Abstract
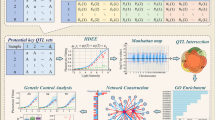
Light is the most important environmental cue signaling the transition from skotomorphogenesis to photomorphogenesis, thus affecting plant development and metabolic activity. How the light response mechanisms of maize seedlings respond to fluctuations in the light environment has not been well characterized to date. In this study, we built a gene coexpression network from a dynamic transcriptomic map of maize seedlings exposed to different light environments. Coexpression analysis identified ten modules and multiple genes that closely correlate with photosynthesis and characterized hub genes associated with regulatory networks, duplication events, domestication and improvement. In addition, we identified that 38% of hub genes underwent duplication events, 74% of which are related to photosynthesis. Moreover, we captured the dynamic expression atlas of gene sets involved in the chloroplast photosynthetic apparatus and photosynthetic carbon assimilation in different light environments, which should help to elucidate the key mechanisms and regulatory networks that underlie photosynthesis in maize. Insights from this study provide a valuable resource to better understand the genetic mechanisms of the response to fluctuations in the light environment in maize.






Similar content being viewed by others
Data availability
Sequence data from this article can be found in the National Center for Biotechnology Information Sequence Read Archive (http://www.ncbi.nlm.nih.gov/sra) under accession number SRP118761.
References
Achard P, Liao L, Jiang C, Desnos T, Bartlett J, Fu X, Harberd NP (2007) DELLAs contribute to plant photomorphogenesis. Plant Physiol 143:1163–1172
Allen JF (2002) Photosynthesis of ATP—electrons, proton pumps, rotors, and poise. Cell 110:273–276
Alter P, Dreissen A, Luo FL, Matsubara S (2012) Acclimatory responses of Arabidopsis to fluctuating light environment: comparison of different sunfleck regimes and accessions. Photosynth Res 113:221–237
Assenov Y, Ramirez F, Schelhorn SE, Lengauer T, Albrecht M (2008) Computing topological parameters of biological networks. Bioinformatics 24:282–284
Ballesteros ML, Bolle C, Lois LM, Moore JM, Vielle-Calzada JP, Grossniklaus U, Chua NH (2001) LAF1, a MYB transcription activator for phytochrome A signaling. Genes Dev 15:2613–2625
Baumgardt RL, Oliverio KA, Casal JJ, Hoecker U (2002) SPA1, a component of phytochrome A signal transduction, regulates the light signaling current. Planta 215:745–753
Benjamini Y, Hochberg Y (1995) Controlling the false discovery rate: a practical and powerful approach to multiple testing. J Roy Statist Soc Ser B 57:289–300
Boeckx T, Winters AL, Webb KJ, Kingston-Smith AH (2015) Polyphenol oxidase in leaves: is there any significance to the chloroplastic localization? J Exp Bot 66:3571–3579
Brohammer AB, Kono TJY, Springer NM, McGaugh SE, Hirsch CN (2018) The limited role of differential fractionation in genome content variation and function in maize (Zea mays L.) inbred lines. Plant J 93:131–141
Buchanan BB, Balmer Y (2005) Redox regulation: a broadening horizon. Annu Rev Plant Biol 56:187–220
Chattopadhyay S, Ang LH, Puente P, Deng XW, Wei N (1998) Arabidopsis bZIP protein HY5 directly interacts with light-responsive promoters in mediating light control of gene expression. Plant Cell 10:673–683
Cockburn W (1983) C3 C4: Mechanisms, and cellular and environmental regulation, of photosynthesis (Book). Plant Cell Environ 6:747–748
Darko E, Heydarizadeh P, Schoefs B, Sabzalian MR (2014) Photosynthesis under artificial light: the shift in primary and secondary metabolism. Philos Trans R Soc Lond B Biol Sci 369:20130243
de Wit M, Galvao VC, Fankhauser C (2016) Light-mediated hormonal regulation of plant growth and development. Annu Rev Plant Biol 67:513–537
Deng X-W, Matsui M, Wei N, Wagner D, Chu AM, Feldmann KA, Quail PH (1992) COP1, an Arabidopsis regulatory gene, encodes a protein with both a zinc-binding motif and a G beta homologous domain. Cell 71:791–801
Deng XW, Matsui M, Wei N, Wagner D, Chu AM, Feldmann KA, Quail PH (1994) Light inactivation of arabidopsis photomorphogenic repressor COP1 involves a cell-specific regulation of its ucleocytoplasmic partitioning. Cell 79:1035–1045
Du Z, Zhou X, Ling Y, Zhang Z, Su Z (2010) agriGO: a GO analysis toolkit for the agricultural community. Nucleic Acids Res 38:W64-70
Eberhard S, Finazzi G, Wollman FA (2008) The dynamics of photosynthesis. Annu Rev Genet 42:463–515
Fitter DW, Martin DJ, Copley MJ, Scotland RW, Langdale JA (2002) GLK gene pairs regulate chloroplast development in diverse plant species. Plant J 31:713–727
Fleet CM, Sun TP (2005) A DELLAcate balance: the role of gibberellin in plant morphogenesis. Curr Opin Plant Biol 8:77–85
Freeling M (2009) Bias in plant gene content following different sorts of duplication: tandem, whole-genome, segmental, or by transposition. Annu Rev Plant Biol 60:433–453
Garrone A, Archipowa N, Zipfel PF, Hermann G, Dietzek B (2015) Plant protochlorophyllide oxidoreductases A and B: catalytic efficiency and initial reaction steps. J Biol Chem 290:28530–28539
Giuliani R, Karki S, Covshoff S, Lin HC, Coe RA, Koteyeva NK, Evans MA, Quick WP, von Caemmerer S, Furbank RT, Hibberd JM, Edwards GE, Cousins AB (2019) Transgenic maize phosphoenolpyruvate carboxylase alters leaf-atmosphere CO2 and (13)CO2 exchanges in Oryza sativa. Photosynth Res. https://doi.org/10.1007/s11120-019-00655-4
Grossman AR, Bhaya D, Apt KE, Kehoe DM (1995) LIGHT-harvesting complexes in oxygenic photosynthesis: diversity, control, and evolution. Ann Rev Genet 29:231–288
Hanada K, Kuromori T, Myouga F, Toyoda T, Li WH, Shinozaki K (2009) Evolutionary persistence of functional compensation by duplicate genes in Arabidopsis. Genome Biol Evol 1:409–414
He JX, Gendron JM, Sun Y, Gampala SS, Gendron N, Sun CQ, Wang ZY (2005) BZR1 is a transcriptional repressor with dual roles in brassinosteroid homeostasis and growth responses. Science 307:1634–1638
Hornitschek P, Kohnen MV, Lorrain S, Rougemont J, Ljung K, Lopez-Vidriero I, Franco-Zorrilla JM, Solano R, Trevisan M, Pradervand S, Xenarios I, Fankhauser C (2012) Phytochrome interacting factors 4 and 5 control seedling growth in changing light conditions by directly controlling auxin signaling. Plant J 71:699–711
Hufford MB, Xu X, van Heerwaarden J, Pyhajarvi T, Chia JM, Cartwright RA, Elshire RJ, Glaubitz JC, Guill KE, Kaeppler SM, Lai J, Morrell PL, Shannon LM, Song C, Springer NM, Swanson-Wagner RA, Tiffin P, Wang J, Zhang G, Doebley J, McMullen MD, Ware D, Buckler ES, Yang S, Ross-Ibarra J (2012) Comparative population genomics of maize domestication and improvement. Nat Genet 44:808–811
Jensen RG (1983) Photosynthesis: C3, C4. mechanisms, and cellular and environmental regulation, of photosynthesis. Science 222:1009–1009
Jin J, Tian F, Yang DC, Meng YQ, Kong L, Luo J, Gao G (2017) PlantTFDB 4.0: toward a central hub for transcription factors and regulatory interactions in plants. Nucleic Acids Res 45:D1040–D1045
Kanehisa M, Furumichi M, Tanabe M, Sato Y, Morishima K (2017) KEGG: new perspectives on genomes, pathways, diseases and drugs. Nucleic Acids Res 45:D353–D361
Kaur N, Li J, Hu J (2013) Peroxisomes and Photomorphogenesis. In: del Río LA (ed) Peroxisomes and their Key Role in Cellular Signaling and Metabolism. Springer, Netherlands, Dordrecht, pp 195–211
Kim B, Jeong YJ, Corvalan C, Fujioka S, Cho S, Park T, Choe S (2014) Darkness and gulliver2/phyB mutation decrease the abundance of phosphorylated BZR1 to activate brassinosteroid signaling in Arabidopsis. Plant J 77:737–747
Kramer DM, Avenson TJ, Edwards GE (2004) Dynamic flexibility in the light reactions of photosynthesis governed by both electron and proton transfer reactions. Trends Plant Sci 9:349–357
Kuhlgert S, Austic G, Zegarac R, Osei-Bonsu I, Hoh D, Chilvers MI, Roth MG, Bi K, TerAvest D, Weebadde P, Kramer DM (2016) MultispeQ Beta: a tool for large-scale plant phenotyping connected to the open PhotosynQ network. R Soc Open Sci 3:160592
Langfelder P, Horvath S (2008) WGCNA: an R package for weighted correlation network analysis. BMC Bioinformatics 9:559
Langmead B, Trapnell C, Pop M, Salzberg SL (2009) Ultrafast and memory-efficient alignment of short DNA sequences to the human genome. Genome Biol 10:R25
Li T, Jia KP, Lian HL, Yang X, Li L, Yang HQ (2014) Jasmonic acid enhancement of anthocyanin accumulation is dependent on phytochrome A signaling pathway under far-red light in Arabidopsis. Biochem Biophys Res Commun 454:78–83
Li QF, Huang LC, Wei K, Yu JW, Zhang CQ, Liu QQ (2017) Light involved regulation of BZR1 stability and phosphorylation status to coordinate plant growth in Arabidopsis. Biosci Rep. https://doi.org/10.1042/BSR20170069
Lu M, Yang G, Li P, Wang Z, Fu S, Zhang X, Chen X, Shi M, Ming Z, Xia J (2018) Bioinformatic and functional analysis of a key determinant underlying the substrate selectivity of the Al transporter, Nrat1. Front Plant Sci 9:606
Luo XM, Lin WH, Zhu S, Zhu JY, Sun Y, Fan XY, Cheng M, Hao Y, Oh E, Tian M, Liu L, Zhang M, Xie Q, Chong K, Wang ZY (2010) Integration of light- and brassinosteroid-signaling pathways by a GATA transcription factor in Arabidopsis. Dev Cell 19:872–883
Menon C, Sheerin DJ, Hiltbrunner A (2016) SPA proteins: SPAnning the gap between visible light and gene expression. Planta 244:297–312
Miao Z, Han Z, Zhang T, Chen S, Ma C (2017) A systems approach to a spatio-temporal understanding of the drought stress response in maize. Sci Rep 7:6590
Murchie EH, Niyogi KK (2011) Manipulation of photoprotection to improve plant photosynthesis. Plant Physiol 155:86–92
Okada K (2009) PetH is rate-controlling in the interaction between PetH, a component of the supramolecular complex with photosystem II, and PetF, a light-dependent electron transfer protein. Biochem Biophys Res Commun 389:394–398
Oyama T, Shimura Y, Okada K (1997) The Arabidopsis HY5 gene encodes a bZIP protein that regulates stimulus-induced development of root and hypocotyl. Genes Dev 11:2983–2995
Pahari S, Cormark RD, Blackshaw MT, Liu C, Erickson JL, Schultz EA (2014) Arabidopsis UNHINGED encodes a VPS51 homolog and reveals a role for the GARP complex in leaf shape and vein patterning. Development 141:1894–1905
Panchy N, Lehti-Shiu M, Shiu SH (2016) Evolution of gene duplication in plants. Plant Physiol 171:2294–2316
Parks BM (2003) The red side of photomorphogenesis. Plant Physiol 133:1437–1444
Ponnu J, Wahl V, Schmid M (2011) Trehalose-6-phosphate: connecting plant metabolism and development. Front Plant Sci 2:70
Robinson MD, McCarthy DJ, Smyth GK (2010) edgeR: a Bioconductor package for differential expression analysis of digital gene expression data. Bioinformatics 26:139–140
Sakata S, Mizusawa N, Kubota-Kawai H, Sakurai I, Wada H (2013) Psb28 is involved in recovery of photosystem II at high temperature in Synechocystis sp. PCC 6803. Biochim Biophys Acta 1827:50–59
Scharf KD, Berberich T, Ebersberger I, Nover L (2012) The plant heat stress transcription factor (Hsf) family: structure, function and evolution. Biochim Biophys Acta 1819:104–119
Schnable JC, Springer NM, Freeling M (2011) Differentiation of the maize subgenomes by genome dominance and both ancient and ongoing gene loss. Proc Natl Acad Sci USA 108:4069–4074
Schottler MA, Toth SZ (2014) Photosynthetic complex stoichiometry dynamics in higher plants: environmental acclimation and photosynthetic flux control. Front Plant Sci 5:188
Schottler MA, Toth SZ, Boulouis A, Kahlau S (2015) Photosynthetic complex stoichiometry dynamics in higher plants: biogenesis, function, and turnover of ATP synthase and the cytochrome b6f complex. J Exp Bot 66:2373–2400
Schumann T, Paul S, Melzer M, Dormann P, Jahns P (2017) Plant growth under natural light conditions provides highly flexible short-term acclimation properties toward high light stress. Front Plant Sci 8:681
Shannon P, Markiel A, Ozier O, Baliga NS, Wang JT, Ramage D, Amin N, Schwikowski B, Ideker T (2003) Cytoscape: a software environment for integrated models of biomolecular interaction networks. Genome Res 13:2498–2504
Shi Q, Kong F, Zhang H, Ye J, Heng S, Liang R, Ma L, Liu J, Lu X, Li P, Li G (2019) Molecular mechanisms governing shade responses in maize. Biochem Biophys Res Commun 516:112–119
Shikanai T (2016) Chloroplast NDH: A different enzyme with a structure similar to that of respiratory NADH dehydrogenase. Biochim Biophys Acta 1857:1015–1022
Smith AM, Zeeman SC, Smith SM (2005) Starch degradation. Annu Rev Plant Biol 56:73–98
Trapnell C, Roberts A, Goff L, Pertea G, Kim D, Kelley DR, Pimentel H, Salzberg SL, Rinn JL, Pachter L (2012) Differential gene and transcript expression analysis of RNA-seq experiments with TopHat and Cufflinks. Nat Protoc 7:562–578
Wang H, Wu G, Zhao B, Wang B, Lang Z, Zhang C, Wang H (2016) Regulatory modules controlling early shade avoidance response in maize seedlings. BMC Genomics 17:269
Wang B, Lin Z, Li X, Zhao Y, Zhao B, Wu G, Ma X, Wang H, Xie Y, Li Q, Song G, Kong D, Zheng Z, Wei H, Shen R, Wu H, Chen C, Meng Z, Wang T, Li Y, Li X, Chen Y, Lai J, Hufford MB, Ross-Ibarra J, He H, Wang H (2020) Genome-wide selection and genetic improvement during modern maize breeding. Nat Genet. https://doi.org/10.1038/s41588-020-0616-3
Weaver LM, Amasino RM (2001) Senescence is induced in individually darkened arabidopsis leaves, but inhibited in whole darkened plants. Plant Physiol 127:876–886
Weise SE, Weber AP, Sharkey TD (2004) Maltose is the major form of carbon exported from the chloroplast at night. Planta 218:474–482
Wise RR, Hoober JK (2006) The Diversity of Plastid Form and Function. Springer, The Netherlands, pp 3–26
Wu SH (2014) Gene expression regulation in photomorphogenesis from the perspective of the central dogma. Annu Rev Plant Biol 65:311–333
Yilmaz A, Nishiyama MY Jr, Fuentes BG, Souza GM, Janies D, Gray J, Grotewold E (2009) GRASSIUS: a platform for comparative regulatory genomics across the grasses. Plant Physiol 149:171–180
Zhang J (2012) Genetic Redundancies and Their Evolutionary Maintenance. In: Soyer OS (ed) Evolutionary Systems Biology. Springer, New York , pp 279–300
Acknowledgements
This study was funded by the Young Scientists Fund of the National Natural Science Foundation of China (Grant No. 31701438) and the Science Foundation for Post Doctorate Research of the Ministry of Science and Technology of China (2018M633588).
Author information
Authors and Affiliations
Contributions
Shutu Xu, Dongwei Guo and Jiquan Xue conceived and designed the experiments. Shutu Xu, Xiaonan Gou, Wenxin Zhang, Ting Li and Jianzhou Qu performed experiments. Jianzhou Qu analyzed and interpreted the data. Jianzhou Qu and Shutu Xu wrote the paper and prepared figures and/or tables. All authors read and approved the final manuscript.
Corresponding author
Ethics declarations
Conflicts of interest
All the authors declare that they have no conflicts of interest.
Ethical approval
This article does not contain any studies with human participants or animals performed by any of the authors.
Additional information
Communicated by Stefan Hohmann.
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
438_2021_1761_MOESM1_ESM.tif
Supplementary Correlation of gene expression level across replicates. For each gene, the expression values were normalized by the log2(FPKM+1). The scale bar shows the degree of correlation between samples. The samples from seedlings under normal light condition (SNL), seedlings after four days of dark treatment (SDT) and four days after seedlings return to normal light (SRNL) (TIF 2312 KB)
438_2021_1761_MOESM2_ESM.tif
Supplementary Comparison of expression patterns of candidate genes by qRT-PCR and RNA-Seq. Histogram showing expression levels of candidate genes based on qRT-PCR (left). Boxplot showing expression levels of candidate genes based on RNA-Seq (right). The samples from seedlings under normal light condition (SNL), seedlings after four days of dark treatment (SDT) and four days after seedlings return to normal light (SRNL) (TIF 3827 KB)
Rights and permissions
About this article
Cite this article
Qu, J., Gou, X., Zhang, W. et al. New insights into the response of maize to fluctuations in the light environment. Mol Genet Genomics 296, 615–629 (2021). https://doi.org/10.1007/s00438-021-01761-6
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00438-021-01761-6