Abstract
Droplet microfluidics is a powerful tool for a diverse range of biomedical and industrial applications such as single-cell biology, synthetic biology, digital PCR, biosafety monitoring, drug screening, and food, feed, and cosmetic industries. As an integral part of droplet microfluidics, on-chip multiplexed droplet sorting has recently gained enthusiasm, since it enables real-time sorting of single droplets containing cells with different phenotypes into multiple bins. However, conventional sorting methods are limited in throughput and scalability. Here, we present high-throughput, scalable, multiplexed droplet sorting by employing a pair of sequentially addressable dielectrophoretic arrays (SADAs) across a microchannel on a microfluidic chip. A SADA is an on-chip array of electrodes, each of which is sequentially activated and deactivated in synchronization to the position and speed of a flowing droplet of interest. The dual-SADA (dSADA) structure enables high-throughput deflection of droplets in multiple directions in a well-controlled manner. For proof-of-concept demonstration and characterization of the dSADA, we performed fluorescence-activated droplet sorting (FADS) with a 3-way dSADA at a high throughput of 2450 droplets/s. Furthermore, to show the scalability of the dSADA, we also performed FADS with a 5-way dSADA at a high throughput of 473 droplets/s.
Similar content being viewed by others
1 Introduction
Droplet microfluidics is a powerful tool for a diverse range of biomedical and industrial applications such as single-cell biology (Mazutis et al. 2013; Wang et al. 2014), synthetic biology (Abatemarco et al. 2017), digital PCR (Hindson et al. 2013), biosafety monitoring (Chaipan et al. 2017), drug screening (Miller et al. 2012), and food, feed, and cosmetic industries (Muschiolik 2007; Yukuyama et al. 2016). It has shown a new paradigm by virtue of its capability to confine, cultivate, and analyze single cells, clones, or exosomes in monodisperse droplets that serve as isolated reaction and transport vessels (Ahn et al. 2006; Brouzes et al. 2009; Guo et al. 2012; Kintses et al. 2012; Best et al. 2016; Kim et al. 2017; Shang et al. 2017; Ding et al. 2019; Matuła et al. 2019; Suea-Ngam et al. 2019). These advantages allow us to study and exploit cellular proliferation, protein secretion, antibody production, and enzyme activity in isolated environments at the single-cell level (Ahn et al. 2006; Brouzes et al. 2009; Guo et al. 2012; Kintses et al. 2012; Best et al. 2016; Kim et al. 2017; Shang et al. 2017; Ding et al. 2019; Matuła et al. 2019; Suea-Ngam et al. 2019). While droplet microfluidics has been exploited over the last decade, its full potential remains untapped, especially for industrial deployment.
As an integral part of droplet microfluidics, on-chip multiplexed droplet sorting has recently gained enthusiasm since it enables real-time sorting of single droplets containing cells with different phenotypes (e.g., growth rate, secretion rate, antibody production rate) into multiple bins (Frenzel and Merten 2017; Caen et al. 2018). While multiplexed droplet sorting is a well-established method in fluorescence-activated cell sorting (FACS) (Cossarizza et al. 2019), its on-chip implementation remains challenging and unexploited in spite of its enormous potential for the above applications. This is because the motion and thus manipulation of droplets in a microchannel are resisted by shear stress on them due to their surrounding liquid medium. To address this need and hence go beyond what is possible with FACS in terms of utility, efforts have been made to develop various approaches for droplet actuation [e.g., magnetic (Zhang et al. 2009), thermal (Baroud et al. 2007), electrical (Mazutis et al. 2013; Ahmadi et al. 2019), acoustic (Li et al. 2013), pneumatic (Wu et al. 2013)], among which dielectrophoretic actuation has been shown the most reliable, robust, and fast (Xi et al. 2017) as it is directly driven by electronics that provides high controllability both spatially and temporally.
Unfortunately, conventional on-chip dielectrophoretic sorting methods are limited in throughput and scalability due to their simple electrode design for sorting actuation, thereby leading to low degrees of freedom in force generation. Specifically, two types of on-chip multiplexed droplet sorters have been previously reported (Frenzel and Merten 2017; Caen et al. 2018). One type is based on serial integration of multiple binary droplet sorters (Frenzel and Merten 2017), in which a single pair of electrodes is employed for each binary droplet sorter. Its throughput is only a few droplets/s. The other type is a droplet sorter with the ability to tune the strength of the applied force (Caen et al. 2018), in which two pairs of electrodes are employed across a microchannel. Its throughput can be up to 200 droplets/s. The electrode design of these pioneering multiplexed droplet sorters has been optimized since their first reports, while its basic concept remains the same as the single-pair-electrode-based droplet sorters, placing a limit on the sorting throughput and/or the number of outlets.
In this article, we present high-throughput, scalable, multiplexed droplet sorting by employing a pair of sequentially addressable dielectrophoretic arrays (SADAs) across a microchannel on a microfluidic chip. A SADA is an on-chip array of electrodes, each of which is sequentially activated and deactivated in synchronization to the position and speed of a flowing droplet of interest (Isozaki et al. 2020b). In addition to this capability, the pair of SADAs, which we refer to as a dual SADA (dSADA), enables high-throughput deflection of droplets in multiple directions in a well-controlled manner. Specifically, the direction and magnitude of the droplet deflection can be controlled by selectively activating electrodes and tuning the number of activated electrodes in the dSADA, respectively. For proof-of-concept demonstration and characterization of the dSADA, we performed a finite element method (FEM)-based simulation and fluorescence-activated droplet sorting (FADS) of 58-pL (48 μm in diameter) droplets with a 3-way dSADA at a high throughput of 2450 droplets/s. Furthermore, to show the scalability of the dSADA, we also performed FADS of 66-pL (50 μm in diameter) droplets with a 5-way dSADA at a high throughput of 473 droplets/s.
2 Results
2.1 Overview of FADS with the dSADA
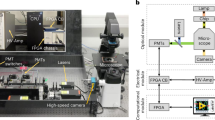
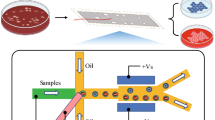
As shown in Fig. 1a, our FADS system consists of (i) a microfluidic chip that combines a droplet generator, a droplet accelerator, and a dSADA sorter containing two electrode arrays, (ii) an optical setup to detect fluorescence signals from cells within droplets, and (iii) an electric controller to activate or deactivate the electrodes of the arrays of the dSADA sorter. The operation of the FADS system is as follows. Using a syringe pump, cells suspended in a medium are injected into the microfluidic chip. The injected cells are encapsulated into droplets by the droplet generator. The generated droplets are accelerated by introducing additional fluids (oil phase) to increase the inter-droplet distance. The accelerated droplets are optically interrogated by the optical setup. The fluorescence signals from the cells are detected by the optical setup composed of a continuous-wave (CW) laser diode and a photomultiplier tube (PMT). The PMT signal is processed by a field-programmable gate array (FPGA)-based electrical control system [see Isozaki et al. (2020b) for details], followed by triggering the activation of the electrodes of the dSADA and generating dielectrophoretic force at a controlled time delay. The dielectrophoretic force is used to manipulate a droplet containing target cell(s) into one of the collection channels on the chip. The detection and sorting processes are monitored with a high-speed camera. The details of each component are described below.
2.2 Functionality of the dSADA
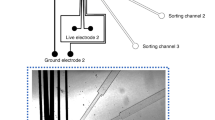
As schematically illustrated in Fig. 1b, the functionality of the dSADA is based on a sequential process that is similar to the sorting mechanism of the SADA [see Isozaki et al. (2020b) for details], but with multiple modifications to realize multiplexed droplet sorting. First, to sort droplets into the upper channel(s) [corresponding to Outlet 1 and Outlet 2 in Fig. 1b], the electrodes of the upper SADA are sequentially activated and deactivated. An activated driving electrode generates a spatially nonuniform electric field with the neighboring grounded electrodes, exerting a localized dielectrophoretic force to pull a target droplet. Note that the localized force is not applied to the neighboring droplets preceding or following the target droplet. As mentioned above, the timings of the activation and deactivation of the electrodes are synchronized to the position and speed of the target droplet passing through the dSADA region. Using all five driving electrodes of the upper SADA, the dSADA can sort droplets into the outer channel [corresponding to Outlet 1 in Fig. 1b], while using the three upstream electrodes, the dSADA can sort droplets into the inner channel [corresponding to Outlet 2 in Fig. 1b] since the sum of the applied dielectrophoretic force used to pull the target droplet is weaker than that in the former case. As the dSADA has a symmetric structure, sorting of droplets into the lower channel(s) (corresponding to Outlet 4 and Outlet 5) can be similarly performed. In the absence of the external force, droplets flow into the middle channel (corresponding to Outlet 3) by default.
2.3 Electrical simulation of FADS with the dSADA
To clarify the functionality of the dSADA, we performed an FEM-based electrical simulation with a commercial software package (Comsol Multiphysics ver. 4.4, COMSOL, U.S.A.). The simulation model is a two-dimensional static model and consists of two electrode arrays, carrier oil in a microchannel, water droplets, and polydimethylsiloxane (PDMS) microchannel walls (Fig. 2a). The material properties of each element are shown in Table 1. The width of the microchannel, the width of the driving electrodes, and the width of the ground electrodes are given by 150 µm, 100 µm, and 40 µm, respectively. The droplet diameter d and droplet pitch p are variable parameters. An 80-kHz 1-kV peak-to-peak AC signal is applied to the upper center driving electrode.
Simulation of the dSADA. a Simulation model. b Simulation of deflecting a single droplet, showing the electrical fields along the microchannel and the dielectrophoretic force on the target droplet, for the following droplet (upper) and preceding droplet (lower). The pitches of the droplets in the upper and lower models are 60 µm and 150 µm, respectively. c Dielectrophoretic force on the target and non-target droplets and the ratio between them as a function of the droplet pitch. d Simulations of deflecting a single droplet. The electrical fields along the microchannel for sorting droplets with various diameters are shown. e Dielectrophoretic force on the target and non-target droplets and the ratio between them as a function of the droplet diameter
Figure 2b shows the simulation results of the distribution patterns of the electric field in the entire dSADA region with a droplet diameter of d = 50 µm and droplet pitches of p = 60 µm and 150 µm. The insets of the figure show the distribution patterns of the dielectrophoretic force on the target droplet and an adjacent droplet. In the case of the small droplet pitch (p = 60 µm), the spatial gradient of the electric field generated by the activated driving electrode affects not only the target droplet, but also the adjacent droplets. Moreover, electrical interactions between the droplets exist, leading to fluctuations in the flow speed of droplets. On the other hand, in the case of the large droplet pitch (p = 150 µm), the dielectrophoretic force applies only to the target droplet, enabling sorting of the target droplet(s) from a stream of droplets.
To quantitatively evaluate the ability to sort a target droplet from a stream of droplets including non-target droplets, we calculated the dielectrophoretic force on the target droplet and adjacent droplets as a function of the pitch of the 50-μm droplets. Furthermore, we calculated the ratio of the dielectrophoretic force on the target droplet (Ft) to the dielectrophoretic force on an adjacent droplet (Fn). As shown in Fig. 2c, Fn exponentially decreases with an increase of the pitch of droplets p, while Ft slowly decreases. Therefore, the ratio exponentially increases. Assuming that this ratio must be more than 30 to effectively sort a single target droplet at a time without affecting other droplets and the flow speed of the droplets flowing at the center of the channel is 0.5 m/s, the throughput of the achievable design specification is derived to be ~ 3000 droplets/s by dividing 0.5 m/s by ~ 0.15 mm. Note that the throughput of the dSADA should be much lower, because the flow speed of the droplets is lower for multiple reasons: the target droplets are attracted to a channel wall where the flow speed is lower than that at the center of the channel and the width of the main channel increases downstream to introduce multiple outlets.
To investigate the effect of the droplet size on the sorting performance, we performed simulations with a constant droplet pitch (p = 150 µm) and varying droplet sizes. As shown in Fig. 2d, the droplet size affects the local electric field distribution with larger droplets affecting it to a larger degree. On the other hand, the impact of the electric field differences owing to the droplet size on the sorting performance is negligible. Specifically, as shown in Fig. 2e, the ratio of Ft to Fn is on the same order of magnitude as a function of the droplet size ranging from 10 to 100 μm. These results indicate that the dSADA can sort droplets with various sizes. Note that the optimization of the microchannel design is necessary for sorting droplets with a certain size. In this article, we optimized the microchannel design so that ~ 50-μm droplets could be sorted at a high throughput.
2.4 Demonstration of multiplexed FADS with the dSADA
To demonstrate multiplexed FADS with the dSADA, we designed, fabricated, and demonstrated a three-way dSADA-based FADS system with three outlets. Here, the width of the main channel was made constant in the dSADA region. To perform sorting experiments with the three-way dSADA sorter, we used deionized water containing fluorescent ink as the dispersed phase such that all droplets generated from the dispersed phase would emit fluorescence that could trigger the electronics via the PMT (Fig. 1a). Moreover, we developed a sorting function program so that one out of five droplets is sorted alternatively into the upper and lower channels as a proof-of-concept demonstration. The sorting operation was monitored and evaluated with the high-speed camera (see Materials and Methods for details of the optical setup and dSADA operation).
Figure 3 shows a sequence of image frames captured by the high-speed camera (see Movie S1 for the complete image sequence). The images show that target droplets are gradually deflected and sorted into the upper or lower outlet while non-target droplets flow into the middle outlet by default. The pitch of droplets, the flow speed of droplets, and the sorting throughput were found to be ~ 312 μm, ~ 0.765 m/s, and 2450 droplets/s, respectively. The obtained sorting throughput is reasonably high enough compared to the throughput of the achievable design specification (~ 3000 droplets/s). Furthermore, we experimentally measured the trajectories of the target droplets and three non-target droplets to quantitatively clarify the functionality of the three-way dSADA-based FADS system. As shown in Fig. 4, at the fifth driving electrode, the total lateral displacement of both target droplets reached 25 μm, which is sufficient to sort droplets into the upper or lower outlet, while the total lateral displacement of the non-target droplets was less than several micrometers, which was negligibly small. By monitoring the sorting process of 21 videos recorded by the high-speed camera, the success rate of the sorting process was found to be 100%. These results firmly indicate that the concept of the dSADA works as expected.
2.5 Demonstration of scalable FADS with the dSADA
To demonstrate the scalability of the dSADA, we designed and fabricated a five-way dSADA-based FADS system and evaluated its performance using deionized water containing fluorescent ink. To increase the number of outlet channels from three to five, we modified the design of the three-way dSADA. Specifically, as shown in Fig. 5, the width of the main channel was made constant in the dSADA region from the first (most upstream) driving electrodes to the third driving electrodes, while the channel was widened in the dSADA region from the fourth driving electrodes to the fifth driving electrodes. The main channel at the fifth driving electrodes was wider than that at the first driving electrodes by a factor of ~ 2.4. With this modification, we improved the spatial resolution of the position of droplets for accurate multiplexed droplet sorting, such that the tolerance of the droplet displacement for sorting was increased. As schematically shown in Fig. 5 as an example, to sort droplets into the upper outer outlet (Outlet 1), all the driving electrodes of the upper SADA were activated, while to sort droplets into the upper inner outlet (Outlet 2), the first, second, and third driving electrodes were activated.
We evaluated the performance of the five-way dSADA-based FADS system using protocols similar to those for the three-way dSADA-based FADS demonstration. Note that we programmed the sorting function so that four consecutive droplets were sorted into Outlet 1, Outlet 2, Outlet 5, and Outlet 4 in sequence. Figure 5 shows the trajectories of the sorted droplets recorded by the high-speed camera (see Movie S2 for the complete image sequence). The image in Fig. 5 was generated by stacking the captured images. These trajectories indicate that the five-way dSADA operation worked well. To quantify the success rate of the sorting process, we took a movie comprising 356 sorting events. The movie shows no sorting failure, meaning that the sorting accuracy was 100% at a high throughput of 473 droplets/s. In addition, we performed droplet sorting using another program that was developed to direct all droplets into a target outlet (Outlet 1, Outlet 2, Outlet 4, or Outlet 5). As shown in Movie S3, the five-way dSADA operation worked well even at a high concentration of target droplets.
To show the utility of the five-way dSADA to biological applications, we used it to demonstrate sorting of biological cells encapsulated in droplets made of a biologically friendly material. Specifically, we performed multiplexed sorting of Chlamydomonas reinhardtii cells encapsulated in 46-μm droplets generated from culture medium (AF-6; conductivity: 39.7 mS/m) according to the number of cells encapsulated in droplets with a high throughput of 667 droplets/s. As shown in Movie S4, culture-medium droplets were deflected by the dSADA in a similar manner to deionized water droplets, indicating the applicability of the dSADA for biological applications where the conductivity of the droplet-forming media is typically higher than that of deionized water.
3 Discussion
As summarized in Fig. 6, our dSADA enabled multiplexed droplet sorting with higher throughputs than previously reported multiplexed droplet sorters (Frenzel and Merten 2017; Caen et al. 2018). Specifically, the throughput of the three-way dSADA-based FADS system is comparable to that of the previously reported two-way SADA-based FADS system (Isozaki et al. 2020b), while the sorting throughput tends to decrease with an increase of the number of outlets. Moreover, the throughput of the five-way dSADA-based FADS system is three times higher than that of the previously demonstrated five-way droplet sorting system by Caen et al. (2018). These results indicate that the dSADA-based FADS opens up the possiblity for sorting samples that consist of multiple populations and rare targets, which is promising for many applications such as pathogen detection for food quality control (Ngamsom et al. 2016), screening for drug sensitivity of bacteria (Boedicker et al. 2008), and multiplexed gene sequencing for studying cell heterogeneity (Kang et al. 2018).
The throughput and scalability of the dSADA can be further improved by taking multiple approaches. First, as discussed in our previous paper (Isozaki et al. 2020b), the number of electrodes in the SADA can be increased without a limit. Second, the microchannel design can be optimized. For example, while in this work, we designed the main channel combining the straight channel region (from the first driving electrodes to the third driving electrodes) and the slanted channel region (from the fourth driving electrodes to the fifth driving electrodes), a slanted channel can be employed over the entire region of the dSADA. Third, the strength of the applied force to target droplets can be tuned via varying the applied voltage to the electrodes, leading to an increased range of deflection patterns. Fourth, a gapped divider that has been previously reported (Sciambi and Abate 2015) can be implemented into the dSADA. Specifically, the gapped divider forms a gap between the channel ceiling and the floor, avoiding droplet breakage by colliding the droplets with the divider such that the droplets can squeeze. Fifth, the dSADA can be employed three dimensionally by forming another dSADA in the orthogonal direction to the first dSADA, thereby increasing the number of outlets significantly, as it enables droplet sorting in three dimensions. This fabrication is feasible with the rapidly-growing 3D-printer technology (Yazdi et al. 2016; Li et al. 2019). With these improvements, the dSADA has the potential for super-multiplexed droplet sorting.
To maximally take advantage of the capability of dSADA-based FADS, multimodal and/or high-content analysis methods are in good synergy with it, because single cells encapsulated in droplets can be separated into multiple bins based on the availability of rich cellular information provided by the analytical methods. For example, analogous to FACS, multicolor fluorescence detection with FADS (Perfetto et al. 2004; Cossarizza et al. 2019) is applicable to dSADA-based FADS and can widen its application range. Also, high-speed Raman spectroscopy can be employed with dSADA-based droplet sorting for high-throughput, label-free identification of intracellular molecules (Butler et al. 2016; Wang et al. 2017; Lindley et al. 2019; Hiramatsu et al. 2020). Furthermore, high-throughput imaging (Diebold et al. 2013; Wong et al. 2014; Barteneva and Vorobjev 2015; Han and Lo 2015; Lau et al. 2016; Mikami et al. 2018a, b, 2020; Nitta et al. 2018; Lei et al. 2018; Gu et al. 2019; Isozaki et al. 2019, 2020a) can be used in conjunction with dSADA-based FADS to provide microscopy-like high-content analytical capabilities. These integrations and additional capabilities hold promise for a new class of opportunities for diverse biomedical and industrial applications such as single-cell biology, synthetic biology, digital PCR, biosafety monitoring, drug screening, and food, feed, and cosmetic industries.
4 Materials and methods
4.1 Optical setup
We used the optical setup described in our previous paper (Isozaki et al. 2020b). Briefly, the optical setup consists of a fluorescence detection module (see Fig. 1a) and an optical monitoring module (not shown in Fig. 1a). The fluorescence detection module consists of a CW laser diode (532 nm), a cylindrical lens (f = − 200 mm), a short-pass dichroic beamsplitter (cutoff wavelength of 550 nm), an objective lens (6.3 ×), a long-pass filter (cutoff wavelength of 575 nm), a spherical lens (f = 175 mm), an iris, and a PMT (H7422-40, Hamamatsu Photonics, Japan). The laser light is focused by the cylindrical lens and objective lens onto the center of the stream of droplets in the microchannel and 200-µm upstream of the first electrode of the dSADA. The fluorescence from each cell in a droplet is detected by the PMT via the same objective lens, dichroic beamsplitter, long-pass filter, spherical lens, and iris. The optical monitoring module consists of a halogen lamp (OSL2, Thorlabs, USA), a band-pass filter (with 20-nm bandwidth centered at 540 nm), a 70:30 beamsplitter, a notch filter (with 17-nm bandwidth centered at 533 nm), a spherical lens (f = 175 mm), a high-speed camera (Phantom v2640, Vision Research, USA), and the same objective lens shared with the fluorescence detection module.
4.2 dSADA chip
The dSADA chip consists of a polydimethylsiloxane (PDMS) slab with microchannel structures and a glass substrate with Cr/Au electrodes fabricated using conventional soft-lithography and photolithography, respectively. See our previous paper (Isozaki et al. 2020b) for the details of the fabrication process. The height and width of the main microchannel are ~ 50 µm and 150 µm, respectively. The width and pitch of the driving electrodes are 100 µm and 200 µm, respectively. The surface of the microchannel was made hydrophobic by washing the microchannel with 0.1% perfluorodecyl dimethylchlorosilane solution.
4.3 Operation parameters of the dSADA
We used a sample solution (deionized water containing fluorescent ink or C. reinhardtii cells in AF-6 medium) as the dispersed phase, hexadecane with span 80 (concentration: 1%) as the continuous phase to generate droplets, and hexadecane with span 80 (concentration: 1% for the deionized water solution or 10% for the AF-6 sample solution) as the continuous phase to accelerate droplets. The high concentration of span 80 in the second continuous phase was for preventing the coalescence of droplets. The flow rates of the dispersed phase, continuous phase for generating droplets, and continuous phase for accelerating droplets were typically 2, 60, and 100 µL/min, respectively. We applied 80.1-kHz sinusoidal-like voltages generated by amplifying 3.3-Vpp square-shaped signals to a high voltage of ~ 2.4 kVpp using home-made amplifiers (Isozaki et al. 2020b). The parameters for the flow rates and voltages were optimized, depending on the target throughput and the type of dSADA (i.e., three-way dSADA or five-way dSADA). All the settings were manually tuned via monitoring droplet sorting processes with the high-speed camera. We operated the dSADA using the aforementioned parameters following the protocols shown in our previous paper (Isozaki et al. 2020b) as a basis, but with some modifications. Briefly, we aligned the position of the excitation laser, tuned the flow rates, checked the fluorescence signals from the droplets or cells, tuned the signal thresholds for activating the electrodes, measured the flow speed of the droplets, input the timing used to apply the voltage to each driving electrode, manually optimized the timing, and tuned the amplitude of the applied voltage.
4.4 Cultivation and preparation of C. reinhardtii cells
C. reinhardtii TKAC1017 was provided by Tsuruoka, Keio, Algae Collection (TKAC) of T. Nakada at the Institute for Advanced Biosciences, Keio University. C. reinhardtii cells were cultured in 10 mL of AF-6 medium in 50-mL flasks (690195, Greiner, Austria). The cells were cultivated in a custom-made growth chamber [see (Isozaki et al. 2020b) for details] at 25 °C under warm-white LED lighting at 120 µmol photons/m2/s with a 12 h/12 h light/dark cycle. The cells were subcultured every 3–4 days at a log growth phase cell concentration. For the droplet sorting experiments, the cells were concentrated by a centrifuge to a concentration of ~ 2.65 × 107 cells/mL. Assuming that the droplet volume and cell concentration are 50 pL and 2.65 × 107 cells/mL, respectively, the probabilities that a droplet contains 0, 1, 2, 3, 4, and > 5 cells were found to be 26.58%, 35.22%, 23.33%, 10.31%, 3.41%, and 1.15%, respectively (Mazutis et al. 2013). No staining process was necessary since the autofluorescence of the cells was used to trigger the dSADA.
Availability of data and materials
The raw data in this manuscript are available from the corresponding author upon request.
Code availablity
The codes in this manuscript are available from the corresponding author upon request.
References
Abatemarco J, Sarhan MF, Wagner JM et al (2017) RNA-aptamers-in-droplets (RAPID) high-throughput screening for secretory phenotypes. Nat Commun 8:332. https://doi.org/10.1038/s41467-017-00425-7
Ahmadi F, Samlali K, Vo PQN, Shih SCC (2019) An integrated droplet-digital microfluidic system for on-demand droplet creation, mixing, incubation, and sorting. Lab Chip 19:524–535. https://doi.org/10.1039/c8lc01170b
Ahn K, Kerbage C, Hunt TP et al (2006) Dielectrophoretic manipulation of drops for high-speed microfluidic sorting devices. Appl Phys Lett 88:024104. https://doi.org/10.1063/1.2164911
Baroud CN, Delville JP, Gallaire F, Wunenburger R (2007) Thermocapillary valve for droplet production and sorting. Phys Rev E 75:046302. https://doi.org/10.1103/PhysRevE.75.046302
Barteneva NS, Vorobjev IA (2015) Imaging flow cytometry: methods and protocols. Springer, New York
Best RJ, Lyczakowski JJ, Abalde-Cela S et al (2016) Label-free analysis and sorting of microalgae and cyanobacteria in microdroplets by intrinsic chlorophyll fluorescence for the identification of fast growing strains. Anal Chem 88:10445–10451. https://doi.org/10.1021/acs.analchem.6b02364
Boedicker JQ, Li L, Kline TR, Ismagilov RF (2008) Detecting bacteria and determining their susceptibility to antibiotics by stochastic confinement in nanoliter droplets using plug-based microfluidics. Lab Chip 8:1265–1272. https://doi.org/10.1039/b804911d
Brouzes E, Medkova M, Savenelli N et al (2009) Droplet microfluidic technology for single-cell high-throughput screening. Proc Natl Acad Sci 106:14195–14200. https://doi.org/10.1073/pnas.0903542106
Butler HJ, Ashton L, Bird B et al (2016) Using Raman spectroscopy to characterize biological materials. Nat Protoc 11:664–687. https://doi.org/10.1038/nprot.2016.036
Caen O, Schütz S, Jammalamadaka MSS et al (2018) High-throughput multiplexed fluorescence-activated droplet sorting. Microsystems Nanoeng 4:33. https://doi.org/10.1038/s41378-018-0033-2
Chaipan C, Pryszlak A, Dean H et al (2017) Single-virus droplet microfluidics for high-throughput screening of neutralizing epitopes on HIV particles. Cell Chem Biol 24:751–757. https://doi.org/10.1016/j.chembiol.2017.05.009
Cossarizza A, Chang H, Radbruch A et al (2019) Guidelines for the use of flow cytometry and cell sorting in immunological studies (second edition). Eur J Immunol 49:1457–1973. https://doi.org/10.1016/j.optmat.2011.11.002
Diebold ED, Buckley BW, Gossett DR, Jalali B (2013) Digitally synthesized beat frequency multiplexing for sub-millisecond fluorescence microscopy. Nat Photonics 7:806–810. https://doi.org/10.1038/nphoton.2013.245
Ding Y, Howes PD, Demello AJ (2019) Recent advances in droplet microfluidics. Anal Chem 92:132–149. https://doi.org/10.1021/acs.analchem.9b05047
Frenzel D, Merten CA (2017) Microfluidic train station: highly robust and multiplexable sorting of droplets on electric rails. Lab Chip 17:1024–1030. https://doi.org/10.1039/c6lc01544a
Gu Y, Zhang AC, Han Y et al (2019) Machine learning based real-time image-guided cell sorting and classification. Cytom Part A 95:499–509. https://doi.org/10.1002/cyto.a.23764
Guo MT, Rotem A, Heyman JA, Weitz DA (2012) Droplet microfluidics for high-throughput biological assays. Lab Chip 12:2146. https://doi.org/10.1039/c2lc21147e
Han Y, Lo Y (2015) Imaging cells in flow cytometer using spatial-temporal transformation. Sci Rep 5:13267. https://doi.org/10.1038/srep13267
Hindson CM, Chevillet JR, Briggs HA et al (2013) Absolute quantification by droplet digital PCR versus analog real-time PCR. Nat Methods 10:1003–1005. https://doi.org/10.1038/nmeth.2633
Hiramatsu K, Yamada K, Lindley M et al (2020) Large-scale label-free single-cell analysis of paramylon in Euglena gracilis by high-throughput broadband Raman flow cytometry. Biomed Opt Express 11:1752–1759. https://doi.org/10.1364/BOE.382957
Isozaki A, Mikami H, Hiramatsu K et al (2019) A practical guide to intelligent image-activated cell sorting. Nat Protoc 14:2370–2415. https://doi.org/10.1038/s41596-019-0183-1
Isozaki A, Mikami H, Tezuka H et al (2020a) Intelligent image-activated cell sorting 2.0. Lab Chip 20:2263–2273. https://doi.org/10.1039/d0lc00080a
Isozaki A, Nakagawa Y, Loo MH et al (2020b) Sequentially addressable dielectrophoretic array for high-throughput sorting of large-volume biological compartments. Sci Adv 6:ab06712. https://doi.org/10.1126/sciadv.aba6712
Kang HM, Subramaniam M, Targ S et al (2018) Multiplexed droplet single-cell RNA-sequencing using natural genetic variation. Nat Biotechnol 36:89–94. https://doi.org/10.1038/nbt.4042
Kim HS, Hsu SC, Han SI et al (2017) High-throughput droplet microfluidics screening platform for selecting fast-growing and high lipid-producing microalgae from a mutant library. Plant Direct 1:e00011. https://doi.org/10.1002/pld3.11
Kintses B, Hein C, Mohamed MF et al (2012) Picoliter cell lysate assays in microfluidic droplet compartments for directed enzyme evolution. Chem Biol 19:1001–1009. https://doi.org/10.1016/j.chembiol.2012.06.009
Lau AKS, Shum HC, Wong KKY, Tsia KK (2016) Optofluidic time-stretch imaging-an emerging tool for high-throughput imaging flow cytometry. Lab Chip 16:1743–1756. https://doi.org/10.1039/c5lc01458a
Lei C, Kobayashi H, Wu Y et al (2018) High-throughput imaging flow cytometry by optofluidic time-stretch microscopy. Nat Protoc 13:1603–1631. https://doi.org/10.1038/s41596-018-0008-7
Li S, Ding X, Guo F et al (2013) An on-chip, multichannel droplet sorter using standing surface acoustic waves. Anal Chem 85:5468–5474. https://doi.org/10.1021/ac400548d
Li F, Macdonald NP, Guijt RM, Breadmore MC (2019) Increasing the functionalities of 3D printed microchemical devices by single material, multimaterial, and print-pause-print 3D printing Feng. Lab Chip 19:35–49. https://doi.org/10.1039/c8lc00826d
Lindley M, Hiramatsu K, Nomoto H et al (2019) Ultrafast simultaneous Raman-fluorescence spectroscopy. Anal Chem 91:15563–15569. https://doi.org/10.1021/acs.analchem.9b03563
Matuła K, Rivello F, Huck WTS (2019) Single-cell analysis using droplet microfluidics. Adv Biosyst 4:1900188. https://doi.org/10.1002/adbi.201900188
Mazutis L, Gilbert J, Ung WL et al (2013) Single-cell analysis and sorting using droplet-based microfluidics. Nat Protoc 8:870–891. https://doi.org/10.1038/nprot.2013.046
Mikami H, Harmon J, Kobayashi H et al (2018a) Ultrafast confocal fluorescence microscopy beyond the fluorescence lifetime limit. Optica 5:117–126. https://doi.org/10.1364/OPTICA.5.000117
Mikami H, Lei C, Nitta N et al (2018b) High-speed imaging meets single-cell analysis. Chemistry 4:2278–2300. https://doi.org/10.1016/j.chempr.2018.06.011
Mikami H, Kawaguchi M, Huang CJ et al (2020) Virtual-freezing fluorescence imaging flow cytometry. Nat Commun 11:1162. https://doi.org/10.1038/s41467-020-14929-2
Miller OJ, El A, Mangeat T et al (2012) High-resolution dose–response screening using droplet-based microfluidics. Proc Natl Acad Sci 109:378–383. https://doi.org/10.1073/pnas.1113324109
Muschiolik G (2007) Multiple emulsions for food use. Curr Opin Colloid Interface Sci 12:213–220. https://doi.org/10.1016/j.cocis.2007.07.006
Ngamsom B, Esfahani MMN, Phurimsak C et al (2016) Multiplex sorting of foodborne pathogens by on-chip free-flow magnetophoresis. Anal Chim Acta 918:69–76. https://doi.org/10.1016/j.aca.2016.03.014
Nitta N, Sugimura T, Isozaki A et al (2018) Intelligent image-activated cell sorting. Cell 175:266–276. https://doi.org/10.1016/j.cell.2018.08.028
Perfetto SP, Chattopadhyay PK, Roederer M (2004) Seventeen-colour flow cytometry: unravelling the immune system. Nat Rev Immunol 4:648–655. https://doi.org/10.1038/nri1416
Sciambi A, Abate AR (2015) Accurate microfluidic sorting of droplets at 30 kHz. Lab Chip 15:47–51. https://doi.org/10.1039/C4LC01194E
Shang L, Cheng Y, Zhao Y (2017) Emerging droplet microfluidics. Chem Rev 117:7964–8040. https://doi.org/10.1021/acs.chemrev.6b00848
Suea-Ngam A, Howes PD, Srisa-Art M, Demello AJ (2019) Droplet microfluidics: from proof-of-concept to real-world utility? Chem Commun 55:9895–9903. https://doi.org/10.1039/c9cc04750f
Wang BL, Ghaderi A, Zhou H et al (2014) Microfluidic high-throughput culturing of single cells for selection based on extracellular metabolite production or consumption. Nat Biotechnol 32:473–478. https://doi.org/10.1038/nbt.2857
Wang X, Ren L, Su Y et al (2017) Raman-activated droplet sorting (RADS) for label-free high-throughput screening of microalgal single-cells. Anal Chem 89:12569–12577. https://doi.org/10.1021/acs.analchem.7b03884
Wong TTW, Lau AKS, Ho KKY et al (2014) Asymmetric-detection time-stretch optical microscopy (ATOM) for ultrafast high-contrast cellular imaging in flow. Sci Rep 4:3656. https://doi.org/10.1038/srep03656
Wu L, Chen P, Dong Y et al (2013) Encapsulation of single cells on a microfluidic device integrating droplet generation with fluorescence-activated droplet sorting. Biomed Microdevices 15:553–560. https://doi.org/10.1007/s10544-013-9754-z
Xi H-D, Zheng H, Guo W et al (2017) Active droplet sorting in microfluidics: a review. Lab Chip 17:751–771. https://doi.org/10.1039/C6LC01435F
Yazdi AA, Popma A, Wong W et al (2016) 3D printing: an emerging tool for novel microfluidics and lab-on-a-chip applications. Microfluid Nanofluidics 20:1–18. https://doi.org/10.1007/s10404-016-1715-4
Yukuyama MN, Ghisleni DDM, Pinto TJA, Bou-Chacra NA (2016) Nanoemulsion: process selection and application in cosmetics - a review. Int J Cosmet Sci 38:13–24. https://doi.org/10.1111/ics.12260
Zhang K, Liang Q, Ma S et al (2009) On-chip manipulation of continuous picoliter-volume superparamagnetic droplets using a magnetic force. Lab Chip 9:2992–2999. https://doi.org/10.1039/b906229g
Acknowledgements
We thank Soo Hyeon Kim for his help on measuring the conductivity of cell media used to generate droplets.
Funding
This work was funded by the ImPACT Program of the Council for Science, Technology, and Innovation (Cabinet Office, Government of Japan), JSPS KAKENHI (18K14101), JSPS Core-to-Core Program, Kanagawa Institute of Industrial Science and Technology (KISTEC), White Rock Foundation, Murata Science Foundation, JST Adaptable and Seamless Technology transfer Program (JPMJTM19Y9), and Nakatani Foundation.
Author information
Authors and Affiliations
Corresponding authors
Ethics declarations
Conflict of interest
A. Isozaki, D. Huang, and K. Goda are inventors on a patent application covering the dSADA sorter. K. Goda is a shareholder of CYBO.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Supplementary file1 (AVI 3378 KB)
Supplementary file2 (AVI 7630 KB)
Supplementary file3 (MP4 18263 KB)
Supplementary file4 (MP4 12364 KB)
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Isozaki, A., Huang, D., Nakagawa, Y. et al. Dual sequentially addressable dielectrophoretic array for high-throughput, scalable, multiplexed droplet sorting. Microfluid Nanofluid 25, 32 (2021). https://doi.org/10.1007/s10404-021-02432-z
Received:
Accepted:
Published:
DOI: https://doi.org/10.1007/s10404-021-02432-z