Abstract
Carbohydrate rich substrates such as lignocellulosic hydrolysates remain one of the primary sources of potentially renewable fuel and bulk chemicals. The pentose sugar d-xylose is often present in significant amounts along with hexoses. Saccharomyces cerevisiae can acquire the ability to metabolize d-xylose through expression of heterologous d-xylose isomerase (XI). This enzyme is notoriously difficult to express in S. cerevisiae and only fourteen XIs have been reported to be active so far. We cloned a new d-xylose isomerase derived from microorganisms in the gut of the wood-feeding beetle Odontotaenius disjunctus. Although somewhat homologous to the XI from Piromyces sp. E2, the new gene was identified as bacterial in origin and the host as a Parabacteroides sp. Expression of the new XI in S. cerevisiae resulted in faster aerobic growth than the XI from Piromyces on d-xylose media. The d-xylose isomerization rate conferred by the new XI was also 72% higher, while absolute xylitol production was identical in both strains. Interestingly, increasing concentrations of xylitol (up to 8 g L−1) appeared not to inhibit d-xylose consumption. The newly described XI displayed 2.6 times higher specific activity, 37% lower KM for d-xylose, and exhibited higher activity over a broader temperature range, retaining 51% of maximal activity at 30 °C compared with only 29% activity for the Piromyces XI.
Similar content being viewed by others
Introduction
Lignocellulosic material continues to be the most promising renewable raw material for the production of sustainable fuels and fine chemicals1. Xylan is the second most abundant biopolymer on earth which contains mostly the pentose sugar d-xylose2. Baker’s yeast or Saccharomyces cerevisiae is the preferred organism for industrial transformation of sugars derived from lignocellulose due to innate resistance to fermentation inhibitors3. Expression of heterologous pathways are necessary for d-xylose utilization as it is not metabolized naturally by S. cerevisiae. d-xylose metabolism remains a metabolic bottleneck in S. cerevisiae despite the development of several types of pathways for the consumption of this sugar4.
d-xylose metabolic pathways can be classified into two main categories, d-xylose reductase/xylitol dehydrogenase (XR/XDH) and d-xylose isomerase (XI). The XR/XDH pathway converts d-xylose to xylitol by reduction with NADPH or NADH followed by an oxidation with NAD+ to xylulose in an overall redox neutral process. Alternatively, the same reaction is carried out by a single XI enzyme without cofactors. The XR/XDH pathway is mainly found in fungi while the XI pathway is common in prokaryotes. The currently most promising d-xylose metabolic pathways are based on the prokaryotic XI route. The reason for this is that although the overall reaction is redox neutral, the XR/XDH pathway suffers from a NAD(P)H cofactor imbalance that has proven hard to remedy5,6,7. However, the XI pathway suffers from low capacity and inhibition by xylitol8. Another issue is that the XI is rather difficult to express heterologously in yeast. Several unsuccessful attempts have been made to express XIs, such as those from Escherichia coli9,10 Bacillus subtilis, Actinoplanes missouriensis11, Lactobacillus pentosus12 and Clostridium thermosulfurogenes (renamed as Thermoanaerobacterium thermosulfurigenes)13. The first successfully expressed XI was a thermostable enzyme from Thermus thermophilus14 followed by an XI originating from the fungus Piromyces sp.15.
Only fourteen different XIs with characterized kinetic parameters have been reported to actively express in S. cerevisiae (Table 1). Interestingly, the fungal XIs (Table 1, entry #2 and #3) both come from the same division (Neocallimastigomycota). These fungi are known for possessing genes originating via lateral gene transfer. Their XIs are of prokaryotic origin and have been recently incorporated in evolutionary terms16. These fungal isomerases were isolated from fungi residing in the intestines of herbivorous mammals, an ecosystem typically poised at 37 °C and with dense populations of co-occurring bacteria. Interestingly, only four XIs have been isolated directly from metagenomes and subsequently expressed in S. cerevisiae (Table 1, #5 and #6, #12, #14). Two XIs were derived from soil samples, one using degenerate primers and another by functional metagenomic screening using E. coli with a nonfunctional xylA17. An additional XI was amplified from bovine rumen contents using degenerate primers targeting relatively conserved XI specific sequences18, and yet another derived from protists in the hindgut of the termite Reticulitermes speratus using a similar approach19.
An alternative and less biased approach to retrieving novel XIs is through the assembly of shotgun metagenomic sequences from environments thought to be enriched in d-xylose utilization capacity. Following gene prediction, in-silico translation and annotation against appropriate databases, a subset of XI genes may then be optimized computationally for a specific host, and then synthesized in-vitro. This strategy avoids the bias associated with PCR amplification and is expected to broaden the discovery of enzymes with novel properties. Using this approach, we have identified 182 putative XIs (coded by xylA genes) from the microbial metagenome in the gut of a passalid beetle (Odontotaenius disjunctus)20, known to subsist on d-xylose rich hardwoods, and to harbor d-xylose utilizing yeasts21. We codon optimized and synthesized three XIs, one of which expressed successfully in S. cerevisiae, highlighting that direct metagenome reconstruction combined with in-vitro gene synthesis is a viable approach for the identification of novel XI genes for expression in S. cerevisiae. In the present work, this new XI was characterized and compared to the homolog from Piromyces sp. The d-xylose isomerization capacity of the expressed XIs were evaluated in-vivo by yeast growth and by d-xylose conversion rates. Enzyme kinetics and optimal temperature were determined in-vitro using cell extracts. Additionally, a phylogenetic analysis of the XIs that have been expressed efficiently or that could not be expressed in yeast was carried out to interpret the role of evolutionary relationships in the expression of these enzymes.
Results
Identification of XI genes from metagenomic information
The metagenome of the wood-feeding beetle O. disjunctus, previously reconstructed by our team members20, was screened for the detection of d-xylose isomerases against the pfamA database. A total of 182 putative XI sequences were detected and phylogenetically placed together with the XI of Piromyces (see Fig. S1, Supplementary Information). Based on their phylogenetic relatedness, 3 metagenome-derived XI sequences were selected (labeled 8054_2, 15405_2, and 1362_6 in Fig. S1) and further characterized using the SWISS-MODEL workspace33. Top SWISS-MODEL templates for each of the target amino acid sequences corresponded to d-xylose isomerases for which crystal structures are available (see Table S1, Supplementary Information). Sequences 1362_6 and 14450_2 had 55.9% and 56.7% sequence homology with the XI of Thermotoga neapolitana, respectively, while 8054_2 had 82.5% homology with the XI of Bacteroides thetaiotaomicron. Reconstructed structure-homology models for each sequence also showed high quality scores between target and template (GMQE) and agreement between model and structure (QMEAN) (see Table S1, Supplementary Information). Based on these results, we proceeded to synthesize codon-optimized versions of these xylA genes for expression in S. cerevisiae.
The codon-optimized synthetic genes corresponding to the sequences of 8054_2, 15405_2, 1362_6 and Piromyces sp. xylA were cloned into the plasmid pLBL3 under the control of a constitutive TEF promoter. Plasmid pLBL3_XR/XDH expressing Scheffersomyces stipitis d-xylose reductase (XR) and d-xylitol dehydrogenase (XDH) genes (GenBank Gene ID: 4839234 and 4852013) corresponding to the initial fungal d-xylose metabolic pathway was also constructed. The resulting plasmids were transformed into S. cerevisiae EBY.VW4000. In parallel, a plasmid containing a partial d-xylose utilization pathway (TKL1, TAL1, RPE1 and RKI1), xylulokinase (XKS1), and a d-xylose/glucose facilitator from Candida intermedia (Gxf1), was constructed using the yeast pathway kit34 in the strain CEN.PK111-61A (Fig. 1). Yeast strains containing d-xylose isomerase genes or XR/XDH pathway were mated to the strain with the partial d-xylose utilization pathway. The resulting diploid strains were plated on solid media with d-xylose as the sole carbon source. The strains expressing the XR/XDH gene pair, the Piromyces sp. xylA, and the 8054_2 gene produced growth clearly distinguishable from strains with the partial d-xylose utilization pathway and the empty pLBL3 vector. Strains expressing the 15405_2 and 1362_6 genes showed no discernible growth.
Strategy for the screening of d-xylose isomerases (XIs) in Saccharomyces cerevisiae. (a) Putative xylA genes from the microbial metagenome in the gut of a passalid beetle (Odontotaenius disjunctus) were codon-optimized, synthesized and cloned in the plasmid pLBL3; the partial d-xylose pathway (TKL1, TAL1, RPE1, RKI1, XKS1, and Gxf1) was constructed in the plasmid pYPK0 under the control of different terminators/promoters (TP). (b) The plasmids containing xylA genes (pLBL3_XI) were carried by the strain EBY.VW400 and the plasmid containing the partial d-xylose pathway (pYPK0_XTTRRG) was carried by the strain CEN.PK111-61A. (c) Each EBY.VW4000 strain clone was mated with the CEN.PK111-61A strain. (d) Candidate XI enzymes were functionally screened by scoring yeast growth on solid media with d-xylose as the sole or main carbon source.
Phylogenetic relationship among yeast (un)expressed XIs
Phylogenetic analysis was used to further distinguish the successfully expressed from the unexpressed metagenome derived XIs and to complement the phylogenetic analysis from the SWISS-MODEL (Fig. 2a). For this purpose, we also included sequences of XIs for which activity has been reported in S. cerevisiae to evaluate any evolutionary relationships that might exist between successfully and unsuccessfully expressed enzymes. The sources of XIs included in the phylogenetic analysis were divided in three main prokaryotic phyla: Firmicutes (Fig. 2a—colored green), Proteobacteria (orange), and Bacteroidetes (blue); two other phyla that contain few representatives: Actinobacteria (purple; Streptomyces rubiginosus, Actinoplanes missouriensis, Bifidobacterium longum) and Deinococcus-Thermus (Thermus thermophilus); and one in Plantae (Arabidopsis thaliana). The XIs from the fungi Piromyces sp. and Orpinomyces sp. share 95% identity and were clustered within the Bacteroidetes phylum. T. thermophilus showed 55% and 59% identity with A. missouriensis and S. rubiginosus, respectively, but these three enzymes show very low identity with the other XIs (< 29%). XIs from metagenomic libraries were grouped within the prokaryotic phyla similar to previous reports17,18,19.
Phylogenetic analysis of the amino acid sequences of XIs that have been reported to actively express or not to express in Saccharomyces cerevisiae. Data from patents was not considered in the analysis. (a) In the phylogenetic tree, Firmicutes are colored green, Proteobacteria are colored orange, Bacteroidetes are colored blue, Actinobacteria are colored purple, and XIs from other sources are colored black. Dashed squares highlight two clusters of XIs belonging to Clostridia class (Firmicutes) and Bacteroidia class (Bacteroidetes) that expressed actively in yeast. (b) Ratio of active and inactive XIs expressed in yeast from the taxonomic groups presented in the panel A. GenBank accession numbers of the amino acid sequences are available in Table S2, Supplementary Information.
In the phylogenetic tree, 8054_2 XI was clustered with Bacteroidetes sharing 82% identity with B. thetaiotaomicron, and 78% with Piromyces sp. Sequence 8054_2 was previously reported as belonging to a metagenome assembled-genome (MAG) identified as Parabacteroides sp. which was highly abundant in the mostly anaerobic anterior hindgut of O. disjunctus20. A BLAST analysis also showed that 8054_2 XI shares the highest identity with XIs from Bacteroidetes phylum, particularly with the Porphyromonadaceae family of the Bacteroidia class. Porphyromonadaceae are commonly found in the gastrointestinal tract and oral cavity of animals35. The 8054_2 XI also shares high identity with the Bacteroides genus, such as Bacteroides timonensis (83%, accession number WP_044271094.1) that is an obligate anaerobe isolated from human feces36. The 15405_2 XI and 1362_6 XI share 94% identity and exhibit low similarity with the other XIs displayed in the tree. A BLAST analysis revealed high identity of both 15405_2 XI and 1362_6 XI with XIs from Leuconostoc genus of the Firmicutes phylum (85% and 86%; accession numbers WP_042252435.1, WP_004912802.1, respectively).
The expression success of XIs in yeast reveals a positive tendency towards the Bacteroidetes phylum (Fig. 2b). The majority of the reported active XIs in yeast are from Bacteroidetes (12 of 21), and most of the XIs from this phylum have been expressed successfully (12 of 13). The only XI belonging to Bacteroidetes that was not expressed in yeast is from Robiginitalea biformata that, coincidentally, is also the most dissimilar enzyme of this phylum, being positioned quite distant from the Bacteroidetes cluster. Inactive XIs (indicated by a red “X”) are comprised mostly of the other phyla: Proteobacteria (6 of 10), Firmicutes (7 of 11), and Actinobacteria (3 of 3). The XIs from Proteobacteria that have been successfully expressed in yeast (Rhizobium radiobacter, Burkholderia cenocepacia, Sorangium cellulosum) belong to different classes and do not share relevant ecological traits in the context of this work. The active XIs from Firmicutes (Clostridium phytofermentans, Clostridium cellulovorans, Ruminococcus flavefaciens) are from anaerobic cellulolytic organisms that belong to Clostridia class (Fig. 2a—Firmicutes’ dashed square). Similarly, actively expressed XIs clustered with Bacteroidetes are from anaerobic species that are commonly associated with the gastrointestinal tract of animals and belong to the same Bacteroidia class (Bacteroidetes’ dashed square), while the inactive XI belongs a different class (Flavobacteriia)37.
Growth on d-xylose under aerobic conditions
Growth performance of yeast diploids carrying the genes 8054_2, xylA from Piromyces sp. (opt.PiXI), and XR/XDH, or an empty vector was assessed in synthetic medium with 2% (w/v) d-xylose as the sole carbon source under aerobic conditions. The results of the growth are shown in Fig. 3. The negative control strain (Fig. 3—Empty vector), lacking XI or XR/XDH enzymes but expressing the partial d-xylose utilization pathway, did not proliferate. The cultures expressing enzymes that convert d-xylose to xylulose grew over time. The strain expressing XR/XDH proliferated at a maximal specific growth rate of 0.10 h−1, the strain expressing 8054_2 XI grew at 0.06 h−1, and the strain expressing opt.PiXI grew at 0.04 h−1. The XR/XDH pathway provided a considerably higher growth rate than the XI pathways as expected, and remarkably, 8054_2 XI conferred 50% faster growth in yeast than opt.PiXI.
Aerobic growth of diploid Saccharomyces cerevisiae strains expressing 8054_2 XI, the codon-optimized XI from Piromyces sp. E2 (opt.PiXI), XR/XDH, or the empty vector. Cultures were grown in shaking-flasks on synthetic medium with 2% (w/v) d-xylose as the sole carbon source. Data points represent an average of 3 biological replicates with standard deviation indicated.
d-Xylose consumption under high cell density cultures
Cultures were inoculated at a high cell density (~ 10 g L−1 DCW) in medium with 4% (w/v) d-xylose as the sole carbon source in shake-flasks at 1:20 liquid-to-air ratio. Both cultures started consuming d-xylose immediately after incubation and showed a steady d-xylose conversion until the end of the experiment (Fig. 4). The strain carrying 8054_2 XI consumed d-xylose at a rate of 0.015 g d-xylose h−1 g(DCW)−1, and the strain carrying opt.PiXI showed a consumption rate of 0.010 g d-xylose h−1 g(DCW)−1. The strain carrying XR/XDH depleted 96% of d-xylose after 18 h, corresponding to at least 0.187 g d-xylose h−1 g(DCW)−1 (see Fig. S2, Supplementary Information). The higher growth rate and d-xylose consumption conferred by the XR/XDH pathway compared to the XI pathway is consistent with previous work38,39. Xylitol was produced throughout the cultivation with a yield of 0.33 g xylitol g−1 d-xylose consumed for 8054_2 XI, and 0.47 g xylitol g−1 d-xylose consumed for opt.PiXI. Therefore, excluding d-xylose converted to xylitol rather than metabolized by the XI system, the rate of d-xylose isomerization was 0.0098 g d-xylose h−1 g(DCW)−1 for 8054_2 XI, and 0.0057 g d-xylose h−1 g(DCW)−1 for opt.PiXI. This represents a 72% higher d-xylose isomerization by 8054_2 XI than opt.PiXI. Absolute xylitol production was identical for both strains (~ 8 g L−1 after 138 h) due to the background activity of aldose reductases (such as GRE3) that reduce d-xylose to xylitol40. Xylitol has been shown to inhibit XI activity in-vitro8,41, although in our experiments, xylitol appears not to affect d-xylose consumption over time as it appears linear for the duration of the experiment for both strains.
The d-xylose consumption by diploid Saccharomyces cerevisiae strains expressing 8054_2 XI or the codon-optimized XI from Piromyces sp. E2 (opt.PiXI). Cultures were grown under high cell density (10 g L−1 DCW) in shake flasks on synthetic medium with 4% (w/v) d-xylose as the sole carbon source. Data points represent an average of 3 biological replicates with standard deviation indicated.
Kinetic parameters and optimal temperature
Kinetic properties of the enzymes 8054_2 XI and opt.PiXI were determined using crude cell extracts prepared from the recombinant diploid strains by a bead beating method. There seem to be no significant differences in the apparent protein expression level between both XIs in the cell extracts (see Fig. S3, Supplementary Information). A Michaelis–Menten curve was adjusted to the experimental values of both enzymes with close fit (Fig. 5). The extracts containing 8054_2 XI converted d-xylose at a maximal rate (Vmax) of 0.0296 U mg−1 total protein, and the extracts containing opt.PiXI showed a Vmax of 0.0112 U mg−1 total protein. The 8054_2 XI exhibited a KM for d-xylose of 19.27 ± 0.74 mM, and opt.PiXI exhibited a KM of 30.76 ± 2.43 mM. Thus, the comparative analysis presented in this study indicates that 8054_2 XI operates at a 2.6 times higher Vmax and with 37% lower KM for d-xylose than opt.PiXI. We also assessed optimal enzyme temperature using a discontinuous colorimetric assay to measure relative activity of XIs in the cell extracts at different temperatures. The optimal temperatures of 8054_2 XI and opt.PiXI were approximately 60 °C and 65 °C, respectively (Fig. 6). Relative activity of both enzymes shows a steady increase from 30 °C, i.e., yeast optimum growth temperature, to the respective peak (100% activity) followed by a strong decrease to the lowest activity at 80 °C. Despite the similar thermophilic behavior, 8054_2 XI retains 51% of the maximal activity at 30 °C, while activity of opt.PiXI was reduced to 29%.
Enzymatic specific activity of the XI enzymes 8054_2 XI or the codon-optimized XI from Piromyces sp. E2 (opt.PiXI). Kinetic parameters were determined from a Michaelis–Menten plot adjusted to the experimental data (R squared > 0.99). Strains were grown in synthetic medium containing 2% (w/v) d-xylose until the exponential growth phase. Crude cell extracts were prepared and used immediately in the activity assays. Data points represent an average of 3 biological replicates.
Relative enzymatic activity of XI enzymes 8054_2 XI and the codon-optimized XI from Piromyces sp. E2 (opt.PiXI) at different temperatures. Strains were grown in synthetic medium containing 2% (w/v) d-xylose until the exponential growth phase. Crude cell extracts were prepared and used immediately in the activity assays. Data points represent an average of 3 biological replicates with standard deviation indicated.
Discussion
Novel metagenomic sequence data as well as sequence data in public databases represent a largely untapped resource for novel catalytic capability for future biotechnological processes. Making use of these data requires accurate assembly of sequence reads, gene identification followed by gene synthesis. In this work, we searched metagenomic data we derived from the hindgut of a wood-feeding beetle20 for similarity to known products of d-xylose isomerases genes, which depends on correctly translating metagenome sequence data in-silico. Gene synthesis remains a comparatively expensive approach relative to PCR amplification of DNA from complex extracts, with synthesis cost, time, and error rates increasing with sequence length. However, as well as eliminating PCR bias, an added benefit of gene synthesis is that its codon usage can be optimized for a specific host. Despite the expense, the use of such optimized genes is growing in popularity as evidenced by the twenty percent of sequences deposited at Addgene being composed of codon optimized genes42. Although some XI enzymes have been successfully expressed in S. cerevisiae allowing more or less efficient d-xylose metabolism, the screening for XIs with significant expression in yeast has proved difficult and often require a suite of optimizations8,14,15,43. Our strategy used here allowed direct selection of one very efficient XI from three synthesized candidate genes chosen from the three closest homologs to a known gene without any further evolutionary adaptation of the S. cerevisiae host. The separation of XI and the other genes necessary for efficient growth on d-xylose facilitate high-throughput functional screening which would enable testing of many more genes when gene synthesis costs are more permissive44.
d-xylose isomerases have been found in lignocellulosic material such as decomposing biomass, the gut of animals or their feces15,17,18,19 presumably due to the opportunity to consume d-xylose. Little is known about the relevance of phylogenetic relationship of different XIs for their efficient expression in S. cerevisiae, or whether certain microbial groups contain a pool of XIs with higher probability of being expressed in yeast. One of three of the XI enzymes screened in this work was successfully expressed (8054_2 XI) and clustered within the Bacteroidetes phylum (Fig. 2a). Curiously, all the previously assayed XIs from Bacteroidia and Clostridia (with exception to T. thermosulfurigenes) exhibited considerable activity in yeast (Fig. 2a—dashed squares). These classes of organisms have crucial ecological traits in common, such as an anaerobic metabolism and a remarkable capacity for degrading polysaccharides37,45,46,47. This indicates that Bacteroidia and Clostridia, particularly species residing in niches where active degradation of lignocellulosic biomass could be considered a selective pressure, may represent a valuable source of new XIs that express efficiently in yeast.
The growth rates on d-xylose presented in this work are comparable to others using non-evolved S. cerevisiae CEN.PK-based strains with identical genetic modifications. Comparatively, 8054_2 XI produced 50% higher growth rates and 72% higher d-xylose isomerization than the XI from Piromyces sp. under identical conditions, which indicates that the 8054_2 XI is an improved option for metabolic engineering of S. cerevisiae to transform d-xylose. Nevertheless, identical strains carrying the XI from Piromyces sp. have exhibited very distinct growth rates, which vary from 0.02 to 0.07 h−18,17,38, or 0.21 to 0.22 h−124,43. The growth rates conferred by the XI enzymes expressed in this work are closer to the first group of values. The second group is among the highest specific growth rates on d-xylose reported in literature for any XI or evolved strain22,23,32,48,49. This discrepancy may be attributed to a wide range of factors including unknown beneficial genetic modifications.
Xylitol has been identified as a competitive inhibitor of XIs41. Interestingly, contradictory results on xylitol inhibition have been reported. According to in-vitro assays, XI from Piromyces sp. is strongly inhibited in-vitro at a concentration of 50 mM xylitol, and an inhibition constant of 4.6 mM was determined8. Similarly, XI from Bacteroides stercoris, with an inhibition constant of 5.1 mM xylitol, showed a 50% decrease in activity at a concentration of 30 mM xylitol, although the fermentation rate only declined by less than 10%28. In our study, xylitol had no noticeable effect on d-xylose isomerization by either enzyme even at a concentration of approximately 50 mM (8 g L−1).
The kinetic parameters of the XI from Piromyces sp. reported in literature are very dissimilar, reflecting the vast experimentation with this enzyme (Table 1, entry #2). To overcome this limitation, in our study, 8054_2 XI and opt.PiXI were compared under identical conditions to eliminate this source of uncertainty. The 8054_2 XI showed significantly higher specific activity and affinity relative to opt.PiXI, both correlated with higher relative activity at 30 °C. As most of the XIs expressed in yeast are thermophilic, enzyme activity is dramatically reduced at industrial temperatures of fermentation that vary from 30 to 40 °C50. Only one yeast-expressed XI in the mesophilic range has been described to date26, with an optimal temperature of 37 °C and retaining 71% of activity at 30 °C, more than twice that we observed for opt.PiXI at 30 °C. We hypothesize that a mesophilic profile of XIs may be a major feature for efficient d-xylose utilization by S. cerevisiae. Although the 8054_2 XI we describe here has a thermophilic temperature optimum, we note its activity over a broad thermal range that extends into mesophilic temperatures. This broad range of temperature activity is perhaps not surprising given its origin from the digestive tract of an invertebrate that cannot regulate its body temperature (i.e. is an ectotherm) unlike the endothermic (elephant) habitat of Piromyces sp. strain E2 from which opt.PiXI was derived51.
To our knowledge, six studies have experimentally compared the XI from Piromyces sp. to other XIs. One XI displayed higher affinity but lower specific activity19, and another exhibited lower affinity and activity but also lower inhibition by xylitol8. A further report identified an enzyme with lower affinity but higher specific activity and another with lower affinity and specific activity18, while two additional studies described enzymes with lower specific activity17,32. Only one new XI showed higher affinity and specific activity relative to the Piromyces sp. XI, which could be further enhanced by evolutionary adaptation23. However, as the XI from Piromyces sp. was not codon optimized, the comparison is hard to interpret directly. Our study reveals, for the first time, a new XI that exhibits clearly superior kinetics than the XI from Piromyces sp. E2 which translates into faster yeast growth and d-xylose consumption in-vivo.
Conclusions
We have synthesized three putative xylA genes from the gut microbiota of a wood-feeding beetle and screened for d-xylose isomerase activity in S. cerevisiae. One of these enzymes, 8054_2 XI, expressed actively and showed high identity with XIs from the Bacteroidia class of the Bacteroidetes phylum. Curiously, the phylogenetic analysis revealed that all XIs from Bacteroidia screened in yeast have expressed successfully. The new enzyme showed higher specific activity and affinity for d-xylose than the current gold-standard from Piromyces sp., as well as substantially higher relative activity at 30 °C. Superior kinetics of 8054_2 XI correlated with higher specific growth rate and d-xylose consumption. The novel XI represents a highly valuable addition to the S. cerevisiae molecular toolbox and shows promise for improved industrial conversion of carbohydrate substrates.
Methods
Selection of XI from the metagenome of O. disjunctus
Previously published predicted genes and proteins from the metagenome of O. disjunctus20 were screened against the pfamA-30 database52 using HMMER’s hmmscan53. Putative metagenome-predicted XI amino acid sequences were aligned together with the XI of Piromyces sp. using ClustalW and a phylogenetic tree reconstructed with IQ-TREE using the evolutionary model LG + G4. After the phylogenetic analysis, three sequences were selected and further characterized in the SWISS-MODEL workspace33 against the SWISS MODEL template library54 and the obtained models evaluated based on amino acid sequence identity, Global Model Quality Estimation (GMQE, evaluated from 0 to 1), and the QMEAN Z-score55 (evaluated from − 4 to 0).
Plasmid constructions
A detailed description of all vectors constructed as a part of this work is available in the form of a collection of Jupyter notebooks56. These notebooks contain Python code describing the details of the cloning with the help of the pydna package57. These notebooks allow reexamination of the cloning strategies in detail by executing the code. The executable documentation is available in a Git repository (https://github.com/MetabolicEngineeringGroupCBMA/Silva_et_al_2020). The plasmid pLBL3 was used to express candidate d-xylose isomerase genes. The pLBL3 is an expression vector with a LEU2 auxotrophic marker with both 2µ and pUC origins of replication. The expressed gene is controlled upstream by a TEF1 promoter (the intergenic sequence between MRL1/YPR079W and TEF1/YPR080W) and downstream by the intergenic sequence between YNL095C and RPS7B/YNL096C. Codon-optimized XI genes were synthesized by Integrated DNA Technologies, Inc (Coralville, IA, USA) and were cloned into pLBL3 by in-vivo gap repair between tailed PCR products of individual genes and the plasmid linearized with AjiI (Thermo Fisher Scientific Inc, Waltham, MA, USA) (Fig. 1a) in S. cerevisiae EBY.VW4000.
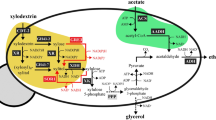
The S. cerevisiae CEN.PK111-61A was transformed with the pYPK0_XTTRRG vector (Fig. 1b) expressing a partial d-xylose utilization pathway. The vector expresses six different genes, a xylulokinase (XKS1), that converts d-xylulose to xylulose 5-phosphate, a d-xylose/glucose facilitator from Candida intermedia (Gxf1), and the four genes of the non-oxidative pentose phosphate pathway TKL1, TAL1, RPE1 and RKI1. TKL1 codes for a transketolase that convert xylulose-5-phosphate and ribose-5-phosphate to sedoheptulose-7-phosphate and glyceraldehyde-3-phosphate; TAL1 codes for a transaldolase that converts sedoheptulose 7-phosphate and glyceraldehyde 3-phosphate to erythrose 4-phosphate and fructose 6-phosphate; RPE1 codes for a d-ribulose-5-phosphate 3-epimerase that converts d-ribulose 5-phosphate to d-xylulose 5-phosphate. RKI1 codes for a ribose-5-phosphate ketol-isomerase that interconverts ribose 5-phosphate and ribulose 5-phosphate. Yeast transformations were performed as described by the high-efficiency protocol using lithium acetate, ssDNA and polyethylene glycol 335058.
Strains and cultivation
Escherichia coli strain XL1-Blue (Stratagene, La Jolla, CA, USA) was used for routine plasmid preparation. E. coli strains were cultivated on lysogeny broth (LB-Lennox) containing 1% (w/v) tryptone (BD biosciences, San Jose, CA, USA), 0.5% (w/v) yeast extract (Panreact AppliChem, Darmstadt, Germany), 0.5% (w/v) sodium chloride and 100 mg L−1 ampicillin (Formedium, King's Lynn, UK). S. cerevisiae strains were cultivated on complex media containing 2% (w/v) bacto-peptone (BD biosciences, San Jose, CA, USA), 1% (w/v) yeast extract, and 2% (w/v) glucose (YPD), maltose (YPM), or d-xylose (YPX). Yeast strains were also cultivated in defined synthetic complete media (SC) containing 0.67% (w/v) yeast nitrogen base without amino acids (BD, Franklin Lakes, NJ, USA), 0.07% amino acid dropout Kaiser mixture59, 50 mM potassium hydrogen phthalate, and 2% (w/v) glucose, maltose or d-xylose. SC media were adjusted to pH 5.5 using sodium hydroxide. Amino acids histidine, uracil and tryptophan were omitted as required for selection of auxotrophic markers. Agar was added to a concentration of 2% (w/v) for solid media. Liquid cultures were incubated with shaking at 200 rpm, yeast at 30 °C and E. coli at 37 °C. S. cerevisiae strains and plasmids used in this work are listed in Table 2.
Yeast mating
Each EBY.VW4000 (MATa) strain carrying a pLBL3_XI plasmid was mated with the CEN.PK111-61A/pYPK0_XTTRRG (MATα) strain (Fig. 1c), resulting in diploid yeast strains. Briefly, EBY.VW4000 (pLBL3_XI) and CEN.PK111-61A (pYPK0_XTTRRG) were cultivated in 5 mL YPD medium overnight. Cultures were then diluted in 5 mL YPD medium to an OD600 of 0.2 and grown for at least two generations, to a final OD600 of 0.8–1.2. Cells were harvested by centrifugation and transferred to 1-mL Eppendorf tube. Cell suspensions were washed twice with deionized water and resuspended in 1 mL rich medium. A hundred microliters of each culture were pooled, mixed and incubated overnight at room temperature. The pooled cultures were washed twice in 1 mL deionized water and resuspended in 1 mL water. A hundred microliters of the washed cultures were spread on SC medium. The resulting diploid clones were collected after two days.
Screening for growth on solid medium
Screening was performed by scoring yeast growth on solid media with d-xylose as the sole, or main, carbon source (Fig. 1d). Recombinant diploids were streaked into 5 mL SC medium containing glucose, lacking leucine, uracil and tryptophan and incubated overnight. Subsequently, 1 mL of culture was washed twice and diluted 1:10 in deionized water. Cell suspensions were incubated at 30 °C overnight and then placed on ice for 2 h. Cultures were serially diluted 1:10 in deionized water. Drops with a volume of 10 µL of dilutions 1, 10–1 and 10–2 were spotted on YPX and SC media with d-xylose. Plates were incubated for 5 days before scoring growth.
Liquid culture growth assays
Individual colonies were used to inoculate SC media containing glucose and grown overnight. Yeast cells were washed and transferred to pre-warmed (30 °C) SC medium containing d-xylose at an OD600 of 0.2 and incubated in 50-mL glass tubes for 24 h. Cells were then transferred to 5 mL identical fresh medium in 50-mL shaking-flasks at an OD600 of 0.2 to start measurements. Absorbance was measured using a NanoDrop 1000 spectrophotometer. Cell growth experiments were performed in triplicate.
d-Xylose consumption
For measuring specific d-xylose consumption, cells from frozen stocks (− 80 °C) were transferred into 50 mL SC media containing glucose and incubated in 500-mL culture flasks for 24 h. Cells were harvested, washed and suspended in 25 mL identical medium containing 4% (w/v) d-xylose, instead of glucose. The inoculation level was 10 g L−1 dry cell weight (DCW). Cultivation was performed in 500-mL flasks and samples were taken every 24 h. d-xylose and xylitol concentrations were quantified by high performance liquid chromatography (Hitachi LaChrom Elite) and detection was done by refractive index with an Elite LaChrom L-2490 RI detector (VWR Hitachi). Samples were prepared by adding 125 µL trichloroacetic acid to a final concentration of 10% (v/v) followed by a 2 h incubation on ice. Samples were centrifuged at 4 °C for 15 min and filtered using a 0.22 µm pore hydrophilic PTFE filter. Samples were separated on a Rezex 8 μm ROA-organic acid H+(8%) column (Phenomenex) with a mobile phase of 2.5 mM H2SO4, and a flow rate of 0.5 mL min−1 at 40 °C. Data was analyzed with the EZChrom Elite 3.3.2 SP2 software. d-xylose consumption experiments were performed in triplicate.
d-xylose isomerase activity assay
Yeast cells expressing XI enzymes (Piromyces sp., 8054_2) were grown overnight in SC media containing glucose. Cultures were diluted in 50 mL same media at an OD600 of 0.3 and incubated for three generations. Cells were harvested, washed twice with water and suspended in 100 mM Tris–HCl buffer pH of 7.5, followed by disruption with glass beads (0.45 mm) using FastPrep FP120 cell disrupter (6.0 oscillations min−1 for 20 s). Cell debris was removed by centrifugation at 16.000×g for 10 min and the supernatant was conserved on ice. Crude cell extracts were used in the enzyme assays immediately after preparation. Protein concentration was measured by the Bradford assay having bovine serum albumin as standard. All of the enzyme assays were performed in triplicate.
Resorcinol method
Resorcinol-based activity assay was performed in this work for determination of optimal temperature of activity of the XI enzymes62,63. Cell extracts were diluted in 100 mM Tris–HCl buffer at pH of 7.5. Experiments were carried out by adding 30 µL cell extracts containing 3 mg mL−1 total protein to 20 µL test solution (100 mM Tris–HCl at pH of 7.5, 30 mM d-xylose, 0.3 mM MnSO4), and then incubated at the testing temperature for an appropriate period of time previously optimized. Sample incubation was carried out into BioRad T100 thermal cycler set at specific temperatures. For color development, 150 µL of a 1:1 mixture (v/v) of solution A (0.05% resorcinol in ethanol) and solution B (0.216 g FeNH4(SO4)2 · 12 H2O in 1 L concentrated HCl) were added to the samples followed by incubation at 80 °C for 40 min. The absorbance measurements were determined on the microplate reader SpectraMax Plus 384 (Molecular Devices) at 630 nm.
d-Sorbitol dehydrogenase (SDH) method
The SDH method was employed to determine specific activity of the XIs. Assays were performed in mixtures containing 100 mM Tris–HCl (pH value of 7.5), 10 mM MgCl2, 0.15 mM NADH, 2 U SDH (Roche Diagnostics, Mannheim, Germany), and 1 mg mL−1 cell extract15. The reactions started with the addition of the mixture to d-xylose solutions that ranged from 5 to 400 mM. The reactions occurred at 30 °C and were monitored spectrophotometrically (SpectraMax Plus 384, Molecular Devices) at 340 nm through the depletion of NADH resulting from the reduction of xylulose to xylitol by SDH. Kinetic parameters were determined from the interpolation of the experimental data to the Michaelis–Menten curve by the least squares fit method using Graphpad Prism 6.
Phylogenetic analysis
The amino acid sequences from the metagenome-derived sequences and those from reported studies were aligned using Clustal Omega with default parameters64. This software generated a percent identity matrix that was used to compare similarity among the sequences of the enzymes, and an alignment file in “ClustalW” format for phylogenetic tree construction. This file was converted to a usable “Phylip” format65. The phylogenetic trees were constructed using PhyML 3.0 on the web server ATGC-Montpellier, using WAG substitution model, mix of NNI/SPR improvements, 8 substitution rate categories, and 1000 bootstrap replicates66. The resulting trees were visualized and edited with the on-line software “Interactive Tree of Life”—iTOL v467. GenBank accession numbers of the amino acid sequences used to generate the phylogenetic tree are available in Table S2, Supplementary Information.
Data availability
All data generated or analyzed during this study are included in this published article, public repositories, or in the Supplementary Information, as indicated in the manuscript.
References
Arevalo-Gallegos, A., Ahmad, Z., Asgher, M., Parra-Saldivar, R. & Iqbal, H. M. N. Lignocellulose: A sustainable material to produce value-added products with a zero waste approach-A review. Int. J. Biol. Macromol. 99, 308–318 (2017).
Dodd, D. & Cann, I. K. O. Enzymatic deconstruction of xylan for biofuel production. Glob. Change Biol. Bioenergy 1, 2–17 (2009).
Cunha, J. T., Romaní, A., Costa, C. E., Sá-Correia, I. & Domingues, L. Molecular and physiological basis of Saccharomyces cerevisiae tolerance to adverse lignocellulose-based process conditions. Appl. Microbiol. Biotechnol. 103, 159–175 (2019).
Moysés, D. N., Reis, V. C. B., de Almeida, J. R. M., de Moraes, L. M. P. & Torres, F. A. G. Xylose fermentation by Saccharomyces cerevisiae: challenges and prospects. Int. J. Mol. Sci. 17, 207 (2016).
Kötter, P., Amore, R., Hollenberg, C. P. & Ciriacy, M. Isolation and characterization of the Pichia stipitis xylitol dehydrogenase gene, XYL2, and construction of a xylose-utilizing Saccharomyces cerevisiae transformant. Curr. Genet. 18, 493–500 (1990).
Kötter, P. & Ciriacy, M. Xylose fermentation by Saccharomyces cerevisiae. Appl. Microbiol. Biotechnol. 38, 776–783 (1993).
Wahlbom, C. F., Eliasson, A. & Hahn-Hägerdal, B. Intracellular fluxes in a recombinant xylose-utilizing Saccharomyces cerevisiae cultivated anaerobically at different dilution rates and feed concentrations. Biotechnol. Bioeng. 72, 289–296 (2001).
Brat, D., Boles, E. & Wiedemann, B. Functional expression of a bacterial xylose isomerase in Saccharomyces cerevisiae. Appl. Environ. Microbiol. 75, 2304–2311 (2009).
Briggs, K. A., Lancashire, W. E. & Hartley, B. S. Molecular cloning, DNA structure and expression of the Escherichia coli D-xylose isomerase. EMBO J. 3, 611–616 (1984).
Sarthy, A. V. et al. Expression of the Escherichia coli xylose isomerase gene in Saccharomyces cerevisiae. Appl. Environ. Microbiol. 53, 1996–2000 (1987).
Amore, R., Wilhelm, M. & Hollenberg, C. P. The fermentation of xylose: An analysis of the expression of Bacillus and Actinoplanes xylose isomerase genes in yeast. Appl. Microbiol. Biotechnol. 30, 351–357 (1989).
Hallborn, J. Metabolic Engineering of Saccharomyces cerevisiae: Expression of Genes Involved in Pentose Metabolism (Lund University, Lund, 1995).
Moes, C. J., Pretorius, I. S. & van Zyl, W. H. Cloning and expression of the Clostridium thermosulfurogenes D-xylose isomerase gene (xylA) in Saccharomyces cerevisiae. Biotechnol. Lett. 18, 269–274 (1996).
Walfridsson, M. et al. Ethanolic fermentation of xylose with Saccharomyces cerevisiae harboring the Thermus thermophilus xylA gene, which expresses an active xylose (glucose) isomerase. Appl. Environ. Microbiol. 62, 4648–4651 (1996).
Kuyper, M. et al. High-level functional expression of a fungal xylose isomerase: the key to efficient ethanolic fermentation of xylose by Saccharomyces cerevisiae?. FEMS Yeast Res. 4, 69–78 (2003).
Murphy, C. L. et al. Horizontal gene transfer as an indispensable driver for evolution of neocallimastigomycota into a distinct gut-dwelling fungal lineage. Appl. Environ. Microbiol. 85, 15. https://doi.org/10.1128/AEM.00988-19 (2019).
Parachin, N. S. & Gorwa-Grauslund, M. F. Isolation of xylose isomerases by sequence- and function-based screening from a soil metagenomic library. Biotechnol. Biofuels 4, 9 (2011).
Hou, J. et al. Characterization and evolution of xylose isomerase screened from the bovine rumen metagenome in Saccharomyces cerevisiae. J. Biosci. Bioeng. https://doi.org/10.1016/j.jbiosc.2015.05.014 (2015).
Katahira, S. et al. Screening and evolution of a novel protist xylose isomerase from the termite Reticulitermes speratus for efficient xylose fermentation in Saccharomyces cerevisiae. Biotechnol. Biofuels 10, 203 (2017).
Ceja-Navarro, J. A. et al. Gut anatomical properties and microbial functional assembly promote lignocellulose deconstruction and colony subsistence of a wood-feeding beetle. Nat. Microbiol. 4, 864–875 (2019).
Suh, S.-O., McHugh, J. V., Pollock, D. D. & Blackwell, M. The beetle gut: A hyperdiverse source of novel yeasts. Mycol. Res. 109, 261–265 (2005).
Lee, S.-M., Jellison, T. & Alper, H. S. Directed evolution of xylose isomerase for improved xylose catabolism and fermentation in the yeast Saccharomyces cerevisiae. Appl. Environ. Microbiol. 78, 5708–5716 (2012).
Hector, R. E., Dien, B. S., Cotta, M. A. & Mertens, J. A. Growth and fermentation of D-xylose by Saccharomyces cerevisiae expressing a novel D-xylose isomerase originating from the bacterium Prevotella ruminicola TC2-24. Biotechnol. Biofuels 6, 84 (2013).
Verhoeven, M. D. et al. Mutations in PMR1 stimulate xylose isomerase activity and anaerobic growth on xylose of engineered Saccharomyces cerevisiae by influencing manganese homeostasis. Sci. Rep. 7, 46155 (2017).
Lee, M., Rozeboom, H. J., Keuning, E., de Waal, P. & Janssen, D. B. Structure-based directed evolution improves S. cerevisiae growth on xylose by influencing in vivo enzyme performance. Biotechnol. Biofuels 13, 5 (2020).
Madhavan, A. et al. Xylose isomerase from polycentric fungus Orpinomyces: gene sequencing, cloning, and expression in Saccharomyces cerevisiae for bioconversion of xylose to ethanol. Appl. Microbiol. Biotechnol. 82, 1067–1078 (2009).
Seike, T. et al. Molecular evolutionary engineering of xylose isomerase to improve its catalytic activity and performance of micro-aerobic glucose/xylose co-fermentation in Saccharomyces cerevisiae. Biotechnol. Biofuels 12, 139 (2019).
Ha, S.-J., Kim, S. R., Choi, J.-H., Park, M. S. & Jin, Y.-S. Xylitol does not inhibit xylose fermentation by engineered Saccharomyces cerevisiae expressing xylA as severely as it inhibits xylose isomerase reaction in vitro. Appl. Microbiol. Biotechnol. 92, 77–84 (2011).
Aeling, K. A. et al. Co-fermentation of xylose and cellobiose by an engineered Saccharomyces cerevisiae. J. Ind. Microbiol. Biotechnol. 39, 1597–1604 (2012).
de Figueiredo Vilela, L. et al. Functional expression of Burkholderia cenocepacia xylose isomerase in yeast increases ethanol production from a glucose-xylose blend. Bioresour. Technol. 128, 792–796 (2013).
Vieira, I. P. V. et al. Understanding xylose isomerase from Burkholderia cenocepacia: Insights into structure and functionality for ethanol production. AMB Express 9, 73 (2019).
Peng, B., Huang, S., Liu, T. & Geng, A. Bacterial xylose isomerases from the mammal gut Bacteroidetes cluster function in Saccharomyces cerevisiae for effective xylose fermentation. Microb. Cell Fact. 14, 70 (2015).
Waterhouse, A. et al. SWISS-MODEL: Homology modelling of protein structures and complexes. Nucleic Acids Res. 46, W296–W303 (2018).
Pereira, F. et al. Yeast pathway kit: A method for metabolic pathway assembly with automatically simulated executable documentation. ACS Synth. Biol. 5, 386–394 (2016).
Sakamoto, M. The family Porphyromonadaceae. In The Prokaryotes: Other Major Lineages of Bacteria and The Archaea (eds Rosenberg, E. et al.) 811–824 (Springer, Berlin Heidelberg, 2014).
Ramasamy, D. et al. Genome sequence and description of Bacteroides timonensis sp. nov. Stand. Genom. Sci. 9, 1181–1197 (2014).
Thomas, F., Hehemann, J.-H., Rebuffet, E., Czjzek, M. & Michel, G. Environmental and gut bacteroidetes: The food connection. Front. Microbiol. 2, 93 (2011).
Karhumaa, K., Sanchez, R. G., Hahn-Hägerdal, B. & Gorwa-Grauslund, M.-F. Comparison of the xylose reductase-xylitol dehydrogenase and the xylose isomerase pathways for xylose fermentation by recombinant Saccharomyces cerevisiae. Microb. Cell Fact. 6, 5 (2007).
Li, X. et al. Comparison of xylose fermentation by two high-performance engineered strains of Saccharomyces cerevisiae. Biotechnol. Rep. 9, 53–56 (2016).
Träff, K. L., Otero Cordero, R. R., van Zyl, W. H. & Hahn-Hägerdal, B. Deletion of the GRE3 aldose reductase gene and its influence on xylose metabolism in recombinant strains of Saccharomyces cerevisiae expressing the xylA and XKS1 genes. Appl. Environ. Microbiol. 67, 5668–5674 (2001).
Yamanaka, K. Inhibition of D-xylose isomerase by pentitols and D-lyxose. Arch. Biochem. Biophys. 131, 502–550 (1969).
Kunjapur, A. M., Pfingstag, P. & Thompson, N. C. Gene synthesis allows biologists to source genes from farther away in the tree of life. Nat. Commun. 9, 4425 (2018).
Kuyper, M. et al. Metabolic engineering of a xylose-isomerase-expressing Saccharomyces cerevisiae strain for rapid anaerobic xylose fermentation. FEMS Yeast Res. 5, 399–409 (2005).
Carlson, R. On DNA and Transistors. Synthesis. http://www.synthesis.cc/synthesis/2016/03/on_dna_and_transistors (2016).
Warnick, T. A. Clostridium phytofermentans sp. nov., a cellulolytic mesophile from forest soil. Int. J. Syst. Evol. Microbiol. 52, 1155–1160 (2002).
Flint, H. J., Bayer, E. A., Rincon, M. T., Lamed, R. & White, B. A. Polysaccharide utilization by gut bacteria: potential for new insights from genomic analysis. Nat. Rev. Microbiol. 6, 121–131 (2008).
Doi, R. H. & Tamaru, Y. The Clostridium cellulovorans cellulosome: an enzyme complex with plant cell wall degrading activity. Chem. Rec. 1, 24–32 (2001).
Shen, Y. et al. An efficient xylose-fermenting recombinant Saccharomyces cerevisiae strain obtained through adaptive evolution and its global transcription profile. Appl. Microbiol. Biotechnol. 96, 1079–1091 (2012).
Lee, S.-M., Jellison, T. & Alper, H. S. Systematic and evolutionary engineering of a xylose isomerase-based pathway in Saccharomyces cerevisiae for efficient conversion yields. Biotechnol. Biofuels 7, 122 (2014).
Carlos, L., Olitta, T. & Nitsche, S. Ethanol production in brazil: the industrial process and its impact on yeast fermentation. In Biofuel Production-Recent Developments and Prospects (ed. Dos Santos Bernardes, M. A.) (InTech, Rijeka, 2011).
Harhangi, H. R. et al. Xylose metabolism in the anaerobic fungus Piromyces sp. strain E2 follows the bacterial pathway. Arch. Microbiol. 180, 134–141 (2003).
El-Gebali, S. et al. The Pfam protein families database in 2019. Nucleic Acids Res. 47, D427–D432 (2019).
Eddy, S. R. A probabilistic model of local sequence alignment that simplifies statistical significance estimation. PLoS Comput. Biol. 4, e1000069 (2008).
Bienert, S. et al. The SWISS-MODEL Repository-new features and functionality. Nucleic Acids Res. 45, D313–D319 (2017).
Studer, G. et al. QMEANDisCo-distance constraints applied on model quality estimation. Bioinformatics 36, 2647 (2020).
Kluyver, T. et al. Jupyter Notebooks: A publishing format for reproducible computational workflows. In Positioning and Power in Academic Publishing: Players, Agents and Agendas (ed. Loizides, F.) 87–90 (IOS Press, Amsterdam, 2016).
Pereira, F. et al. Pydna: a simulation and documentation tool for DNA assembly strategies using python. BMC Bioinform. 16, 142 (2015).
Gietz, R. D. & Woods, R. A. Transformation of yeast by lithium acetate/single-stranded carrier DNA/polyethylene glycol method. Methods Enzymol. 350, 87–96 (2002).
Kaiser, C., Michaelis, S. & Mitchell, A. Methods in Yeast Genetics: A Cold Spring Harbor Laboratory Course Manual (CSHL Press, New York, 1994).
Wieczorke, R. et al. Concurrent knock-out of at least 20 transporter genes is required to block uptake of hexoses in Saccharomyces cerevisiae. FEBS Lett. 464, 123–128 (1999).
Entian, K.-D. & Kötter, P. 25 Yeast genetic strain and plasmid collections. Methods Microbiol. 36, 629–666 (2007).
Schenk, M. & Bisswanger, H. A microplate assay for D-xylose/ D-glucose isomerase. Enzyme Microb. Technol. 22, 721–723 (1998).
Kulka, R. G. Colorimetric estimation of ketopentoses and ketohexoses. Biochem. J. 63, 542–548 (1956).
Madeira, F. et al. The EMBL-EBI search and sequence analysis tools APIs in 2019. Nucleic Acids Res. 47, W636–W641 (2019).
Bikandi, J., San Millán, R., Rementeria, A. & Garaizar, J. In silico analysis of complete bacterial genomes: PCR, AFLP-PCR and endonuclease restriction. Bioinformatics 20, 798–799 (2004).
Guindon, S. et al. New algorithms and methods to estimate maximum-likelihood phylogenies: Assessing the performance of PhyML 3.0. Syst. Biol. 59, 307–321 (2010).
Letunic, I. & Bork, P. Interactive Tree Of Life (iTOL) v4: recent updates and new developments. Nucleic Acids Res. 47, W256–W259 (2019).
Acknowledgements
This work was supported by the project FatVal PTDC/EAM-AMB/32506/2017 (POCI-01-0145-FEDER-032506), co-funded by the European Regional Development Fund (ERDF), through the Operational Programme for Competitiveness and Internationalization (COMPETE 2020), under Portugal 2020, and by the Fundação para a Ciência e a Tecnologia—FCT I.P through national funds. CBMA was supported by the “Contrato-Programa” UIDB/04050/2020 funded by national funds through the FCT I.P. PCS is recipient of a FCT PhD fellowship (SFRH/BD/140039/2018), and was supported by a Fulbright Scholarship Portugal grant from January to May 2020 at Lawrence Berkeley National Laboratory, Berkeley, CA, USA. BJ was awarded a Fulbright grant from The Swedish Fulbright Commission for Visiting Lecturers and Research Scholars between September 2014 and January 2015 visiting Lawrence Berkeley National Laboratory, Berkeley, CA, USA. This work was supported in part by the United States Department of Energy’s Genomic Science Program (grant SCW1039). Part of this work was performed at Lawrence Berkeley National Laboratory under US Department of Energy contract number DE-AC02-05CH11231. DNA sequencing was performed at the Vincent J. Coates Genomics Sequencing Laboratory at the University of California Berkeley, supported by NIH S10 Instrumentation grants S10RR029668 and S10RR027303.
Author information
Authors and Affiliations
Contributions
All authors contributed to the design of the project. P.C.S., J.C.N., F.A. and U.K. performed the experiments and analyzed the data. P.C.S., J.C.N., E.L.B. and B.J. drafted the manuscript. E.L.B. and B.J. revised and edited the final version. All authors read and approved the final manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher's note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Silva, P.C., Ceja-Navarro, J.A., Azevedo, F. et al. A novel d-xylose isomerase from the gut of the wood feeding beetle Odontotaenius disjunctus efficiently expressed in Saccharomyces cerevisiae. Sci Rep 11, 4766 (2021). https://doi.org/10.1038/s41598-021-83937-z
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-021-83937-z
This article is cited by
-
Impact of xylose epimerase on sugar assimilation and sensing in recombinant Saccharomyces cerevisiae carrying different xylose-utilization pathways
Biotechnology for Biofuels and Bioproducts (2023)
-
Low-abundance populations distinguish microbiome performance in plant cell wall deconstruction
Microbiome (2022)
-
Xylose Metabolism in Bioethanol Production: Saccharomyces cerevisiae vs Non-Saccharomyces Yeasts
BioEnergy Research (2022)
-
Xylitol production by Pseudomonas gessardii VXlt-16 from sugarcane bagasse hydrolysate and cost analysis
Bioprocess and Biosystems Engineering (2022)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.