Abstract
Context
Ensemble of small models (ESMs) is a technique to overcome the problem of few occurrence points. Applying the ESMs in a spatially hierarchical framework could increase the accuracy of predictions and conclusions by restricting available habitat at sequentially finer spatial scales.
Objectives
Our objective was to show how applying ESMs in a hierarchical habitat selection framework could help to understand rare species’ niches at various scales. We compared the accuracy of ESMs made by committee averaging and weighted averaging methods. We also compared the predictive power of ESMs made by various modeling techniques for Virginia’s warbler (Leiothlypis virginiae) at its northeastern range limit.
Methods
We defined biologically relevant hierarchical orders of habitat selection for Virginia’s warbler in the Black Hills, U.S.A. We modeled habitat suitabity at the broadest scale as a function of bioclimatic covariates and at finer scales as functions of landcover, soil group and landscape covariates.
Results
The performance of modeling techniques varied among scales. Using the committee averaging method led to more accurate results than weighted averaging. At the broadest order, Virginia’s warbler had a narrow climatic niche. The importance of covariates changed across finer orders, such that at broader orders many covariates were important whereas at finer orders certain covariates became more important than others.
Conclusion
We conclude that applying ESMs within a hierarchical framework can lead to detailed information about rare species’ niches, limiting factors at each habitat selection order, and potential distribution, which could help inform multiscale management.
Similar content being viewed by others
Introduction
Recognizing the spatial distribution of species is one of the primary steps in their conservation and management. Species Distribution Models (SDMs) are powerful tools for estimating habitat preferences and for projecting species realized niches onto geographic space (Guisan et al. 2013, 2017). However, application of SDMs for rare species has been challenging because the accuracy of models decreases with lower numbers of occurrence records (Hernandez et al. 2006). Inclusion of a high number of covariates in models with few occurrences increases the risk of overfitting and collinearity between variables (Dormann et al. 2013). These problems reduce the ability to generalize from the model. One approach to address the overfitting problem is to reduce the number of covariates in the model to those with greatest probable contributions to model fit by relying on expert knowledge about the species’ niche, but such knowledge may not be available for rare species. In addition, excluding covariates for better generalizability could reduce the chances of capturing more dimensions of a species’ niche and limit our understanding of niche requirements (Guillera-Arroita et al. 2015).
Lomba et al. (2010) introduced a new strategy to deal with overfitting in rare species distribution modeling which appeared to result in more accurate spatial projections. This strategy is based on fitting bivariate models for all binary combinations of covariates and creating an ensemble prediction by weighted averaging of predictions based on their performances. They proposed their strategy in a spatially hierarchical framework in which the climatic niche was modeled at the scale of the entire species’ range and then was filtered by regional model predictions based on finer scale covariates. Breiner et al. (2015, 2018) named this approach as Ensembles of Small Models (ESMs) and found that this approach outperformed a wide range of standard SDMs for rare species. They recommended use of tuned ESMs made by Artificial Neural Networks (ANN) or Generalized Boosted Models (GBM) when the goal is high predictive performance and use of ESMs-Maxent with tuned features when projection to other regions is the main objective. Assuming that these recommendations also lead to the best results for other taxa, ESMs have been applied to model habitat suitability of some rare species of insects (Della Rocca et al. 2019), Caucasian grouse (Habibzadeh and Ludwig 2019), bats (Scherrer et al. 2019), and invasive squirrels (Di Febbraro et al. 2019), but ESMs were not compared with other modeling approaches in these studies. Moreover, the modeling recommendations of Breiner et al. (2015, 2018) were based on ESMs made for some rare plant species and may not lead to the best results for other taxa, especially for highly mobile species (Guisan and Thuiller 2005). In addition, committee averaging as an alternative for weighted averaging has not been previously employed in ESMs studies. One of the advantages of the committee averaging method is that it provides a measure of uncertainty along with predictions. Predictions near 0 or 1 mean that all models agree to predict absence or presence, respectively, and a prediction around 0.5 means that half of the models predict absence and half predict presence (Araújo and New 2007).
Rare species often have restricted ranges and do not occupy all available suitable habitat, leading to a patchy distribution (Pacifici et al. 2012). Field surveys for detecting rare species are mostly conducted in areas where the presence of the species had been previously detected. Thus, occurrence records may not represent the full range of the species niche nor do absences necessarily mean that habitat is unsuitable (Guisan et al. 2006; McCune 2016). One approach to deal with such imbalanced data is to apply presence-background modeling techniques (Guisan et al. 2017). However, disregarding true absence points obtained from field surveys can lead to loss of useful information (Guillera-Arroita et al. 2015). The geographic extent of the background, from which pseudo-absence points should be selected, can influence model parametrization and the accuracy of the predictions (Merow et al. 2013). Many species, especially highly mobile species like birds, perceive the landscape at multiple scales when selecting habitat and restrict available habitat based on information gathered at each scale (Mayor et al. 2009; Zimmerman et al. 2009; Jedlikowski et al. 2016). Having a restricted range and being a habitat specialist (because of specific habitat requirements, interspecific competition, etc.) could imply that hierarchical habitat selection is more focused on specific habitat features and, thereby, is more pronounced for rare than for generalist species (Devictor et al. 2008; Razgour et al. 2011). Applying the ESMs approach in a spatially hierarchical framework based on habitat selection concepts (Johnson 1980) could increase the accuracy of predictions by restricting available habitat (i.e., background extent) sequentially from coarse to finer spatial scales. Such an approach also allows incorporation of both true and pseudo-absences and increases chances of capturing more dimensions of the niche of rare species (Lomba et al. 2010; McGarigal et al. 2016).
In this study, we apply ESMs and a hierarchical Bayesian approach in a spatially hierarchical framework to model breeding habitat suitability of Virginia’s warbler (Leiothlypis virginiae) in the Black Hills of South Dakota and Wyoming. The breeding range of Virginia’s warbler covers mountainous regions of the southwestern U.S and extreme northern Mexico (Olson and Martin 1999). Previous studies indicate that the northeastern-most breeding population is isolated in the southern Black Hills, where it is considered rare and a species of conservation concern (Swanson et al. 2000, 2016). Genetic data are consistent with recent range expansion of Virginia’s warblers into the Black Hills (Bubac and Spellman 2016), which suggests that not all suitable habitat in the Black Hills may be currently occupied. We employ the hierarchical habitat selection concept (Johnson 1980) to increase the accuracy of model predictions and show how this approach could help represent habitat selection of a rare species at various scales. Subsequently, we compare the accuracy of ESMs made by the weighted averaging method with those made by the committee averaging method. We also compare the predictive power of ESMs made by various modeling techniques (ANN, GLM, GBM, GAM, RF, Maxent, and MARS).
Materials and methods
Study area and occurrence records
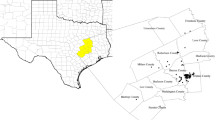
The study was conducted in the Black Hills which is an isolated mountain range located in western South Dakota and extending into northeastern Wyoming, U.S.A. The dominant tree species over much of the Black Hills is ponderosa pine (Pinus ponderosa), with small areas of white spruce (Picea glauca), paper birch (Betula papyrifera) and quaking aspen (Populus tremuloides) mainly at higher elevations or along canyon bottoms (Orr 1975). Previous studies documented that Virginia’s warbler in South Dakota exists in a unique pine-juniper-shrub habitat within the southwestern Black Hills. This habitat occurs on steep rocky terrain covered by scattered ponderosa pine and/or Rocky Mountain juniper (Juniperus scopulorum) as dominant overstory species, with a shrubby understory of mountain mahogany (Cercocarpus montanus) and skunkbush sumac (Rhus aromatica) (Swanson et al. 2016). We used 76 presence and 114 absence points from point count surveys (with the aid of broadcast songs) in July 2016 and June 2018 to build subsequent models (Fig. 1; see Appendix S1 in supplementary materials for details).
Modeling framework
Following Johnson (1980) and Meyer and Thuiller (2006), we considered four hierarchical orders of habitat selection. From broad to fine scales, these included, Order 0: Species range order; Order 1: Population order; Order 2: Individual order; and Order 3: Individual life requirement order. For each order we defined (see below for details) an extent (available habitat). For population and individual orders, we used suitable habitats defined for the preceding order as their extents and for the individual life requirement order, we used our presence-absence points rather than defining an a priori extent. For the population order we selected the optimum scale-of-effect from a range of scales (Jackson and Fahrig 2015). For the species range order, we fixed the scale of analysis to incorporate the known range of the Virginia’s warbler; for individual and individual life requirements orders, we fixed the scale of analysis at 90-m radii to reflect the home range size of Virginia’s warbler (see below for details).
Habitat selection extents
As the broadest scale, the species range order accounts for the geographic distribution of a species within a region. We assumed that Virginia’s warbler has a narrow niche and its potential suitable habitats at the broadest scale are limited. Thus, we applied a 75-km buffer around the Black Hills National Forest as the species range order extent, since such a buffer includes all the preferred pine-juniper-shrub habitats of the Virginia’s warbler within the Black Hills region (Swanson et al. 2000). Because habitat selection at finer scales is constrained by selection at broader scales (Manly et al. 2007), the next two extents (population and individual orders) were defined based on suitable habitats recognized from their previous scales (i.e., suitable habitats of species range order as the extent of population order and suitable habitats of population order as the extent of individual order). The population order extent accounts for selection of habitats by populations within the geographic range. Previous studies identified the southwestern Black Hills as suitable habitat for Virginia’s warbler populations (Swanson et al. 2016). We considered the population order based on the assumption that potential suitable habitats other than current known habitats for Virginia’s warbler populations may exist within the species range order because of recent colonization of the Black Hills so that not all suitable habitats are occupied (Bubac and Spellman 2016). The individual order extent accounts for home range requirements. This order was considered based on the assumption that requirements for Virginia’s warbler at the home range scale may only occur at a subset (discrete patches) of the entire population order extent, resulting in patchy home ranges. The individual life requirement order accounts for life requirements within home range patches. We assumed that there should be some specific habitat requirements in an area to be considered as a home range by Virginia’s warbler. For this extent we used our real absence points obtained from our surveys rather than deriving pseudo-absence points from the available habitat (extent).
Independent variables and scale-of-effect
Climate conditions govern distribution of birds at broad scales directly (Root 1988; Gaston and Blackburn 2008) or through formation of plant communities and food availability (Suárez-Seoane et al. 2002). Thus, for the species range order, we employed WorldClim data, version 2, with a 1-km spatial resolution because this dataset is one of the most recent and widely used climate datasets and provides climate data at a spatial resolution appropriate for the modeling approach that we employed. In addition to 19 conventional bioclimatic variables that represent annual data, we used mean monthly climate data for minimum, average, and maximum temperatures and for precipitation for June and July of 1970–2000 (Fick and Hijmans 2017; Appendix S2).
Habitat components may have different functions in defining the species niche at different scales if limiting factors change with scale (Rettie and Messier 2000; Fletcher and Fortin 2018). Hence, at finer scales than species range order, based on our literature review of Virginia’s warbler habitat associations, we considered a suite of landcover, soil groups and landscape covariates (Appendix S2). We expected that like many other terrestrial species (Boyce et al. 2003; Martínez et al. 2003), decision making by Virginia’s warbler regarding importance of habitat covariates is more general at broader than at finer scales. Also, following Rettie and Messier (2000) and Fletcher and Fortin (2018), we hypothesized that at broader scales Virginia’s warbler spatially avoids those factors that most strongly limit fitness. For instance, we expect this species to avoid grassland-dominated, closed canopy cover and mesic habitats at broader scales since Virginia’s warbler exists in a unique pine-juniper-shrub habitat in the Black Hills (Swanson et al. 2016). At finer scales, we expect factors with less importance at broader scales to become more important because limiting factors at broader scales have already been avoided. For example, at finer scales, vegetation cover, slope, and mountain mahogany density should have higher ranks of variable importance than density of grasslands and high canopy cover, which have already been avoided at broader scales.
To determine the optimal scale-of-effect for the population order, we followed Bellamy et al. (2013) in examining various scales-of-effect. We calculated the proportion of each category for categorical covariates (vegetation types and soil groups) and the mean of continuous covariates within concentric windows around each pixel using the focal statistics tool in ArcGIS 10.7 (ESRI 2017). The radius of windows ranged from 90 m, which is consistent with the maximum Virginia’s warbler home range radius (Fischer 1978), to 990 m (approximate species range order scale-of-effect), with an interval of 90 m. To identify the best window radius for each covariate, we developed univariate models for each combination of covariate and window radius using Maxent 3.3.3 (Phillips et al. 2006) with the R-package BIOMOD2 (Thuiller et al. 2009). The maximum number of background points was randomly selected within the population range extent such that the minimum distance between points was 30 m, so that only one point was included for a 30 × 30 m pixel. Models were cross validated using 50 split-sample runs (80% of points for training and 20% for testing). The best window radius for each covariate was selected based on the area under the curve (AUC) from receiver operation-characteristic (ROC) plots (Fielding and Bell 1997; Appendix S3).
For the individual and individual life requirements orders, we considered the maximum home range size for Virginia’s warbler (90-m radius, Fischer 1978) as the scale-of-effect because the goal was to compare used and unused sites at the home range scale. We calculated the proportion of categorial covariates and the mean of continuous covariates within this radius for each occurrence record.
Multicollinearity
To avoid multicollinearity, we ran a hierarchical clustering method based on a distance measure (1—the absolute value of the Pearson’s product-moment correlation) between variables using R-package KlaR (Weihs et al. 2005) to group correlated variables (correlation > 0.7; Dormann et al. 2013) into clusters. We then excluded binary models from covariates within clusters from the suite of all possible binary combinations of variables. Using a hierarchical clustering method instead of removing one of the paired correlated variables allowed us to investigate the role of all candidate covariates in the Virginia’s warbler’s niche (Appendix S4).
Modeling
We developed a suite of ESMs with seven modeling techniques for species range, population, and individual orders. We developed generalized linear models (GLMs), generalized additive models (GAM; Hastie and Tibshirani 1987), generalized boosting models (GBMs), multivariate adaptive regression splines (MARS; Friedman 1991), artificial neural networks (ANN; Ripley 1996), random forests (RF; Breiman 2001) and Maximum entropy models (Maxent; Phillips et al. 2006). Binary models for the ESMs-Maxent were tuned by employing the “ENMevaluate” function in the “ENMeval” R-package (Muscarella et al. 2014). Binary models of the remaining modeling techniques were tuned using the “BIOMOD-tuning” function implemented in the “escopat” R-package (see Guisan et al. 2017 for details of tuning parameters; Breiner et al. 2018). We also made an ensemble of predictions of all ESMs (EP-ESMs) weighted by their AUC values. To reduce the effect of sampling bias (Phillips et al. 2009), we removed all but one presence or absence point within each 1 km × 1 km (species range order) and 30 m × 30 m (the remaining orders) pixel (Boria et al. 2014; Goljani Amirkhiz et al. 2018). For species range, population, and individual orders, we defined the maximum number of background points such that each pixel was represented by a single point. Background points were down-weighted to equal presence points (prevalence of 0.5; Ferrier et al. 2002). We applied tenfold split sampling (90% for training and 10% for testing) to calculate Somers’ D (2 × AUC-0.5) to evaluate and weight average small models into ESMs. Small models with Somers’ D values ≤ 0 were considered as random models and were not included in ESMs.
To define extent for population and individual orders, in addition to the weighted average method, we applied a committee averaging method (Araújo and New 2007) to the prediction maps of the best ESMs for the previous order based on model evaluation results. Committee averaging is an unweighted averaging method requiring continuous prediction maps to be reclassified into binary maps. Binary maps developed from each model were then summed and the total divided by the total number of binary maps to obtain a single binary map as the output (Araújo and New 2007). We reclassified the prediction maps of small models by calculating their optimal thresholds maximizing the sum of sensitivity and specificity and meeting a required specificity in the R-package “PresenceAbsence” (Freeman and Moisen 2008). We employed maximum sum of sensitivity and specificity since it serves as a robust method for classification of prediction maps for presence-background and rare species distribution models (Liu et al. 2013, 2016). We used required specificity because our goal in defining extent was the conservative removal of unsuitable areas based on the previous order’s models. Required specificity considers the lowest threshold that met a required specificity, defined as the proportion of observations correctly predicted as absence points (Wilson et al. 2005). We considered 85% as the required specificity (Freeman and Moisen 2008). We next divided the sum of binary prediction maps by the total number of binary prediction maps. Thus, for the best modeling technique for each order based on model evaluation results, we obtained three separate ESMs: ESMs-Weighted average, ESM- Committee averaging (maximum sum of sensitivity and specificity) and ESMs-Committee averaging (required specificity). We then used true absence points from our field surveys, along with presence points, to determine the optimum threshold resulting in the best binary map. To accomplish this, we calculated 12 thresholds based on all available criteria in the R-package “PresenceAbsence” (Freeman and Moisen 2008) and chose the threshold providing the highest scores for kappa and true skill statistic indices, which are robust methods for evaluating binary prediction maps (Segurado and Araújo 2004; Allouche et al. 2006).
For the individual life requirement order, using our absence points, we developed a hierarchical Bayesian logistic regression model using the R-package brms (Bürkner 2017). First, we clipped out those absence points falling in unsuitable areas recognized in previous orders. We next applied a Gaussian mixture model (Banfield and Raftery 1993) corresponding to predictor variable values for occurrence records to discover clusters of occurrence records which share a degree of similarity in at least some niche dimensions. We assumed there are some unobserved patterns within clusters which did not result from our predictor variables. We considered these clusters as random effects in the hierarchical Bayesian logistic regression model. We determined the optimum number of clusters and Gaussian mixture model parameters based on the Bayesian Information Criterion (BIC), which works well in model-based clustering (Dasgupta and Raftery 1998). We used the R-package mclust (Scrucca et al. 2016) to implement the Gaussian mixture model. Rare species have narrow niches and usually occupy a small portion of the available habitat (Lomba et al. 2010). Our preliminary models for various orders also indicated that Virginia’s warbler has a narrow niche. Thus, we considered Student's t distribution with a mean of zero (because we standardized predictor variables by their standard deviations) as a weakly informative prior for the predictor’s coefficients. Using the Widely Applicable Information Criterion (WAIC; Watanabe 2010), we selected the optimum Degrees of Freedom (DF) and Standard Deviation (SD) by comparing models with a range of degrees of freedom (3 to 7 with an increment of 0.5) and a narrow range of standard deviations (1 to 4 with an increment of 0.5) as recommended for the weakly informative priors (Ghosh et al. 2018). We then applied an indicator variable selection approach to select the best subset of predictors by removing predictor variables with a posterior mean close to zero in a stepwise manner (O’Hara and Sillanpaa 2009; Hooten and Hobbs 2015). We compared models with and without predictors with a posterior mean close to zero using tenfold cross-validation and estimating expected log pointwise predictive density (elpd; Vehtari et al. 2017). To run hierarchical Bayesian logistic regression models, we considered four chains each with 12,000 iterations, with 6000 iterations to prime the models. To determine the importance of each variable in the model, we applied a projection predictive variable selection approach using the R-package “projpred” (Piironen and Vehtari 2017).
Models evaluation, variable importance, and response curves
To evaluate the accuracy of predictions of ESMs, we used the block method in the R-package “ENMeval” (Muscarella et al. 2014) to split occurrence records into four spatially independent bins and used half the data for training models and half for testing them independently. Using partitioned data, we calculated threshold-independent and threshold-dependent model evaluation metrics using R-packages “PresenceAbsence” (Freeman and Moisen 2008) and “escopat” (Breiner et al. 2018). We used averaged AUC (AUC.mean) as a threshold-independent method to evaluate overall model performance (Peterson et al. 2008). We also calculated the AUC difference (AUC.diff). AUC.diff represents the difference between AUCs calculated on training data and testing data and allows evaluation of model overfitting (Warren and Seifert 2011). We also used the continuous Boyce index (Boyce.mean) as a background points-independent index (Hirzel et al. 2006). For threshold-dependent methods, we used the mean true skill statistic across blocks (Allouche et al. 2006). We found the optimum threshold for mean true skill statistic using maximum sum of sensitivity and specificity.
To obtain the relative importance of variables of the best ESMs, we calculated the mean of permutation importance metrics through all binary models using the “variables_importance” function in the R-package Biomod2, version 3.3-7.1 (Thuiller et al. 2009). We used Mann–Whitney U tests to compare importance of variables used in their corresponding small models. Variables were grouped if there was no significant difference among their variable importance scores and then ranked based on the mean variable importance. To produce response curves for each covariate, we plotted the mean of predicted probability of binary models including the covariate (when the other covariate in the binary model was kept at its median value) against the corresponding values of the covariate. We used the “response” function in the R-package dismo (Hijmans et al. 2017) to create response curves.
Results
Species range order
ESMs-Maxent had better evaluation metrics than other models and it was more accurate based on visual inspection (see Appendix S5a for details of evaluation metrics and tuned parameters). An ESMs-Maxent with committee averaging, in which small model prediction maps were classified using maximum sum of sensitivity and specificity resulted in the most accurate binary map. This binary map was used to define the extent of the population order (see Appendix S5b for details). According to this map, suitable in-range habitats for the species range order are located in the southern, southwestern and western Black Hills, extending into Wyoming. Additional potentially suitable habitats may also occur in the northwestern Black Hills and northwest of the study area (Fig. 2). The maximum temperature of June and July (BIO23) and mean temperature of the wettest quarter (BIO8) had the highest contributions to the Virginia’s warbler’s niche at this order. However, based on a Mann–Whitney U test these two variables did not show significantly higher importance than annual precipitation (BIO12), and the maximum temperature and precipitation of the warmest month (BIO5 and BIO18; P > 0.05). Thus, these five variables had the greatest contributions to defining the niche of Virginia’s warblers (Appendix 5c).
Prediction maps of a species range order, b population order and c Individual order in Black Hills region. d Provides background extents for ensemble of small models of a to c. These maps depict the hierarchical habitat selection by Virginia’s warbler in the Black Hills, U.S.A. Maps depict predictions of ensembles of small models made by modeling techniques that performed better (a maximum entropy, b generalized boosting models, c artificial neural networks) than other modeling techniques (generalized linear models, generalized additive models, multivariate adaptive regression splines, and random forests) at each hierarchical order of habitat selection. The suitable habitats recognized at each order were used as the background extent for the subsequent order
Response curves for the top five most important variables showed that the Virginia’s warbler has a narrow niche regarding climate variables. All response curves had quadratic shapes around narrow ranges of variables at the species range order background extent. These curves showed that Virginia’s warblers prefer areas with a moderate maximum temperature in June and July, in the wettest quarter of the year and in the warmest month of the year. This species shows higher occupancy in areas with lower annual precipitation and moderate precipitation in the warmest quarter of the year (Appendix 5d).
Population order
Regarding the selected window radius according to AUC values, the majority of univariate models on soil groups showed their best performance at a 1-km window radius. Conversely, most of the landcover covariates had their best performance at a window radius of 90 m. Topographic and vegetation type covariates showed their best performance at various windows radii (Appendix S3). At the population order, ESMs-GBM performed better than other models (Appendix S6a). Using a committee averaging approach in which small models were classified using maximum sum of sensitivity and specificity resulted in the most accurate binary map, which was used to define the background extend of the individual order (see Appendix S6b for details). Based on a Mann–Whitney U test, variables with significant contributions to the Virginia’s warbler’s niche (P < 0.05) were height of shrubs (VH-Shrub), Terrain Ruggedness Index (TRI), Heat Load Index (HLI), frequency of Haplustlfs and Argiustolls soil groups, Northwestern Great Plains mixed grass prairie (VT3141), Western Great Plains floodplain forest and woodland (VT3162), Western Great Plains sand prairie grassland (VT3148), and northern Rocky Mountain montane-foothill deciduous shrubland (VT3106; Appendix S6c). The prediction map showed that the most suitable habitats for Virginia’s warbler at this order are located in southern areas of the species range order with additional potentially suitable habitats in the western and northern regions (Fig. 2).
Individual order
At the individual order, ESMs-ANN had the best performance among all models based on all metrics (Appendix S7a) and was used to explore the Virginia’s warbler’s niche. The prediction map revealed that within the suitable areas for the population order, most suitable habitats at the individual order level were located in southern and western regions of the Black Hills, but some small suitable habitats occurred in northwestern areas (Fig. 2). The most important covariate, based on a Mann–Whitney U test, at the individual order was terrain ruggedness index and the second most important was Integrated Moisture Index (IMI, P < 0.05). Other important covariates were frequency of Calciustolls and Ustorthent soil groups, vegetation cover variation (VCVar), frequency of Inter-Mountain Basins Big Sagebrush Steppe (VT3125), Northwestern Great Plains mixed grass prairie (VT3141), ground canopy cover (GCC), and Northwestern Great Plains-Black Hills Ponderosa Pine Woodland and Savanna (VT3179), height of shrubs, and height of trees (VH-Tree). These latter covariates did not differ significantly from each other (P > 0.05) so can all be considered as the third most important variables (Appendix 7b).
Individual life requirements order
The Gaussian mixture model showed two clusters of occurrence records which were used as random effects. Based on WAIC, the best prior for the coefficients was a Student's t distribution with DF and SD of 3 and 2.25, respectively. The optimum DF and SD for the intercept was 4 and 1, respectively. The tenfold cross-validation showed that a model including the best subset of covariates with a varying intercept had the best performance (Appendix S8a). The best model included 15 covariates (six with positive, eight with negative and one with a quadratic relationship with the probability of occurrence, Appendix S8b). The increase in expected log pointwise predictive density when adding each predictor variable to the model showed that terrain ruggedness index is the most important variable in this model, while Inter-Mountain Basin Big Sagebrush Steppe (VT3125) and Northern Rocky Mountain Montane-Foothill Deciduous Shrubland (VT3106) had the least contributions to the model (Table 1).The marginal effect plots showed that the amount of uncertainty in prediction in areas with lower habitat suitability was generally higher (Fig. 3).
Response curves of top ranked covariates of population, individual and individual life requirement orders. The numbers at the top right of each figure refer to the importance score (most important = 1, second most important = 2, etc.) for each covariate at the corresponding habitat selection order. This figure illustrates that probability of occurrence varies in relation to changes in predictor variables and how the importance of variables changes at different habitat selection orders
Top-ranked covariates and niche characteristics at population, individual and individual life requirement orders
Terrain ruggedness index (TRI) is the most important variable for all three orders (Fig. 3). The quadratic shape of the terrain ruggedness index response curve at the population order showed that available habitats are limited to low to medium values of terrain ruggedness index. However, at the next finer scale levels (individual and and individual life requirment orders), areas with the highest terrain ruggedness index are preferred, implying that other factors limit selection of areas with the most suitable terrain ruggedness index at the population order. Eight other covariates occurred at the same level of importance as terrain ruggedness index at the population order (Fig. 3), so these variables may interact to limit the use of optimum levels of other covariates (e.g., terrain ruggedness index and height of shrubs). Virginia’s warblers prefer areas with low to medium shrub height at the population order. However, the importance of shrub height decreases at the individual order and becomes unimportant at the individual life requirement order, allowing response to optimum levels of other variables. Such a decrease in importance at lower levels is also evident for other top-ranked covariates at the population order. For example, heat load index (HLI), shows a bimodal pattern at population and individual orders with both colder and warmer slopes being preferred. However, at the individual life requirement order, only moderate to warmer areas (higher heat load index) are preferred. Virginia’s warblers avoid areas with Haplustalf and Argisutull soils, Western Great Plains Floodplain Forest and Woodland (VT3162) and Western Great Plains Sand Prairie Grassland (VT3148) at species range and population orders, but these variables play no role at the individual life requirement order. Haplustalf and Argiustoll soil groups support grasslands and our results showed that these habitats are unsuitable breeding habitat for Virginia’s warbler, so they are avoided at the broader orders. Low coverages of Northwestern Great Plains Mixed Grass Prairie (VT3141) and Northern Rocky Mountain Montane-Foothill Deciduous Shrubland (VT3106) are preferred at all three orders, implying that the presence of other vegetation types is also required to select an area as breeding habitat. The integrated moisture index (IMI) is not a top-ranked variable at the population order, but its influence becomes greater at finer scales. Low levels of integrated moisture index are favored at all three orders below species range order. The Ustifluvents soil group is less important at broader orders but becomes a top-ranked variable at the individual life requirement order. Distance to ridge and areas underlain by Hapludolls soils were significant predictors only at the individual life requirement order showing that they are considered by Virginia’s warbler only at finer scales (see Fig. 3 for top-ranked covariates and appendix S9 for the rest of the covariates).
Discussion
Virginia’s Warbler niche requirements
At broader habitat selection orders below the species range order (population and individual order), Virginia’s warbler prefers areas with intermediate vegetation cover height and variation, low humidity levels, and intermediate slopes and avoids grassland-dominated habitats (probably due to lack of sufficient cover components). Previous studies also reported that this species tends to occupy xeric habitats (Olson and Martin 1999). With less importance, Virginia’s warblers also prefer habitats with a mix of tree and shrub cover at lower elevations at broader habitat selection scales. These habitats have intermediate NDVI values and shallow to moderately deep and rocky soils which are characteristics of shrub habitat types with mountain mahogany, the favored vegetation type for Virginia’s warbler in the Black Hills (Swanson et al. 2000, 2016). At the individual life requirement order, Virginia’s warblers occupy steep areas close to ridges with moderate to warm temperatures, consistent with the idea that Virginia’s warblers use ridges as territory boundaries (Swanson et al. 2016) and prefer habitats that are warmer than surrounding areas (Olson and Martin 1999). At this order, Virginia’s warbler occupy regions with Hapludoll and Ustifluvent soils. Typical vegetation cover of Hapludoll soils is tall grass with a tree overstory (Soil Survey Staff 2015) and this vegetation could serve as a cover component when Virginia’s warblers select their nest sites, if mountain mahogany is also present. Ustifluvent soil groups are favored by Virginia’s warbler at the population order and occur along streams (Soil Survey Staff 2015), which usually occur in areas with high terrain ruggedness, a feature preferred by Virginia’s warbler. Thus, at population and individual orders, an intermediate frequency of the Ustifluvent soil group is preferred, since higher frequencies occur close to streams in moister habitats than those preferred by Virginia’s warbler. However, at the individual life requirement order the highest frequencies of Ustifluvent soil groups are favorable since unsuitable frequencies have already been disregarded at higher habitat selection orders.
Ensemble of small models within hierarchical habitat selection framework
Our study suggests that choosing extents and scales-of-effect based on habitat selection concepts (Johnson 1980; Mayor et al. 2009) could improve understating of rare species niche requirements and increase the accuracy of predictions. Our results are consistent with the hypothesis that covariates can have different roles, interpretations, and influences on the occurrence of Virginia’s warbler at different scales. Many studies show that disregarding hierarchy theory (O’Neill et al. 1986) in habitat selection and suitability studies may lead to misinterpretation of the niche (e.g., Syfert et al. 2013; Vergara et al. 2016). For instance, although terrain ruggedness is one of the most important variables at three habitat selection orders, its relative influence on Virginia’s warbler habitat preference varies at different scales. Some covariates are only influential at broader scales (e.g., shrub height), whereas others are important only at the finest scale (e.g., distance to the ridge). At broader scales (species range and population orders) Virginia’s warblers make general decisions, as many covariates show high importance in the corresponding models. However, at finer scales certain covariates become more important than others (terrain ruggedness index and integrated moisture index at population order; terrain ruggedness index, Hapludolls and Ustifluvents soil groups at individual order). Response curves showed that the Virginia’s warbler habitat selection pattern at larger scales is to avoid unsuitable habitats (e.g., Haplustalfs and Argiustolls), whereas less important covariates at larger scales become important at the finest scale (distance to the ridge and Hapludolls). Such patterns are also reported by studies of habitat selection for other birds (MacFaden and Capen 2002; Martínez et al. 2003). The importance of multiscale modeling could be magnified in rare species with narrow niches as a result of specific ecological adaptations and life history strategies (Guisan and Thuiller 2005). Habitat selection could, therefore, provide a theoretical framework to specify appropriate extents and scales-of-effect when ESMs is employed in rare species distribution studies. We acknowledge that these results are based on correlations but identified associations can lead to future studies investigating mechanisms that drive multi-scale habitat selection and influence fitness of nesting Virginia’s warblers.
Our modeling method is complementary to and builds upon those developed by Lobma et al. (2010) and Breiner et al. (2015, 2018). In our study, tuned ESMs-Maxent, ESMs-GBM and ESMs-ANN performed best at the three broadest habitat selection orders, which is consistent with the results of Breiner et al. (2015). However, our study showed that the performance of different modeling techniques could vary among scales. While Maxent had the best performance at the species range order, it did not perform as well as GBM at the population order. Whereas GBM performed best at the population order, it had a weak performance at the individual order. ANN was the best technique at the individual order, but not at other orders.
Conclusions about the relative merits of using presence-absence based methods vs. presence-background methods (Brotons et al. 2004) generally depend on the sample size and/or statistical methods (Elith et al. 2006). A spatial hierarchical framework can take advantage of strengths of both approaches to test habitat selection hypotheses for rare species. The projections of our preliminary ensembles of small models (using only presence and absence points, and not including pseudo-absence points) into our broadest scale (species range order) resulted in weak and unrealistic predictions (Appendix S10). For Virginia’s warbler, our absence points did not represent the entire available habitat. Thus, using pseudo-absence (background) points was a better choice to test our hypotheses at the broadest three habitat selection orders. Response curves and prediction maps of the best ESMs supported our expectation that Virginia’s Warbler occupies a narrow niche with limited regional potential distribution at the species range order. Incorporating pseudo-absence points identified some potential suitable habitats separated from the known range at the population order. In addition, the entire suitable population order extent does not encompass all habitat requirements at the individual order. However, at the individual life requirement order, using absence points helped delineate which covariates served as specific habitat requirements for the establishment of a breeding territory by Virginia’s warblers (e.g., vicinity to ridges, warmer slopes and higher moisture). Although other modeling techniques (e.g., logistic regression) could be applied to presence-absence data at this order, applying a hierarchical Bayesian approach facilitated variable selection by shrinking coefficients of unimportant covariates to zero, resulting in a suite of the most important covariates in the model. This approach also quantified that uncertainty in coefficient estimations is much higher for presence points than for absence points, suggesting that more data describing unsuitable habitats at the individual life requirements order is required to better define the Virginia’s warbler niche at finer scales.
Ensemble of small models through committee averaging
Previous ESMs studies used the weighted averaging method to create an ensemble of small models (e.g. Lomba et al. 2010; Breiner et al. 2018). Our study showed that the committee averaging method may lead to more accurate results, especially when creating binary prediction maps is an objective. In this study, potential suitable habitats at each order served as the background extent for the next order. To define background extents, we reclassified the prediction map for each order into a binary map by selecting the best threshold criterion. In this way we could select background extents based on the objectives of our study, leading to biologically meaningful response curves (Merow et al. 2013).
Management implication and future studies guideline
Applying an ESMs approach within a hierarchical framework led to detailed information about the Virginia’s warbler’s niche, limiting factors at each habitat selection order, and potential distribution, which could be helpful for multiscale management and future research. Limiting factors have an important role in habitat selection and, consequently, could shape management actions (Rettie and Messier 2000). The species range order model showed that Virginia’s warbler has a narrow niche regarding bioclimatic variables, which restricts its distribution to some parts of the southern, western and northwestern Black Hills. Climate change could potentially affect future Virginia’s warbler distribution in the Black Hills. Indeed, some models predict an increase in suitable habitats for Virginia’s warbler in the Black Hills as a function of climate change, which may buffer range-wide population declines (van Riper III et al. 2014), especially if Virginia’s warblers continue to expand their range to occupy these new habitats (Bubac and Spellman 2016). Future studies can investigate the impact of climate change on Virginia’s warbler distribution in the Black Hills by comparing its Black Hills niche with the climatic niches of southern populations of this species.
Virginia’s warblers mostly avoid unsuitable habitats at the population order characterized by highly rugged areas dominated by grasslands and shrublands with little tree cover. As a consequence, wildfire events, extensive tree harvesting, or drought could decrease the suitability of current habitats for Virginia’s warblers. The population order model indicated that some other habitats outside of the current known Virginia’s warbler distribution could potentially serve as suitable habitats in the Black Hills. More field surveys and source-sink population studies could provide additional evidence about presence of Virginia’s warblers in these potential habitats. At the individual order, avoidance of some habitat features such as high moisture and dense grasslands compel Virginia’s warbler to search for suitable habitats lacking these features. Such suitable habitats are characterized by rugged terrain, higher vegetation cover and variation (including moderate densities of trees and shrubs), and low grassland density. Maintaining appropriate vegetation cover variation in suitable habitats should be considered in management plans. At the individual life requirement order, areas which are close to ridges and have moderate vegetation cover variation, including trees, shrubs, and tall grasses, have the highest suitability. Thus, at this scale, landcover configuration is important and should be considered, especially in case of disturbances. At the finest scale, our model showed that more data are needed to better delineate habitat features associated with unsuitable breeding territories (e.g., biotic interactions (Swanson et al. 2016) and vegetation composition and configuration).
References
Allouche O, Tsoar A, Kadmon R (2006) Assessing the accuracy of species distribution models: prevalence, kappa and the true skill statistic (TSS). J Appl Ecol 43:1223–1232
Araújo MB, New M (2007) Ensemble forecasting of species distributions. Trends Ecol Evol 22:42–47
Banfield JD, Raftery AE (1993) Model-based Gaussian and non-Gaussian clustering. Biometrics 49:803–821
Bellamy C, Scott C, Altringham J (2013) Multiscale, presence-only habitat suitability models: fine-resolution maps for eight bat species. J Appl Ecol 50:892–901
Boria RA, Olson LE, Goodman SM, Anderson RP (2014) Spatial filtering to reduce sampling bias can improve the performance of ecological niche models. Ecol Modell 275:73–77
Boyce MS, Mao JS, Merrill EH, Fortin D, Turner MG, Fryxell J, Turchin P (2003) Scale and heterogeneity in habitat selection by elk in Yellowstone National Park. Écoscience 10:421–431
Breiman L (2001) Random forests. Mach Learn 45:5–32
Breiner FT, Nobis MP, Bergamini A, Guisan A (2018) Optimizing ensembles of small models for predicting the distribution of species with few occurrences. Methods Ecol Evol 9:802–808
Brotons L, Thuiller W, Araújo MB, Hirzel AH (2004) Presence-absence versus presence-only modelling methods for predicting bird habitat suitability. Ecography (Cop) 27:437–448
Bubac CM, Spellman GM (2016) How connectivity shapes genetic structure during range expansion: Insights from the Virginia’s Warbler. Auk 133:213–230
Bürkner P-C (2017) brms: an R package for Bayesian multilevel models using stan. J Stat Softw 1(1):1–8
Dasgupta A, Raftery AE (1998) Detecting features in spatial point processes with clutter via model-based clustering. J Am Stat Assoc 93:294–302
Della Rocca F, Bogliani G, Breiner FT, Milanesi P (2019) Identifying hotspots for rare species under climate change scenarios: improving saproxylic beetle conservation in Italy. Biodivers Conserv 28:433–449
Devictor V, Julliard R, Jiguet F (2008) Distribution of specialist and generalist species along spatial gradients of habitat disturbance and fragmentation. Oikos 117:507–514
Di Febbraro M, Menchetti M, Russo D et, Ancillotto L, Aloise G, Roscioni F, Preatoni DG, Loy A, Martinoli A, Bertolino S, Mori E (2019) Integrating climate and land-use change scenarios in modelling the future spread of invasive squirrels in Italy. Divers Distrib 25:644–659
Dormann CF, Elith J, Bacher S, Buchmann C, Carl G, Carré G, García Marquéz JR, Gruber B, Lafourcade B, Leitão PJ, Münkemüller T, McClean C, Osborne PE, Reineking B, Schröder B, Skidmore AK, Zurell D, Lautenbach S (2013) Collinearity: a review of methods to deal with it and a simulation study evaluating their performance. Ecography (Cop) 36:27–46. https://doi.org/10.1111/j.1600-0587.2012.07348.x
Elith J, Graham CH, Anderson RP, Dudík, Ferrier SDM, Guisan A, Hijmans RJ, Huettmann F, Leathwick JR, Lehmann A, Li J, Lohmann LG, Loiselle BA, Manion G, Moritz C, Nakamura M, Nakazawa Y, Overton McCMJ, Townsend Peterson A, Phillips JS, Richardson K, Scachetti‐Pereira R, Schapire RE, Soberón J, Williams S, Wisz MS, Zimmermann NE (2006) Novel methods improve prediction of species’ distributions from occurrence data. Ecography (Cop) 29:129–151
ESRI (2017) ArcGIS desktop: release 107. Environmental Systems Research Institute, Redlands
Ferrier S, Drielsma M, Manion G, Watson G (2002) Extended statistical approaches to modelling spatial pattern in biodiversity in northeast New South Wales II Community-level modelling. Biodivers Conserv 11:2309–2338
Fick SE, Hijmans RJ (2017) WorldClim 2: new 1-km spatial resolution climate surfaces for global land areas. Int J Climatol 37:4302–4315
Fielding AH, Bell JF (1997) A review of methods for the assessment of prediction errors in conservation presence/absence models. Environ Conserv 24:38–49
Fischer JM (1978) A natural history study of the Virginia’s Warbler. Northern Arizona University, Arizona
Fletcher R, Fortin M-J (2018) Space use and resource selection. Spatial ecology and conservation modeling: applications with R. Springer International Publishing, Cham, pp 271–320
Freeman EA, Moisen G (2008) Presence absence: an r package for presence absence analysis. J Stat Softw 1:11
Friedman JH (1991) Multivariate adaptive regression splines. Ann Stat 19:1–67
Gaston K, Blackburn T (2008) Pattern and process in macroecology. Blackwell Publishing Inc, Maden
Ghosh J, Li Y, Mitra R (2018) On the use of cauchy prior distributions for bayesian logistic regression. Bayesian Anal 13:359–383
Goljani Amirkhiz R, Frey JK, Cain JW, Breck SW, Bergman DL (2018) Predicting spatial factors associated with cattle depredations by the Mexican wolf (Canis lupus baileyi) with recommendations for depredation risk modeling. Biol Conserv. https://doi.org/10.1016/j.biocon.2018.06.013
Guillera-Arroita G, Lahoz-Monfort JJ, Elith J Gordon A, Kujala H, Lentini PE, McCarthy MA, Tingley R, Wintle BA (2015) Is my species distribution model fit for purpose? Matching data and models to applications. Glob Ecol Biogeogr 24:276–292
Guisan A, Thuiller W (2005) Predicting species distribution: offering more than simple habitat models. Ecol Lett 8:993–1009
Guisan A, Broennimann O, Engler R, Vust M, Yoccoz NG, Lehmann A, Zimmermann NE (2006) Using niche-based models to improve the sampling of rare species. Conserv Biol 20:501–511
Guisan A, Tingley R, Baumgartner JB et al (2013) Predicting species distributions for conservation decisions. Ecol Lett 16:1424–1435
Guisan A, Thuiller W, Zimmermann NE (2017) Habitat suitability and distribution models: with applications in R. Cambridge University Press, Cambridge
Habibzadeh N, Ludwig T (2019) Ensemble of small models for estimating potential abundance of Caucasian grouse (Lyrurus mlokosiewiczi) in Iran. Ornis Fenn 96:77–89
Hastie T, Tibshirani R (1987) Generalized additive models: some applications. J Am Stat Assoc 82:371–386
Hernandez PA, Graham CH, Master LL, Albert DL (2006) The effect of sample size and species characteristics on performance of different species distribution modeling methods. Ecography (Cop) 29:773–785
Hijmans RJ, Phillips S, Leathwick J, Elith J (2017) Species distribution modeling. R package version 1.1–4
Hirzel AH, Le Lay G, Helfer V, Randin Ch, Guisan A (2006) Evaluating the ability of habitat suitability models to predict species presences. Ecol Modell 199:142–152
Hooten MB, Hobbs NT (2015) A guide to Bayesian model selection for ecologists. Ecol Monogr 85:3–28
Jackson HB, Fahrig L (2015) Are ecologists conducting research at the optimal scale? Glob Ecol Biogeogr 24:52–63
Jedlikowski J, Chibowski P, Karasek T, Brambilla M (2016) Multi-scale habitat selection in highly territorial bird species: exploring the contribution of nest, territory and landscape levels to site choice in breeding rallids (Aves: Rallidae). Acta Oecologica 73:10–20
Johnson DH (1980) The comparison of usage and availability measurements for evaluating resource preference. Ecology 61:65–71
Liu C, White M, Newell G (2013) Selecting thresholds for the prediction of species occurrence with presence-only data. J Biogeogr 40:778–789
Liu C, Newell G, White M (2016) On the selection of thresholds for predicting species occurrence with presence-only data. Ecol Evol 6:337–348
Lomba A, Pellissier L, Randin C, Vicente J, Moreira F, Honrado J, Guisan A (2010) Overcoming the rare species modelling paradox: a novel hierarchical framework applied to an Iberian endemic plant. Biol Conserv 143:2647–2657
MacFaden SW, Capen DE (2002) Avian habitat relationships at multiple scales in a New England forest. For Sci 48:243–253
Manly BF, McDonald L, Thomas DL, McDonald TL, Erickson WP (2007) Resource selection by animals: statistical design and analysis for field studies. Springer, Amsterdam
Martínez JA, Serrano D, Zuberogoitia I (2003) Predictive models of habitat preferences for the Eurasian eagle owl Bubo bubo: a multiscale approach. Ecography (Cop) 26:21–28
Mayor SJ, Schneider DC, Schaefer JA, Mahoney SP (2009) Habitat selection at multiple scales. Écoscience 16:238–247
McCune JL (2016) Species distribution models predict rare species occurrences despite significant effects of landscape context. J Appl Ecol 53:1871–1879
McGarigal K, Wan HY, Zeller KA, Timm BC, Cushman SA (2016) Multi-scale habitat selection modeling: a review and outlook. Landsc Ecol 31:1161–1175
Merow C, Smith MJ, Silander JA (2013) A practical guide to MaxEnt for modeling species’ distributions: what it does, and why inputs and settings matter. Ecography (Cop) 36:1058–1069
Meyer CB, Thuiller W (2006) Accuracy of resource selection functions across spatial scales. Divers Distrib 12:288–297
Muscarella R, Galante PJ, Soley-Guardia M, Boria RA, Kass JM, Uriarte M, Anderson RP (2014) ENMeval: an R package for conducting spatially independent evaluations and estimating optimal model complexity for Maxent ecological niche models. Methods Ecol Evol 5:1198–1205
O’Hara RB, Sillanpaa MJ (2009) A review of Bayesian variable selection methods: what, how and which. Bayesian Anal 4:85–117
O’Neill RV, Deangelis DL, Waide JB, Allen TFH (1986) A hierarchical concept of ecosystems. Princeton University Press, Princeton
Olson CR, Martin TE (1999) Virginia’s Warbler (Oreothlypis virginiae). In: The birds of North America. Cornell Lab of Ornithology, New York
Orr HK (1975) Watershed management in the Black Hills: the status of our knowledge. USDA Forest Service, Fort Collins CO
Pacifici K, Dorazio RM, Conroy MJ (2012) A two-phase sampling design for increasing detections of rare species in occupancy surveys. Methods Ecol Evol 3:721–730
Peterson AT, Papeş M, Soberón J (2008) Rethinking receiver operating characteristic analysis applications in ecological niche modeling. Ecol Modell 213:63–72
Phillips SJ, Anderson RP, Schapire RE (2006) Maximum entropy modeling of species geographic distributions. Ecol Modell 190:231–259
Phillips SJ, Dudik M, Elith J, Graham CH, Lehmann A, Leathwick J, Ferrier S (2009) Sample selection bias and presence-only distribution models: implications for background and pseudo-absence data. Ecol Appl 19:181–197
Piironen J, Vehtari A (2017) Comparison of Bayesian predictive methods for model selection. Stat Comput 27:711–735
Razgour O, Hanmer J, Jones G (2011) Using multi-scale modelling to predict habitat suitability for species of conservation concern: the grey long-eared bat as a case study. Biol Conserv 144:2922–2930
Rettie WJ, Messier F (2000) Hierarchical habitat selection by woodland caribou: its relationship to limiting factors. Ecography (Cop) 23:466–478
Ripley BD (1996) Pattern recognition and neural networks. Cambridge University Press, Cambridge
Root T (1988) Environmental factors associated with avian distributional boundaries. J Biogeogr 15:489–505
Scherrer D, Christe P, Guisan A (2019) Modelling bat distributions and diversity in a mountain landscape using focal predictors in ensemble of small models. Divers Distrib 25:770–782
Scrucca L, Fop M, Murphy TB, Raftery AE (2016) mclust 5: clustering, classification and density estimation using gaussian finite mixture models. R J 8:289–317
Segurado P, Araújo MB (2004) An evaluation of methods for modelling species distributions. J Biogeogr 31:1555–1568
Suárez-Seoane S, Osborne PE, Alonso JC (2002) Large-scale habitat selection by agricultural steppe birds in Spain: identifying species-habitat responses using generalized additive models. J Appl Ecol 39:755–771
Swanson DL, Palmer JS, Liknes ET, Dean KL (2000) A breeding population of Virginia’s Warblers in the southwestern Black Hills of South Dakota. Southwest Nat 45:39–44
Swanson DL, Dixon MD, Palmer JS (2016) A re-assessment of the distribution of Virginia’s warbler in the Black Hills of South Dakota
Syfert MM, Smith MJ, Coomes DA (2013) The effects of sampling bias and model complexity on the predictive performance of MaxEnt species distribution models. PLoS ONE 8:e55158
Thuiller W, Lafourcade B, Engler R, Araújo MB (2009) BIOMOD – a platform for ensemble forecasting of species distributions. Ecography (Cop) 32:369–373
Van Riper IIIC, Hatten JR, Giermakowski JT, Mattson D, Holmes JA, Johnson MJ, Nowak EM, Ironside K, Peters M, Heinrich P, Cole KL, Truettner C, Schwalbe CR (2014) Projecting climate effects on birds and reptiles of the Southwestern United States. Reston, VA
VanDerWal J, Shoo LP, Johnson CN, Williams SE (2009) Abundance and the environmental niche: environmental suitability estimated from niche models predicts the upper limit of local abundance. Am Nat 174:282–291
Vehtari A, Gelman A, Gabry J (2017) Practical Bayesian model evaluation using leave-one-out cross-validation and WAIC. Stat Comput 27:1413–1432
Vergara M, Cushman SA, Urra F, Ruiz-González A (2016) Shaken but not stirred: multiscale habitat suitability modeling of sympatric marten species (Martes martes and Martes foina) in the northern Iberian Peninsula. Landsc Ecol 31:1241–1260
Warren DL, Seifert SN (2011) Ecological niche modeling in Maxent: the importance of model complexity and the performance of model selection criteria. Ecol Soc Am 21:335–342
Watanabe S (2010) Asymptotic Equivalence of Bayes Cross Validation and Widely Applicable Information Criterion in Singular Learning Theory. http://arxiv.org/abs/1004.2316
Weihs C, Ligges U, Luebke K, Raabe N (2005) klaR analyzing German business cycles. In: Baier D, Decker R, Schmidt-Thieme L (eds) Data analysis and decision support. Springer-Verlag, Berlin, pp 335–343
Wilson KA, Westphal MI, Possingham HP, Elith J (2005) Sensitivity of conservation planning to different approaches to using predicted species distribution data. Biol Conserv 122:99–112
Zimmerman GS, Gutiérrez RJ, Thogmartin WE, Banerjee S (2009) Multiscale habitat selection by ruffed grouse at low population densities. Condor 111:294–304
Acknowledgements
We would like to thank J. Wesner for providing feedback regarding model construction. This work was funded by NSF OIA-1632810 and a wildlife diversity Grant (UP1500121) from the South Dakota Department of Game, Fish and Parks.
Author information
Authors and Affiliations
Corresponding author
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Amirkhiz, R.G., Dixon, M.D., Palmer, J.S. et al. Investigating niches and distribution of a rare species in a hierarchical framework: Virginia’s Warbler (Leiothlypis virginiae) at its northeastern range limit. Landscape Ecol 36, 1039–1054 (2021). https://doi.org/10.1007/s10980-021-01217-7
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10980-021-01217-7