, Brutscher B (2005) Very fast two-dimensional NMR spectroscopy for real-time investigation of dynamic events in proteins on the time scale of seconds. J Am Chem Soc 127:8014–8015. https://doi.org/10.1021/ja051306e
Sekhar A, Kay LE (2013) NMR paves the way for atomic level descriptions of sparsely populated, transiently formed biomolecular conformers. Proc Natl Acad Soc USA 110:12867–12874. https://doi.org/10.1073/pnas.1305688110
Shaka AJ, Barker PB, Freeman R (1985) Computer-optimized decoupling scheme for wideband applications and low-level operation. J Magn Reson 64:547–552. https://doi.org/10.1016/0022-2364(85)90122-2
Shaka AJ, Keeler J, Frenkiel T, Freeman R (1983) An improved sequence for broadband decoupling: WALTZ-16. J Magn Reson 52:335–338. https://doi.org/10.1016/0022-2364(83)90207-X
Vallurupalli P, Bouvignies G, Kay LE (2012) Studying “invisible” excited protein states in slow exchange with a major state conformation. J Am Chem Soc 134:8148–8161. https://doi.org/10.1021/ja3001419
Vallurupalli P, Kay LE (2013) Probing slow chemical exchange at carbonyl sites in proteins by chemical exchange saturation transfer NMR spectroscopy. Angew Chem Int Ed Engl 52:4156–4159. https://doi.org/10.1002/anie.201209118
Vallurupalli P, Sekhar A, Yuwen T, Kay LE (2017) Probing conformational dynamics in biomolecules via chemical exchange saturation transfer: a primer. J Biomol NMR 67:243–271. https://doi.org/10.1007/s10858-017-0099-4
Xie T, Saleh T, Rossi P, Kalodimos CG (2020) Conformational states dynamically populated by a kinase determine its function. Science 370:eabc2754. https://doi.org/10.1126/science.abc2754
Zhou Z, Kümmerle R, Qiu XH, Redwine D, Cong R, Taha A, Baugh D, Winniford B (2007) A new decoupling method for accurate quantification of polyethylene copolymer composition and triad sequence distribution with 13C NMR. J Magn Reson 187:225–233. https://doi.org/10.1016/j.jmr.2007.05.005
Acknowledgements
We thank Lewis E. Kay, Ph.D., for original pulse sequence, advice and discussion, Zhilian Xia for providing the 13CHD2-labeled sample and Nisha Badders, PhD, ELS, for scientific editing. This work is financially supported by USA NIH Grants R35 GM122462 to C.G.K. and by the Biomolecular NMR Center, St. Jude Children’s Research Hospital.
Author information
Authors and Affiliations
Corresponding authors
Ethics declarations
Conflict of interest
The authors have no conflicts of interest to declare.
Additional information
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Xia, Y., Yuwen, T., Liu, A. et al. Removal of 2H-decoupling sidebands in 13CHD2 13C-CEST profiles. J Biomol NMR 75, 133–142 (2021). https://doi.org/10.1007/s10858-021-00362-0
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10858-021-00362-0