Abstract
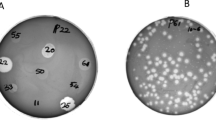
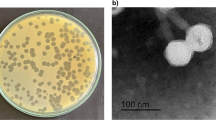
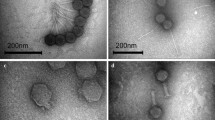
Klebsiella pneumoniae causes a variety of human infections including pneumonia. Herein, 3 lytic bacteriophages specific for K. pneumoniae designated ΦKpnM-vB1, ΦKpnP-vB2 and ΦKpnM-vB3 were isolated and characterized. Transmission electron microscopy (TEM) analysis revealed that both ΦKpnM-vB1 and ΦKpnM-vB3 belong to family Myoviridae, while ΦKpnP-vB2 is a member of family Podoviridae. The one-step growth curve showed that ΦKpnM-vB1, ΦKpnP-vB2 and ΦKpnM-vB3 exhibited latent period of 10 min. The average burst sizes were 100, 150 and 120 PFU/cell, respectively. Isolated phages showed high thermal and pH stability. The genomic analysis indicated that ΦKpnM-vB1, ΦKpnP-vB2 and ΦKpnM-vB3 contain dsDNA genome with estimated sizes of 55, 40 and 50 Kbp, respectively. Isolated phages had lytic activity on K. pneumoniae and Escherichia coli strains. Isolated phages were highly efficient in reduction of Klebsiella biofilm suggesting their use to control biofilm formation caused by this pathogen. Isolated ΦKpnM-vB1, ΦKpnP-vB2 and ΦKpnM-vB3 are proposed to be suitable candidates for phage therapy applications. These phages offer an effective solution for treatment of infections caused by these drug-resistant bacteria.







Similar content being viewed by others
REFERENCES
Manohar, P., Tamhankar, A.J., Lundborg, C.S., and Nachimuthu, R., Front. Microbiol., 2019, vol. 10, p. 574.
Podschun, R. and Ullmann, U., Clin. Microbiol. Rev., 1998, vol. 11, no. 4. pp. 589–603.
Akers, K.S., Mende, K., Cheatle, K.A., Zera, W.C., Yu, X., Beckius, M.L., et al., BMC Infect. Dis., 2014, vol. 14, p. 190.
Paczosa, M.K. and Mecsas, J., Microbiol. Mol. Biol. Rev., 2016, vol. 80, no. 3. pp. 629–661.
Nagel, T.E., Chan, B.K., De Vos, D., El-Shibiny, A., Kang’ethe, E.K., Makumi, A., and Pirnay, J.P., Front. Microbiol., 2016, vol. 7, p. 882.
Hung, C.H., Kuo, C.F., Wang, C.H., Wu, C.M., and Tsao, N., Antimicrob. Agents Chemother., 2011, vol. 55, no. 4. pp. 1358–1365.
Chibani-Chennoufi, S., Bruttin, A., Dillmann, M.L., and Brussow, H., J. Bacteriol., 2004, vol. 186, no. 12. pp. 3677–3686.
El-Shibiny, A., El-Sahhar, S., and Adel, M., J. Appl. Microbiol., 2017, vol. 123, no. 2. pp. 556–567.
Vandenheuvel, D., Lavigne, R., and Brussow, H., Annu. Rev. Virol., 2015, vol. 2, no. 1. pp. 599–618.
Principi, N., Silvestri, E., and Esposito, S., Front. Pharmacol., 2019, vol. 10, p. 513.
Cao, F., Wang, X., Wang, L., Li, Z., Che, J., Wang, L., et al., Biomed. Res. Int., 2015, vol. 2015, p. 752930.
Tan, D., Zhang, Y., Cheng, M., Le, S., Gu, J., Bao, J., et al., Viruses, 2019, vol. 11, no. 11.
Forbes, B.A., Sahm, D.F., Weissfeld, A.S., and Bailey, W.R. Bailey and Scott’s Diagnostic Microbiology, 12th ed., St. Louis: Elsevier Mosby, 2007.
Clifford, R.J., Milillo, M., Prestwood, J., Quintero, R., Zurawski, D.V., Kwak, Y.I., et al., PLoS One, 2012, vol. 7, no. 11. p. e48558.
Bauer, A.W., Kirby, W.M., Sherris, J.C., and Turck, M., Am. J. Clin. Pathol., 1966, vol. 45, no. 4. pp. 493–496.
Clinical and Laboratory Standards Institute (CLSI), 26 ed., Pennyslvania, USA: Wayne, 2016.
Cabral, A.B., Melo Rde, C., Maciel, M.A., and Lopes, A.C., Rev. Soc. Bras. Med. Trop., 2012, vol. 45, no. 5. pp. 572–578.
Jamalludeen, N., Johnson, R.P., Friendship, R., Kropinski, A.M., Lingohr, E.J., and Gyles, C.L., Vet. Microbiol., 2007, vol. 124, nos. 1–2. pp. 47–57.
Goodridge, L., Gallaccio, A., and Griffiths, M.W., Appl. Environ. Microbiol., 2003, vol. 69, no. 9. pp. 5364–5371.
Jamal, M., Hussain, T., Das, C.R., and Andleeb, S., J. Med. Microbiol., 2015, vol. 64, Pt. 4. pp. 454–462.
Kwiatek, M., Mizak, L., Parasion, S., Gryko, R., Olender, A., and Niemcewicz, M., Folia Microbiol. (Praha), 2015, vol. 60, no. 1. pp. 7–14.
Gill, J.J. and Hyman, P., Curr. Pharm. Biotechnol., 2010, vol. 11, no. 1. pp. 2–14.
Chhibber, S., Nag, D., and Bansal, S., BMC Microbiol., 2013, vol. 13, no. pp. 174.
Kesik-Szeloch, A., Drulis-Kawa, Z., Weber-Dabrowska, B., Kassner, J., Majkowska-Skrobek, G., Augustyniak, D., et al., Virol. J., 2013, vol. 10, pp. 100.
Taha, O.A., Connerton, P.L., Connerton, I.F., and El-Shibiny, A., Front. Microbiol., 2018, vol. 9, p. 2127.
Fernandez, L., Gutierrez, D., Garcia, P., and Rodriguez, A., Antibiotics (Basel), 2019, vol. 8, no. 3.
Chaturongakul, S. and Ounjai, P., Front. Microbiol., 2014, vol. 5, p. 442.
Anand, T., Virmani, N., Kumar, S., Mohanty, A.K., Pavulraj, S., Bera, B.C., et al., J. Glob. Antimicrob. Resist., 2019, vol. 21, pp. 34–41.
Mah, T.F., Pitts, B., Pellock, B., Walker, G.C., Stewart, P.S., and O’Toole, G.A., Nature, 2003, vol. 426, no. 6964. pp. 306–310.
Verma, V., Harjai, K., and Chhibber, S., Biofouling, 2010, vol. 26, no. 6. pp. 729–737.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Ethical approval: This study was approved by Ethics Committee, Zagazig University. All participants provided written informed consent prior to enrolment in the study.
Competing interests: The authors declare that they have no competing interests.
Rights and permissions
About this article
Cite this article
Askoura, M., Saed, N., Enan, G. et al. Characterization of Polyvalent Bacteriophages Targeting Multidrug-Resistant Klebsiella pneumonia with Enhanced Anti-Biofilm Activity. Appl Biochem Microbiol 57, 117–126 (2021). https://doi.org/10.1134/S000368382101004X
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1134/S000368382101004X