Abstract
During the decomposition process of soil organic carbon (SOC), microbial products such as microbial necromass and microbial metabolites may form an important stable carbon (C) pool, called microbially derived C, which has different decomposition patterns from plant-derived C. However, current Earth System Models do not simulate this microbially derived C pool separately. Here, we incorporated the microbial necromass pool to the first-order kinetic model and the Michaelis–Menten model, respectively, and validated model behaviors against previous observation data from the decomposition experiments of 13C-labeled necromass. Our models showed better performance than existing models and the Michaelis–Menten model was better than the first-order kinetic model. Microbial necromass C was estimated to be 10–27% of total SOC in the study soils by our models and therefore should not be ignored. This study provides a novel modification to process-based models for better simulation of soil organic C under the context of global changes.
Similar content being viewed by others
Introduction
The pool size of global soil organic carbon (SOC) to 1 m depth is estimated to be 1417–1469.5 Pg C [1, 2], which is much larger than the amounts of carbon (C) in plants and atmosphere. The uncertainty of current process-based models in estimating global SOC size is 50% [3], meaning that about 630–735 Pg C still cannot be accurately simulated. Measures have been proposed to improve Earth System Models, such as consideration of nitrogen constraints [4, 5], or incorporation of density-dependent formulation of microbial turnover [6]. Although other sources besides our understanding of the potential processes may also contribute to the model uncertainty such as error in observation data given the low precision of soil carbon measurement, uncertainty in initial conditions, and uncertainty in parameters due to the limited available data for constraining model parameters, it is still important to incorporate the mechanisms already confirmed in observation experiments into process-based models for better simulation and prediction of C cycling and climate changes.
SOC pools in process-based models are usually divided into different parts based on their decomposition rates, such as the slow pool and the fast pool. However, SOC is composed of a very complex mixture of polymers of microbial and plant residues and degradation products [7], and different compounds not only differ in decomposition rate, but also differ in decomposition pattern. For example, plant-derived organic N and microbially derived organic N have been found to have different decomposition curves [8]. In addition, the microbial necromass pool (i.e., mass from the cell death and subsequent lysis and fragmentation of soil microbes) may interact with soil minerals more easily than the plant residues [9], making it necessary to simulate the microbial necromass pool separately from the plant residue pool. Although the living microbial biomass in the soil is a relatively small C pool, microbially derived SOC may accumulate to form a large proportion pool if microbial biomass turnover rate is higher than plant litter input rate [10], or the chemical nature of microbial necromass is not labile or microbially derived SOC is protected by the soil mineral matrix [11,12,13,14]. Recently, a variety of methods have been used to estimate the contribution of microbial necromass C to SOC in different soils, and a range of 24–80% was determined [11, 15,16,17]. Therefore, the role of microbial necromass in the formation of organic matter may have been seriously underestimated, and should be considered in models [18].
In most process-based models, the decomposition of SOC is expressed by the first-order kinetic equation [19]. The decomposition coefficient is generally determined by the environmental factors [20], and the roles of the microbial community play in the decomposition of organic matter have not been well represented in these models [5]. Recently, the Michaelis–Menten equation was used to describe the decomposition rate of organic matter in many models, such as the MEND [21,22,23], GER [24], AWB [25, 26], MIMICS [27], RESOM [28, 29], CORPSE [30], ORCHIMIC [5] models. These models consider the microbial biomass carbon (MBC) pool separately and have been found to provide better simulation results than the first-order kinetic models [3, 31]. However, the microbial necromass pool has not been considered as a separate pool with a different decomposition pattern from plant residue in these models. We propose a conceptual four-pool model including MBC and microbial necromass pools (Supplementary Fig. 1) because once MBC pool is separated, it is undoubtedly logical to separate the necromass pool to better represent the formation process of microbially derived SOC.
Recent findings have made it possible to realize the conceptual model because the decomposition pattern of microbial necromass has been observed [8] and the responses of the microbial necromass pool to environmental factors have been studied [32]. In addition, although fungi, gram-positive bacteria (including Actinomycetes), and gram-negative bacteria have different molecular structures, their turnover rates were found to be similar, and the quality of microbial necromass was found to have no significant effect on their decomposition rates [33], suggesting that the same decomposition rate can be used for different groups of microbial necromass in the model.
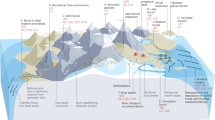
In this study, we established a new model accounting for the microbial necromass pools, and demonstrated its validity and feasibility using observation data from previous experiments of the decomposition of 13C-labeled microbial necromass [33,34,35]. Two archetypal models were selected [24,25,26] and the microbial necromass pools were added into these two models and we named these two models as the Michaelis–Menten necromass decomposition (MIND) model and the first-order necromass decomposition (FOND) model (Fig. 1). The aims of this study were: (1) to evaluate the necessity of incorporating the microbial necromass pool into ESMs and to compare the performance of MIND and FOND models; (2) to investigate the decomposition rate of microbial necromass C in different soils; and (3) to estimate the contribution of microbial necromass C to soil organic C under steady state. Our results suggest that microbial necromass C is an essential, but overlooked part of stable SOC, and model performance is improved by representing the microbially derived organic carbon pool in process-based models.
A The Michaelis–Menten necromass decomposition model (MIND); B the first-order necromass decomposition model (FOND). The red dotted line represents the paths involved when 13C-labeled microbial necromass is added to the system. The valve symbol and black dashed line represent the regulation of the size of MBC pool on the processes according to Michaelis–Menten kinetics. CP plant-derived carbon, CB microbial biomass carbon, CNF fast pool of microbial necromass carbon, CN-MAOM mineral-associated pool of microbial necromass carbon, CDP plant-derived dissolved organic carbon, CDN microbially derived dissolved organic carbon (color figure online).
Method
Model structure
The model structures are shown in Fig. 1. In MIND model, the Michaelis–Menten equation was used to describe the nonlinear variation of substrate decomposition rate with microbial biomass. The FOND model assumed that the variation of the decomposition rate was controlled by environmental factors and in our simulations the decomposition rate did not change because we used incubation experiment data under controlled environment. In both models, after the death of soil microorganisms they are transferred to the fast pool and the mineral-associated pool of microbial necromass proportionally. We minimized the number of additional C pools to avoid introducing more parameters and overfitting.
MIND was expressed by the following equations:
where CP, CB, CNF, CN-MAOM, and CN are pool sizes (mg C g−1 soil) of plant-derived organic carbon, microbial biomass carbon (MBC), microbial fast necromass carbon, microbial mineral-associated necromass carbon, total microbial necromass carbon (MNC), respectively; I is the annual average carbon input rate (mg C g−1 soil h−1) to soil from plant-derived litter, which was calculated based on litterfall minus litter heterotrophic respiration (g C m−2 s−1) and then converted to mg C g−1 soil h−1 using bulk density (0–15 cm) data; Vmax,P, Vmax,N are maximum assimilation rate (mg C mg−1 MBC h−1) of CP and CN, respectively; KM,P, KM,N are half-saturation for assimilation (mg C g−1 soil) of CP and CN, respectively; CUEP, CUEN are carbon use efficiency (unitless) of CP and CN, respectively; kB is average mortality rate of the microbial community (h−1); fBNF is proportion of fast pool in MBC (unitless); RMAOM-F is ratio of the decomposition rate of the mineral-associated pool to that of the fast pool of microbial necromass (unitless).
When the 13C-labeled microbial necromass C was added to the system, we can get the following equations:
where superscript u stands for “unlabeled” and l stands for “labeled” pool; the size of a pool was the sum of labeled proportion and unlabeled proportion. For example, total MBC was the sum of labeled MBC and unlabeled MBC.
The analytical steady-state solutions of MIND were as follows:
FOND was expressed by the following equations:
where CD is dissolved organic carbon pool (DOC, mg C g−1 soil), which is the sum of plant-derived DOC (CDP) and microorganism-derived DOC (CDN); f is the fraction of inputs into CP (unitless); kP, kDP, kDN, and kNF are the decomposition rate (mg C mg−1 C h−1) of CP entering CDP, CDP entering CP, CDN entering CN-MAOM, CNF entering CDN, respectively; fP, fDP, and fN are fraction (unitless) of CP entering CDP, CDP entering CP, CN entering CDN, respectively; kB is turnover rate constant of CB (h−1); kuptake is uptake rate constant of CD (mg C g−1 DOC h−1); CUE is carbon use efficiency (unitless); fBNF is proportion of fast pool in MBC (unitless); RMAOM-F is the decomposition rate ratio of the mineral-associated pool to that of the fast pool (unitless).
When 13C-labeled microbial necromass C was added to the system, we can get the following equations:
The analytical steady-state solutions of FOND were as follows:
The mean residence time (MRT) for the necromass 13C in FOND model was calculated according to the following equation according to literatures [32, 36]:
Models MIND-O and FOND-O had the same structure as MIND and FOND except that the MNC pool was not considered (Supplementary Fig. 2). The equations for MIND-O and FOND-O models are listed in the figure legend of Supplementary Fig. 2.
Observation data
We searched the Web of Science for observation data on the decomposition of 13C-labeled microbial necromass and found three articles [33,34,35] and four data sets (named studies 1–4; Supplementary Tables 1 and 2 and Supplementary Fig. 3). In short, soil microorganisms were cultured in a matrix with resources enriched in 13C. The microorganisms used 13C as their carbon sources and were then killed to form 13C-enriched microbial necromass. The 13C-enriched microbial necromass was then put into soil for lab incubation in studies 1 and 2 and for in situ decomposition experiments in studies 3 and 4. Studies 1 and 2 used 13C-labeled E. coli necromass, and studies 3 and 4 used 13C-labeled microbial groups including fungi, gram-positive bacteria (including Actinomycetes), and gram-negative bacteria. The soils in studies 1 and 2 were from temperate farmland and forests ecosystems, respectively, and the soils in studies 3 and 4 were incubated in temperate and tropical forest ecosystems, respectively.
Parameter estimation
The model parameterization was based on the 13C recovery data from each observation study. Some parameters had fixed values from previous findings, while most parameters were given a range first, within which the parameter had uniform distribution (Supplementary Table 3). Then these parameters without reference values were estimated using the nonlinear least squares method. We first used the ODE45 solver to calculate the integration of the differential equations y = f(t,y) from t0 to tn, with an initial condition of y0. The ODE45 solver was based on an explicit Runge–Kutta (4,5) equation (the Dormand–Prince pair). It is a single-step solver, which needs only the solution at the immediate preceding time, y(tn–1) [37, 38]. Then, the lsqnonlin function was used to fit estimated 13C recovery to measured data to get the best combination of parameters.
The lsqnonlin function uses nonlinear least squares to fit the data and solve the problems with constraints of parameters within the given ranges. The lsqnonlin function uses the trust-region-reflective algorithm [39, 40] by default to adapt to the problem that Levenberg–Marquardt algorithm [41] cannot constrain the ranges of parameters.
Model validation
We used the leave-one-out cross-validation (LOOCV) [42], which is for model validation of small sample size like our data. The learning algorithm was applied once for each instance, using all other instances as a training set and then the selected instance was used as a single-item test set. This step continued until each sample was treated as a validation.
We also tested the model with the recovery of 13C in respired CO2 values in each observation study, as well as SOC and MBC values under steady-state conditions from each observation studies. Linear regression was performed between estimated data and measured data.
In the four studies on the decomposition of 13C-labeled necromass, the units of the decomposition rates of microbial necromass in their calculations were different from those in our study and therefore cannot be compared to our values. Therefore, we compared the ratio of the decomposition rate of the mineral-associated pool to that of the fast pool (RMAOM-F) of microbial necromass and the proportion of fast pool in MBC (fBNF) of modeled results to those originally estimated in their studies because these are unitless values. The exponential model was used to calculate the decomposition rates of the fast and mineral-associated pools of necromass in studies 1 and 2 [34, 35]. In studies 3 and 4, the initial decomposition rate and the final decomposition rate were given, which approximately corresponded to the decomposition rate of fast and mineral-associated pools in our study, respectively [33]. Since fBNF was not given in studies 3 and 4, we only compared it with the values in studies 1 and 2.
Slope, R2, root mean squared error (RMSE), and mean absolute error (MAE) were calculated to quantify the accuracy of the model estimates, following equations:
where \(\hat y\) is simulation value, y is observations value, \(\bar y\) is average value, n is number of observations.
Uncertainty analysis
The Lsqnonlin function starts from the initial value y0, and iteratively finds the minimum value of the sum of squares of the function between the measured value and the modeled value. Because the Lsqnonlin function depends on the settings of initial values and iteration intervals, it was a local optimal solution instead of global optimal solution. Thus, it is possible that the best solution was missed. We used the Monte Carlo random sampling method to extract subintervals and initial values randomly within the parameter range for 1000 times. The distribution of the parameter was selected when R2 between modeled results and observation data was >0.9.
The Box–Cox transformation was used to normalize the data when mean of the parameter distribution was not in the center of the interval, and the data did not obey the normal distribution:
where Yi is the initial value, λ is the transformation parameter.
The uncertainty of the parameter was presented as the 95% confidence interval of the normal distribution.
Sensitivity analysis
Sensitivity analysis was performed by changing from the lower end to the higher end of 95% confidence interval of the parameter distribution and analyzing the changes in the sizes of different carbon pools in the model. The sensitivity was expressed as follows Allison et al. [25]:
Global scale estimation
Global microbial necromass C under steady state was estimated using Eqs. (18)–(20). We used our data of the linear relationships between microbial carbon use efficiency (CUE) and mean annual temperature (MAT) and mean annual precipitation (MAP) to estimate microbial CUE of plant residue (CUEp) in different areas (Eq. (51)): [43]
Microbial CUE of microbial necromass (CUEN) was assumed to be 0.1 higher than CUEp because the C:N ratio of microbial necromass is lower than that of plant residue [44,45,46]. We established linear relationships between fBNF, RNSF, kB and MAT or MAP based on the four observation studies (Eqs. (52)–(54)) and extrapolated these parameters to global estimates:
Vmax,N, KM,N were estimated according to the Arrhenius equations (Eqs. (55)–(57)) using soil temperature (T) according to literatures [21, 25]:
The decomposition rate of microbial residues was estimated by soil moisture, soil pH, and soil clay content according to literatures [5, 20, 47, 48]:
where T is the soil temperature (K); Tref is the reference temperature, set as 285.15 K; Ea is the activation energy of substrate decomposition (kJ mol−1) and was 50 and 30 kJ mol−1 for Vmax,N and KM,N, respectively [30, 49]. R is the ideal gas constant (0.008314 kJ mol−1 K−1); Vmax,0 and KM,0 are the pre-exponential coefficient, respectively, and were estimated to be 240 (mg C mg−1 MBC day−1) and 300 (mg C g−1 soil), respectively, based on the four observation studies. θ is soil moisture (%); pHopt is the optimal pH for substrate decomposition and was set as 6 [21]. pHsen is the sensitivity parameter of substrate decomposition and was set as 1.66 according to the average value in a previous study [48]. Clay is the soil clay content (%). MAT is the mean annual temperature (°C). MAP is the mean annual precipitation (mm).
Data sources
Input rate I was from the community land model (CLM) spatial output data, and CLM5.0 data, which is from the National Center for Atmospheric Research (NCAR) (https://www.earthsystemgrid.org). The file of monthly litterfall rate was clm50_r270_1deg_GSWP3V1_iso_newpopd_hist.clm2.h0.LITFALL.185001-201412.nc and the file of monthly litter heterotrophic respiration was clm50_r270_1deg_GSWP3V1_iso_newpopd_hist.clm2.h0.LITTERC_HR.185001-201412.nc. We chose the subset between 2000 and 2014 to obtain the mean annual data.
Climate data MAT and MAP were from the worldclim database (http://www.worldclim.org) [50]. Soil moisture data were from the gap-free global annual soil moisture for 1991–2018 [51]. Soil temperature was derived from NCEP/NCAR 40 years reanalysis data [52]. Soil pH, soil clay content, and soil bulk density were obtained from harmonized data set of derived soil properties for the world (WISE30sec) [53]. SOC content was from a global 3D soil information system [54], and the average value of the top 0–15 cm was used. Global biomes were based on the classification by Olson and Dinerstein [55], with the associated spatial data available at http://maps.tnc.org/files/shp/terr-ecoregions-TNC.zip. All data were resampled to a spatial resolution of 0.125° × 0.125° for calculation.
Results
Model simulation
Generally, the decomposition curve of 13C-labeled microbial necromass C can be well simulated by both MIND and FOND models (Fig. 2 and Supplementary Fig. 4). The R2 of the regression of the recovery in soil 13C of the MIND simulation results with observations was >0.9, the RMSEs were between 0.88 and 2.66, while the R2 of the regression of FOND simulation results with measured data was >0.68, and the RMSEs were between 1.90 and 4.93 (Table 1).
MIND model showed that the recovery of 13C in MBC experienced a rapid growth stage at the beginning, and then gradually declined for the four data sets (Fig. 3 and Supplementary Fig. 5). Additionally, 10–25% of the microbial necromass C was quickly transferred to MBC (Fig. 3), and then was transferred to the mineral-associated pool of microbial necromass gradually as microbes died. In contrast, the FOND model showed that the recovery of 13C in MBC of four data sets increased only slightly and <5% of the MNC was transferred to MBC (Fig. 3).
In all the four studies, the fast pool of MNC (CNF) was simulated to be quickly decomposed within 100 days (Fig. 3). The difference among different studies was that the proportion of MBC transferred to the fast and mineral-associated pools of necromass was different (Table 2). The MIND model simulated that the recovery of 13C in CNF still had a little at the later stage, while the FOND model simulated that the recovery of 13C in CNF was almost 0 at 50–100 days of decomposition (Fig. 3).
The temporal variation of simulated mineral-associated pool of labeled MNC (CN-MAOM) was relatively small in all four studies (Fig. 3). Because of a new round of microbial biomass turnover, MIND model simulated that the recovery of 13C in CN-MAOM exhibited a slightly increasing trend, while FOND model simulated that the recovery of 13C in CN-MAOM did not change much with decomposition time.
Both models estimated that CN-MAOM had very slow decomposition rate, and the decomposition rate of the fast pool of microbial necromass C was at least 1000 times faster than that of the mineral-associated pool of microbial necromass C (Table 2, RMAOM-F being 1.08 × 10−7 to 8.26 × 10−3 for MIND model, and 6.14 × 10−5 to 1.89 × 10−3 for FOND model), which were different from the original estimations in these four studies.
Because CORPSE model is the only model that simulates “dead microbes” in current process-based models, we compared our estimated parameters to theirs (Supplementary Table 4) and found that the mean residence time of the mineral-associated microbial necromass estimated by our models (24–67 years) were similar to theirs (45–75 years) except for study 4.
About 44–99% of the microbial necromass C was transferred to the fast necromass C pool simulated by MIND model (Table 2, fBNF). About 35–88% of the microbial necromass C was transferred to the fast necromass C pool simulated by FOND model. Because a portion of microbial necromass C was transferred to DOC in FOND model, but remained in CNF and CN-MAOM in MIND model, it was reasonable that this fBNF was less estimated by FOND than by MIND model.
Model validation
Based on LOOCV and observed recovery of 13C in respired CO2, modeled and observed SOC pool and MBC pool at steady state in four studies, we found that both models had relatively high accuracy (Figs. 4 and 5 and Supplementary Figs. 6 and 7). The validation results of short-term studies studies 1 and 2 based on MIND and FOND were basically consistent. For the long-term studies study 3 and 4, their errors were greater than those of short-term ones, and the errors of FOND were larger than those of MIND. This result suggested that MIND had better simulation ability and our model had larger errors when simulating long-term observation data (Figs. 4 and 5). However, it is also possible that the observation data themselves had larger errors in the long-term experiments due to the changes in soil characteristics after long-term incubation. SOC was better simulated by both models than MBC (Supplementary Fig. 6).
The y-axis is the absolute value of the difference between simulation and observation. Box shows the upper and lower quartile with the line inside representing median. Whiskers show upper and lower extremes. Studies 1–4 are four decomposition experiments of 13C-labeled microbial necromass in different ecosystems and the details of the site information of studies 1–4 are in Supplementary Table 2.
In the four studies simulated by the two models, the percentage of MBC out of total SOC was between 1 and 1.6%, and the percentage of MNC out of total SOC was between 10 and 27% (Table 3 and Supplementary Table 5) at steady state. MIND and FOND models estimated different values for the proportion of MNC for study 1, but estimated similar values for the rest three studies.
Long-term and large-scale simulation results
Both models showed that the remaining microbial necromass 13C was within 0.15% after 1000 years of decomposition, and remaining microbial necromass 13C was within 20% after 100 years of decomposition (Fig. 6 and Supplementary Fig. 8). FOND model estimated that the averaged mean residence time (MRT) of necromass C was 11.1, 23.7, 21.6, and 64.5 years for studies 1–4, respectively.
We estimated the average MNC content (0–15 cm) of 13 biomes and calculated the proportion of MNC out of total SOC content (Supplementary Table 6). “Montane grasslands and shrublands” had the highest MNC/SOC (51%) and “Tropical and subtropical broadleaf forests” had the lowest MNC/SOC (2%). Overall, the proportion of MNC in SOC decreased from the cold areas to the tropical areas (Supplementary Table 6).
Comparison of simulation results between models with and without the microbial necromass pool
In order to evaluate if the addition of the microbial necromass pool in the models improved models’ performance, we used another two models without the microbial necromass pool (MIND-O and FOND-O corresponding to MIND and FOND, respectively) to simulate recovery of 13C in soil using data of the four studies. Results showed that MIND-O and FOND-O had very poor performance and the uncertainties were extremely high which cannot be presented in the figure (Supplementary Fig. 9 and Supplementary Table 7). Based on the four indexes of R2, RMSE and MAE, FOND and MIND models were generally better than the other two models (Table 1).
Sensitivity of model parameters
Sensitivity analysis showed that Vmax,P, KM, P were the two dominant parameters of the decomposition of organic matter in MIND model; CNF was most sensitive to Vmax,N, KM,N, fBNF, and kB; CN-MAOM was most sensitive to RMAOM-F because it determines the amount of microbial carbon input to the mineral-associated pool (Supplementary Table 8).
In FOND model, CP was most sensitive to kP; CD was most sensitive to kuptake; CNF was most sensitive to fBNF and kNF because they determine the proportion of microbial necromass input to CNF and the decomposition rate of CNF, respectively (Supplementary Table 8).
Discussion
Representation of the microbial necromass pool in process-based models
We found our newly proposed models representing the microbial necromass pool more effectively simulated the four experimental data set of the decomposition of 13C-labeled microbial necromass than models without the microbial necromass pool (Table 1). In models without the microbial necromass pool, microbial necromass was considered as a part of total soil organic matter and had the same decomposition pattern as plant-derived organic matter. Plant residues are mainly composed of polysaccharides and lignin, aliphatic biopolymers and tannins; while the cell walls of microbes are mainly composed of homopolysaccharides and heteropolysaccharides [56]. Some fungal cell walls also contain relatively high proportion of proteins, lipids and melanin [56]. Therefore, due to the difference in molecular structures of plant-derived and microbially derived organic matter, the decomposition pattern and rate are also expected to be different. Although the average decomposition rate of microbial necromass in soil was found to be faster than that of plant residues [8], some component of microbial necromass such as some glucosamine, galactosamine, or sorbic acid may accumulate for a long term in soil [56]. For example, melanin has been found to inhibit the decomposition of microbial cell wall [57], and more peptidoglycans in bacterial cell walls usually result in slower decomposition rates [56]. In addition, because microbes generally live on the surface of a particle in soil [11, 58, 59], microbial necromass is easily protected by minerals [11, 32, 60,61,62]. Once the necromass interacts with soil minerals, they can become inaccessible to microbes and have very slow decomposition rates [63, 64]. Microbial necromass has been found to bind to clay particles and promote the formation of micro-aggregates with <50 μm [65], and could be more tightly bound in soil matrix than plant debris [9, 66]. Therefore, the unprotected parts of microbial necromass (i.e., fast pool of microbial necromass) could be decomposed much more rapidly than the protected parts (i.e., mineral-associated necromass pool) [67, 68], showing different pattern from plant residues. The mineral-associated pool of microbial necromass had at least 1000 times slower decomposition rate than the fast pool in our simulation results, which supported this viewpoint. By adding the microbial necromass pool and dividing the pool into fast and mineral-associated components, process-based models can better simulate the decomposition pattern of SOC. Sulman et al. [30], in their model CORPSE, integrated two compartments of microbial necromass, called “unprotected dead microbe C” and “protected dead microbe C.” However, because the “protected dead microbe C” pool was not differentiated from “protected plant residue” and they had the same decomposition pattern and rates, we believe our model is the first attempt to realize this purpose of separately simulating microbial necromass C from plant-derived organic C decomposition processes. Between the two models representing the microbial necromass pool, the MIND model performed better than the FOND model. The reason was because microbial controls on SOC decomposition was better simulated in MIND model due to the consideration of microbial biomass effect on decomposition [3, 5, 21, 22, 69]. The Michaelis–Menten model has been found to better simulate the response of soil respiration to the drying-rewetting cycle than the first-order kinetic model because the consumption of SOC depends on microbial biomass [31]. We believe that by adding the microbial necromass pool to the Michaelis–Menten model, the paradoxical role of soil microbes in regulating SOC as both consumers of (via the Michaelis–Menten equation) and contributors to (via the necromass pool) SOC can be better represented and more accurately simulated. Furthermore, the Michaelis–Menten model may be more sensitive to disturbance such as warming and raising CO2 [3, 30]. Therefore, under the context of global changes, we believe MIND model considering microbial controls on SOC decomposition could be more effective in the long term.
Temporal variations of 13C recovery in MBC were better simulated by MIND model than by FOND model (Fig. 3) because 13C recovery in MBC was estimated to be 6.8–7% at day 224 by MIND model in study 1, which is similar to the PLFA measurement results of 11 ± 9% in the original study [60]. The 13C recovery in MBC only increased a little in FOND simulation results and remained very low in models without the necromass pool (Fig. 3), both of which seemed unrealistic.
Analysis of model parameters
In order to incorporate our model in Earth System Models, we need to know more on the major model parameters newly introduced in our model such as the decomposition rate of microbial necromass and the microbial CUE of microbial necromass. The Vmax and KM of the Michaelis–Menten equation for SOM decomposition given in previous studies were 0.01 mg C mg−1 MBC h−1 and 250 mg C g−1 soil, respectively [6, 26]. In MIND model, our results showed that Vmax,P was generally lower than 0.01 and Vmax,N was higher than 0.01 mg C mg−1 MBC h−1. Moreover, the KM,P and KM,N in MIND model were above and below previous estimation, respectively, except in study 4 (Table 2). We believe these results are reasonable because SOM is a mixture of plant-derived and microbially derived organic matter and the values of Vmax and KM for the whole SOM should be between the values for plant residue and those for microbial necromass. Vmax is the maximal reaction rate and decreases with decreasing substrate quality and quantity [70]. KM is an indicator of the affinity of an enzyme has for its substrate [71] and an increase in KM indicates a decrease in overall enzyme function [24]. Vmax and KM for microbial necromass should be higher and lower than Vmax and KM for plant residue, respectively, because most components of microbial necromass are faster than plant residue. In the first-order kinetic model, the decomposition rate of SOC was reported to be 5.6 × 10−6 mg C mg−1 C h−1 in previous studies [6, 26]. In FOND model, we found the decomposition rate of plant residue was approximately equal to this number but the decomposition of the fast pool of microbial necromass was higher than this value and the decomposition of the mineral-associated pool of microbial necromass was much lower than this value (Table 2, kP and kNF). This result further agrees with the viewpoint discussed above that the fast part of microbial necromass can be quickly used while the mineral-associated part of microbial necromass may accumulate for a very long period. RMAOM-F was the most sensitive parameter for the CN-MAOM pool (Supplementary Table 8), suggesting the importance of this parameter and should be studied in different soils and under different conditions in the future.
Additionally, we also compared other major parameters existing in previous models. The decomposition rate of microbially derived DOC (kDN) in our models was estimated to be 0.003–0.005 mg C mg−1 C h−1 (Table 2) and it is similar to the decomposition rate constant of DOC (kD) in other models, which was estimated to be 0.001 mg C mg−1 C h−1 in previous studies [6, 26]. It was slightly larger in our study, probably because that DOC from microbial necromass is easier to be protected by soil minerals than plant-derived DOC [9]. The turnover rate constant of microbial biomass (kB) was estimated to be 0.16–3.15 year−1 in our study (Table 2) and it was 2.45 year−1 in the previous modeling studies [6, 26] and 0.2–20 year−1 in previous observation studies [72, 73]. The fraction of fast content in microbial biomass (fBNF) was estimated to be 0.53 in FOND for study 1 and 0.52 in their original estimation [34]. Very few previous studies estimated this parameter, and the only available study estimated that the average fBNF was 0.51 based on their modeling of temperature sensitivity of microbial respiration [74]. In a different study, the proportion of microbial necromass C transferred to DOC was estimated to be 0.172 [62], which is lower than our estimation because our fast pool of microbial necromass contains not only DOC, but also other compounds whose decomposition rates may be lower than DOC.
Our estimation of the mean residence time of mineral-associated microbial necromass was 24–67 years for studies 1–3, which is similar to CORPSE model estimates (45–75 years) (Supplementary Table 4), suggesting the stability of microbial necromass once it is protected by soil minerals. For study 4, our estimates of the mean residence time of mineral-associated microbial necromass was very high (533 years), which was because that the protection rate of MNC was very low (kP in Supplementary Table 4, i.e., only a very small amount of microbial necromass entered the mineral-associated pool because the study was conducted in tropical areas). Therefore, although the decomposition rate of this pool was very low in study 4, the long-term accumulation (proportion) of MNC in SOC was comparable to other studies (Table 3). Overall, our estimated parameters were all within the reasonable range and future studies should be focused on how these parameters vary with temperature and moisture for model development.
Implication for long-term and large-scale modeling
After running the models for 1000 years, some 13C still remained in the soil (Fig. 6 and Supplementary Fig. 8) and the average MRT of microbial necromass was 11.1–64.5 years, suggesting the stabilization of microbial necromass. Although this result seems to be surprising, it is consistent with recent observation on microbial necromass decomposition, which found 33.1–39.5% of labeled necromass N remained in soil after 2.2 years of decomposition [8]. Under steady-state conditions, the estimated microbial necromass C accounted for 10–27% of the total SOC, which is at the lower end range of previous estimations, which were 24–80% [11, 15,16,17]. Some previous studies used the content of amino sugars in soil to estimate microbial necromass content and found microbial necromass C accounted for 23.6–58.3% of total SOC [16, 17]. However, the conversion factors used to convert amino sugar to necromass are still under debate. Other previous studies used the relative abundance of fatty acids and amino acids in microbial biomass and estimated the fraction of necromass C to SOC to be about 40% [11]. Fatty acids and amino acids exist not only in microbial necromass, but also in living microbial cells, plant cells, and root exudates [75, 76], and the fraction of necromass C to SOC was probably overestimated based on this method and therefore higher than our estimations. Sulman et al. [30] estimated that “dead microbe carbon” was 0.34% of total SOC based on CORPSE model, but their “dead microbe carbon” pool corresponds to our labile MNC pool because once microbial carbon enters the “protected carbon” pool (which is the sum of chemical resistant carbon, simple carbon, and dead microbe carbon), they are no longer counted as “dead microbe carbon” in their model. Our labile MNC pool was 0.01–0.06% of total SOC, which is comparable to their estimation.
Alternatively, it is possible that the different findings among different studies were due to site difference because the contribution of microbial necromass to soil organic C is affected by many factors, such as soil pH, salinity, clay content, fungal bacterial ratio, environmental stress, ecological type, land use change, soil C:N, microbial C:N, and nutrient status [16, 61, 77]. But it should be noted that of 13C-labeled necromass decomposition studies [33,34,35] used for our models only contain ~20 isolates, and therefore the necromass C decomposition rate for the whole microbial community should be examined in future studies.
Our estimation of the proportion of MNC out of total SOC content in 13 biomes suggested that “montane grasslands and shrublands” and “tundra” biomes had highest proportion of necromass C in SOC (31–51%, Supplementary Table 6), which is close to an estimation based on soil amino sugar content in the Tibet Plateau [78]. Generally, these areas have high altitude and low temperature, and the plant productivity is low [79, 80]. Due to the higher microbial CUE [81] and the better adaptability of microbes than plants under extreme conditions, it is reasonable that the proportion of microbial necromass out of total soil organic matter is high. In tropical areas, plant productivity is high and microbial CUE is low [81], microbial necromass C was estimated to be <14% of SOC by our models (Supplementary Table 6). Previous experiments also found that the contribution of microbial residues to SOC decreased from tropical to boreal forests [82]. Although these results at the large spatial scale are preliminary because the estimates of some parameters were based on four observation studies, it can still shed light on the importance of microbial necromass pool and the variations among different biomes.
A very interesting phenomenon is: century-scale projection using MIND showed some oscillation compared to FOND (Fig. 6). Usually more C pools in Michaelis–Menten-based microbial models would reduce such oscillation [26], and Georgiou et al. [6] reduced this oscillation by considering the turnover of microorganisms at the population level. We hope the addition of the necromass pool could also realize it and found that changes in carbon input, temperature and microbial CUE [6, 26] could still induce oscillation of SOC simulation results for both MIND and FOND models although the addition of necromass pool alleviated such oscillation. The amplitude and period of the oscillation depended on microbial CUE, Vmax and KM [83]. Future studies considering the turnover of microorganisms at the population level [6] may reduce the oscillation and further improve model performance.
Limitations and future studies
At present, only four sets of observation data based on the decomposition experiments of the 13C-labeled microbial necromass were available to calibrate our models. Although the four studies were conducted in different ecosystems and somewhat represent different conditions, more experiments are needed for better parameterization of the models. In addition, the four observation data were based on both lab incubation experiments (studies 1 and 2) and field experiments (studies 3 and 4) and whether or not the experiments were conducted in the field is important. In the future, more field-based microbial necromass decomposition studies are needed to better parameterize the model and better understand the behavior of microbial necromass decomposition. Especially, experiments under the changing environment are needed to test whether model behavior under climate change is consistent with observation and this will be the immediate next step of our work.
The number of observation on some parameters like microbial CUE has increased tremendously in recent 5 years [46, 84]. More direct measurements of the decomposition parameters for the microbial necromass pool (i.e., the decomposition rate of microbial necromass, the turnover rate of microbial biomass, etc.) will substantially increase the effort of parameterization. Previous studies suggested that the decomposition rates of microbial necromass nitrogen from different microbial groups were not significantly different [8, 32], suggesting that substrate quality may not influence the decomposition rates of microbial necromass. However, soil properties such as soil temperature, soil moisture, and soil clay content may influence these parameters and more experiments are needed to estimate these parameters under different conditions.
Nevertheless, our study provides an independent model estimation of microbial necromass C contribution to stable SOC, supplementary to previous model estimations [15, 85, 86]. Although slighter lower than previous estimations, this significant contribution ratio further proved that it is necessary to incorporate the microbial necromass pool in Earth System Models to better simulate carbon cycling under the context of global changes.
Data availability
The data used can be found in Supplementary Information.
Code availability
Code used to model runs is available at https://github.com/fanlei21/fanlei21.github.io.
References
Hiederer R, Köchy M. Global soil organic carbon estimates and the harmonized world soil database. EUR. 2011;79:25225.
Scharlemann JPW, Tanner EVJ, Hiederer R, Kapos V. Global soil carbon: understanding and managing the largest terrestrial carbon pool. Carbon Manag. 2014;5:81–91.
Wieder WR, Bonan GB, Allison SD. Global soil carbon projections are improved by modelling microbial processes. Nat Clim Chang. 2013;3:909–12.
Schimel JP, Weintraub MN. The implications of exoenzyme activity on microbial carbon and nitrogen limitation in soil: a theoretical model. Soil Biol Biochem. 2003;35:549–63.
Huang Y, Guenet B, Ciais P, Janssens IA, Soong JL, Wang Y, et al. ORCHIMIC (v1.0), a microbe-mediated model for soil organic matter decomposition. Geosci Model Dev. 2018;11:2111–38.
Georgiou K, Abramoff RZ, Harte J, Riley WJ, Torn MS. Microbial community-level regulation explains soil carbon responses to long-term litter manipulations. Nat Commun. 2017;8:1223.
Kelleher BP, Simpson AJ. Humic substances in soils: are they really chemically distinct? Environ Sci Technol. 2006;40:4605–11.
Wang C, Wang X, Pei G, Xia Z, Peng B, Sun L, et al. Stabilization of microbial residues in soil organic matter after two years of decomposition. Soil Biol Biochem. 2020;141:107687.
Cotrufo MF, Wallenstein M, Boot C, Denef K, Paul E. The microbial efficiency-matrix stabilization (MEMS) framework integrates plant litter decomposition with soil organic matter stabilization: do labile plant inputs form stable soil organic matter? Glob Change Biol. 2013;19:988–95.
Zhu X, Jackson RD, DeLucia EH, Tiedje JM, Liang C. The soil microbial carbon pump: from conceptual insights to empirical assessments. Glob Change Biol. 2020;26:6032–9.
Miltner A, Bombach P, Schmidt-Brücken B, Kästner M. SOM genesis: microbial biomass as a significant source. Biogeochemistry. 2012;111:41–55.
Torn MS, Trumbore SE, Chadwick OA, Vitousek PM, Hendricks DM. Mineral control of soil organic carbon storage and turnover. Nature. 1997;389:170–3.
Dwivedi D, Riley WJ, Torn MS, Spycher N, Maggi F, Tang JY. Mineral properties, microbes, transport, and plant-input profiles control vertical distribution and age of soil carbon stocks. Soil Biol Biochem. 2017;107:244–59.
Mikutta R, Kleber M, Torn MS, Jahn R. Stabilization of soil organic matter: association with minerals or chemical recalcitrance? Biogeochemistry. 2006;77:25–56.
Liang C, Balser TC. Microbial production of recalcitrant organic matter in global soils: Implications for productivity and climate policy. Nat Rev Microbiol. 2011;9:75–75.
Khan KS, Mack R, Castillo X, Kaiser M, Joergensen RG. Microbial biomass, fungal and bacterial residues, and their relationships to the soil organic matter C/N/P/S ratios. Geoderma. 2016;271:115–23.
Liang C, Amelung W, Lehmann J, Kästner M. Quantitative assessment of microbial necromass contribution to soil organic matter. Glob Chang Biol. 2019;25:3578–90.
Kögel-Knabner I. The macromolecular organic composition of plant and microbial residues as inputs to soil organic matter: fourteen years on. Soil Biol Biochem. 2017;105:A3–8.
Todd-Brown KEO, Randerson JT, Post WM, Hoffman FM, Tarnocai C, Schuur EAG, et al. Causes of variation in soil carbon simulations from CMIP5 Earth System Models and comparison with observations. Biogeosciences. 2013;10:1717–36.
Parton WJ, Schimel DS, Cole CV, Ojima DS. Analysis of factors controlling soil organic matter levels in great plains grasslands. Soil Sci Soc Am J. 1987;51:1173–9.
Wang G, Post WM, Mayes MA. Development of microbial‐enzyme‐mediated decomposition model parameters through steady‐state and dynamic analyses. Ecol Appl. 2013;23:255–72.
Wang G, Mayes MA, Gu L, Schadt CW. Representation of dormant and active microbial dynamics for ecosystem modeling. PLoS ONE. 2014;9:e89252.
Wang G, Jagadamma S, Mayes MA, Schadt CW, Steinweg JM, Gu L, et al. Microbial dormancy improves development and experimental validation of ecosystem model. ISME J. 2015;9:226–37.
German D, Marcelo K, Stone M, Allison S. The Michaelis–Menten kinetics of soil extracellular enzymes in response to temperature: a cross-latitudinal study. Glob Change Biol. 2012;18:1468–79.
Allison SD, Wallenstein MD, Bradford MA. Soil-carbon response to warming dependent on microbial physiology. Nat Geosci. 2010;3:336–40.
Li J, Wang G, Allison SD, Mayes MA, Luo Y. Soil carbon sensitivity to temperature and carbon use efficiency compared across microbial-ecosystem models of varying complexity. Biogeochemistry. 2014;119:67–84.
Wieder WR, Grandy AS, Kallenbach CM, Bonan GB. Integrating microbial physiology and physio-chemical principles in soils with the MIcrobial-MIneral Carbon Stabilization (MIMICS) model. Biogeosciences. 2014;11:3899–917.
Tang J, Riley WJ. Weaker soil carbon–climate feedbacks resulting from microbial and abiotic interactions. Nat Clim Chang. 2015;5:56–60.
Sulman BN, Moore JA, Abramoff R, Averill C, Kivlin S, Georgiou K, et al. Multiple models and experiments underscore large uncertainty in soil carbon dynamics. Biogeochemistry. 2018;141:109–23.
Sulman BN, Phillips RP, Oishi AC, Shevliakova E, Pacala SW. Microbe-driven turnover offsets mineral-mediated storage of soil carbon under elevated CO2. Nat Clim Change. 2014;4:1099–102.
Lawrence C, Neff J, Schimel J. Does adding microbial mechanisms of decomposition improve soil organic matter models? A comparison of four models using data from a pulsed rewetting experiment. Soil Biol Biochem. 2009;41:1923–34.
Wang X, Wang C, Cotrufo MF, Sun L, Jiang P, Liu Z, et al. Elevated temperature increases the accumulation of microbial necromass nitrogen in soil via increasing microbial turnover. Glob Change Biol. 2020;26:5277–89.
Throckmorton HM, Bird JA, Dane L, Firestone MK, Horwath WR. The source of microbial C has little impact on soil organic matter stabilisation in forest ecosystems. Ecol Lett. 2012;15:1257–65.
Kindler R, Miltner A, Richnow H-H, Kästner M. Fate of gram-negative bacterial biomass in soil—mineralization and contribution to SOM. Soil Biol Biochem. 2006;38:2860–70.
Schweigert M, Herrmann S, Miltner A, Fester T, Kästner M. Fate of ectomycorrhizal fungal biomass in a soil bioreactor system and its contribution to soil organic matter formation. Soil Biol Biochem. 2015;88:120–7.
Derrien D, Amelung W. Computing the mean residence time of soil carbon fractions using stable isotopes: impacts of the model framework. Eur J Soil Sci. 2011;62:237–52.
Dormand JR, Prince PJ. A family of embedded Runge-Kutta formulae. J Comput Appl Math. 1980;6:19–26.
Shampine LF, Reichelt MW. The MATLAB ODE suite. Siam J Sci Comput. 1997;18:1–22.
Coleman TF, Li Y. On the convergence of reflective newton methods for large-scale nonlinear minimization subject to bounds. Math Program. 1994;67:189–224.
Coleman TF, Li Y. An interior trust region approach for nonlinear minimization subject to bounds. SIAM J Optim. 1996;6:418–45.
Moré JJ. The Levenberg–Marquardt algorithm: implementation and theory. In: Watson GA (ed). Numerical Analysis. Springer: Berlin, Heidelberg, 1978, p. 105–16.
Leave-one-out cross-validation. In: Sammut C, Webb GI, editors. Encyclopedia of machine learning. Boston, MA: Springer USA; 2010. p. 600–1.
Wang C, Qu L, Yang L, Liu D, Morrissey E, Miao R, et al. Large-scale importance of microbial carbon use efficiency and necromass to soil organic carbon. Glob Chang Biol. 2021.
Farrell M, Prendergast-Miller M, Jones DL, Hill PW, Condron LM. Soil microbial organic nitrogen uptake is regulated by carbon availability. Soil Biol Biochem. 2014;77:261–7.
Hagerty SB, Allison SD, Schimel JP. Evaluating soil microbial carbon use efficiency explicitly as a function of cellular processes: implications for measurements and models. Biogeochemistry. 2018;140:269–83.
Qiao Y, Wang J, Liang G, Du Z, Zhou J, Zhu C, et al. Global variation of soil microbial carbon-use efficiency in relation to growth temperature and substrate supply. Sci Rep. 2019;9:5621.
Krinner G, Viovy N, de Noblet-Ducoudré N, Ogée J, Polcher J, Friedlingstein P, et al. A dynamic global vegetation model for studies of the coupled atmosphere-biosphere system. Glob Biogeochem Cycles. 2005;19:GB1015.
Wang G, Post WM, Mayes MA, Frerichs JT, Sindhu J. Parameter estimation for models of ligninolytic and cellulolytic enzyme kinetics. Soil Biol Biochem. 2012;48:28–38.
Davidson EA, Janssens IA. Temperature sensitivity of soil carbon decomposition and feedbacks to climate change. Nature. 2006;440:165–73.
Fick SE, Hijmans RJ. WorldClim 2: new 1-km spatial resolution climate surfaces for global land areas. Int J Climatol. 2017;37:4302–15.
Guevara M, Taufer M, Vargas R. Gap-free global annual soil moisture: 15 km grids for 1991–2018. Earth Syst Sci Data. 2020;2020:1–65.
Kalnay E, Kanamitsu M, Kistler R, Collins W, Deaven D, Gandin L, et al. The NCEP/NCAR 40-year reanalysis project. Bull Am Meteorol Soc. 1996;77:437–72.
Batjes NH. Harmonized soil property values for broad-scale modelling (WISE30sec) with estimates of global soil carbon stocks. Geoderma. 2016;269:61–8.
Hengl T, Mendes de Jesus J, Heuvelink GBM, Ruiperez Gonzalez M, Kilibarda M, Blagotić A, et al. SoilGrids250m: global gridded soil information based on machine learning. PLoS ONE. 2017;12:e0169748.
Olson DM, Dinerstein E. The Global 200: a representation approach to conserving the earth’s most biologically valuable ecoregions. Conserv Biol. 1998;12:502–15.
Kögel-Knabner I. The macromolecular organic composition of plant and microbial residues as inputs to soil organic matter. Soil Biol Biochem. 2002;34:139–62.
Fernandez CW, Koide RT. Initial melanin and nitrogen concentrations control the decomposition of ectomycorrhizal fungal litter. Soil Biol Biochem. 2014;77:150–7.
Hemkemeyer M, Dohrmann AB, Christensen BT, Tebbe CC. Bacterial preferences for specific soil particle size fractions revealed by community analyses. Front Microbiol. 2018;9:149.
Mills A. Keeping in touch: microbial life on soil particle surfaces. Adv Agron. 2003;78:1–43.
Kindler R, Miltner A, Thullner M, Richnow H-H, Kästner M. Fate of bacterial biomass derived fatty acids in soil and their contribution to soil organic matter. Org Geochem. 2009;40:29–37.
Huang Y, Liang C, Duan X, Chen H, Li D. Variation of microbial residue contribution to soil organic carbon sequestration following land use change in a subtropical karst region. Geoderma. 2019;353:340–6.
Ahrens B, Braakhekke MC, Guggenberger G, Schrumpf M, Reichstein M. Contribution of sorption, DOC transport and microbial interactions to the 14C age of a soil organic carbon profile: insights from a calibrated process model. Soil Biol Biochem. 2015;88:390–402.
Nguyen RT, Harvey HR. Preservation via macromolecular associations during Botryococcus braunii decay: proteins in the Pula Kerogen. Org Geochem. 2003;34:1391–403.
Kallenbach CM, Frey SD, Grandy AS. Direct evidence for microbial-derived soil organic matter formation and its ecophysiological controls. Nat Commun. 2016;7:13630.
Puget P, Angers DA, Chenu C. Nature of carbohydrates associated with water-stable aggregates of two cultivated soils. Soil Biol Biochem. 1998;31:55–63.
Schmidt MWI, Torn MS, Abiven S, Dittmar T, Guggenberger G, Janssens IA, et al. Persistence of soil organic matter as an ecosystem property. Nature. 2011;478:49–56.
Spence A, Simpson AJ, McNally DJ, Moran BW, McCaul MV, Hart K, et al. The degradation characteristics of microbial biomass in soil. Geochim Cosmochim Acta. 2011;75:2571–81.
Drigo B, Anderson IC, Kannangara GSK, Cairney JWG, Johnson D. Rapid incorporation of carbon from ectomycorrhizal mycelial necromass into soil fungal communities. Soil Biol Biochem. 2012;49:4–10.
Wang G, Chen S. A review on parameterization and uncertainty in modeling greenhouse gas emissions from soil. Geoderma. 2012;170:206–16.
Blagodatskaya Е, Blagodatsky S, Khomyakov N, Myachina O, Kuzyakov Y. Temperature sensitivity and enzymatic mechanisms of soil organic matter decomposition along an altitudinal gradient on Mount Kilimanjaro. Sci Rep. 2016;6:22240.
German DP, Weintraub MN, Grandy AS, Lauber CL, Rinkes ZL, Allison SD. Optimization of hydrolytic and oxidative enzyme methods for ecosystem studies. Soil Biol Biochem. 2011;43:1387–97.
Wu J, Xiao H. Measuring the gross turnover time of soil microbial biomass C under incubation. Acta Pedol Sin. 2004;41:401–7.
Cheng W. Rhizosphere priming effect: Its functional relationships with microbial turnover, evapotranspiration, and C–N budgets. Soil Biol Biochem. 2009;41:1795–801.
Luo Z, Tang Z, Guo X, Jiang J, Sun OJ. Non-monotonic and distinct temperature responses of respiration of soil microbial functional groups. Soil Biol Biochem. 2020;148:107902.
de Graaff M-A, Classen AT, Castro HF, Schadt CW. Labile soil carbon inputs mediate the soil microbial community composition and plant residue decomposition rates. New Phytol. 2010;188:1055–64.
Paul EA. The nature and dynamics of soil organic matter: plant inputs, microbial transformations, and organic matter stabilization. Soil Biol Biochem. 2016;98:109–26.
Crowther TW, Sokol NW, Oldfield EE, Maynard DS, Thomas SM, Bradford MA. Environmental stress response limits microbial necromass contributions to soil organic carbon. Soil Biol Biochem. 2015;85:153–61.
Ding X, Chen S, Zhang B, He H, Filley TR, Horwath WR. Warming yields distinct accumulation patterns of microbial residues in dry and wet alpine grasslands on the Qinghai-Tibetan Plateau. Biol Fertil Soils. 2020;56:881–92.
Mao D, Luo L, Wang Z, Zhang C, Ren C. Variations in net primary productivity and its relationships with warming climate in the permafrost zone of the Tibetan Plateau. J Geogr Sci. 2015;25:967–77.
Wu J, Feng Y, Zhang X, Wurst S, Tietjen B, Tarolli P, et al. Grazing exclusion by fencing non-linearly restored the degraded alpine grasslands on the Tibetan Plateau. Sci Rep. 2017;7:15202.
Li J, Wang G, Mayes MA, Allison SD, Frey SD, Shi Z, et al. Reduced carbon use efficiency and increased microbial turnover with soil warming. Glob Change Biol. 2019;25:900–10.
Chen G, Ma S, Tian D, Xiao W, Jiang L, Xing A, et al. Patterns and determinants of soil microbial residues from tropical to boreal forests. Soil Biol Biochem. 2020;151:108059.
Wang YP, Chen BC, Wieder WR, Leite M, Medlyn BE, Rasmussen M, et al. Oscillatory behavior of two nonlinear microbial models of soil carbon decomposition. Biogeosciences. 2014;11:1817–31.
Soares M, Rousk J. Microbial growth and carbon use efficiency in soil: links to fungal-bacterial dominance, SOC-quality and stoichiometry. Soil Biol Biochem. 2019;131:195–205.
Liang C, Cheng G, Wixon DL, Balser TC. An Absorbing Markov Chain approach to understanding the microbial role in soil carbon stabilization. Biogeochemistry. 2011;106:303–9.
Fan Z, Liang C. Significance of microbial asynchronous anabolism to soil carbon dynamics driven by litter inputs. Sci Rep. 2015;5:9575.
Acknowledgements
This study was funded by the National Key R&D Program of China (2019YFA0607301), the National Natural Science Foundation of China (No. 41971058), and the National Program for Support of Top-notch Young Professionals (to EB).
Author information
Authors and Affiliations
Contributions
XF and EB designed the study and analyzed the experiments and wrote and edited the paper. XF collected and organized the data and built and run the model. DG, CZ, CW, YQ, and JZ contributed to the discussion of results and revision of the paper.
Corresponding author
Ethics declarations
Conflict of interest
The authors declare no competing interests.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
About this article
Cite this article
Fan, X., Gao, D., Zhao, C. et al. Improved model simulation of soil carbon cycling by representing the microbially derived organic carbon pool. ISME J 15, 2248–2263 (2021). https://doi.org/10.1038/s41396-021-00914-0
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41396-021-00914-0
This article is cited by
-
Soil bacteria mediate organic carbon stability during alpine wetland biogeomorphic succession in the arid region of Central Asia
Plant and Soil (2024)
-
The structure and development of Loess Critical Zone and its soil carbon cycle
Carbon Neutrality (2024)
-
Integrating microbial community properties, biomass and necromass to predict cropland soil organic carbon
ISME Communications (2023)
-
Differential effects of nitrogen addition on the organic carbon fractions of rhizosphere and bulk soil based on a pot experiment
Journal of Soils and Sediments (2023)
-
Microbial properties determine dynamics of topsoil organic carbon stocks and fractions along an age-sequence of Mongolian pine plantations
Plant and Soil (2023)