Abstract
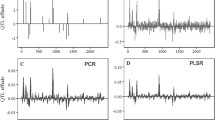
There is a dearth of studies on the genes engaged in citrus taste. Unraveling the major genes involved in pathways related to the taste of citrus (sweet or acidic) is highly important for developing new genotypes with favorable taste. Pivotal genes linked to citrus taste can be extracted through mining a large number of expression data. To attain this objective, 10 different attribute weighting algorithms (AWAs) were applied on the expression data from three lemon (Citrus limon) genotypes differing in terms of fruit acidity. As a result, a total of 170 probe sets were identified by more than eight AWAs as the most discriminative probe sets. Subsequently, principal component analysis and hierarchical clustering heatmaps were implemented for validation of the 170 top-ranked probe sets. Noticeably, the identified top 170 probe sets significantly contributed to accurate discrimination between sweet and acidic lemon samples, which indicate the significance and accuracy of prediction of probe sets. According to the results, some genes like citrate synthase, malate dehydrogenase, proton-pumping ATPase, and flavanone 3-hydroxylase had distinct roles in differentiation of the studied genotypes and acidity. Among all of the genes, malate dehydrogenase was the most informative. To the best of our knowledge, this is the first report on identifying the most important genes contributing to lemon taste using supervised and unsupervised data learning methods.


Similar content being viewed by others
Data availability
All data has been provided with the manuscript. If any additional information is required, please contact the corresponding author.
References
Aprile A, Federici C, Close TJ, De Bellis L, Cattivelli L, Roose ML (2011) Expression of the H+-ATPase AHA10 proton pump is associated with citric acid accumulation in lemon juice sac cells. Funct Integr Genomics 11:551–563. https://doi.org/10.1007/s10142-011-0226-3
Bai Y-X, Hussain SB, Wei X, Shi CY, Liu D-H, Liu Y-Z (2020) Identification and transcript analysis of CsAPD2 reveal its potential role in citric acid accumulation in citrus fruits. Sci Hortic. https://doi.org/10.1016/j.scienta.2020.109607
Baldwin EA (1993) Citrus fruit. In: Seymour GB, Taylor JE, Tucker GA (eds) Biochemistry of fruit ripening. Chapman and Hall, New York, pp 107–149
Clare A, Karwath A, Ougham H, King R (2006) Functional bioinformatics for Arabidopsis thaliana. Bioinformatics 22:1130–1136. https://doi.org/10.1093/bioinformatics/btl051
Du L, Song J, Forney C, Palmer LC, Fillmore S, Zhang Z (2016) Proteome changes in banana fruit peel tissue in response to ethylene and high-temperature treatments. Hortic Res 3:1–12. https://doi.org/10.1038/hortres.2016.12
Etienne A, Génard M, Lobit P, Mbeguié-A-Mbéguié D, Bugaud C (2013) What controls fleshy fruit acidity? A review of malate and citrate accumulation in fruit cells. J Exp Bot 64:1451–1469. https://doi.org/10.1093/jxb/ert035
Forkmann G, Heller W, Grisebach H (1980) Anthocyanin biosynthesis in flowers of Matthiola incana flavanone 3-and flavonoid 3′-hydroxylases. Zeitschrift für Naturforschung C 35:691–695. https://doi.org/10.1515/znc-1980-9-1004
Givan CV (2007) Evolving concepts in plant glycolysis: two centuries of progress. Biol Rev 74:277–309. https://doi.org/10.1111/j.1469-185X.1999.tb00188.x
Irizarry RA, Hobbs B, Collin F, Beazer-Barclay YD, Antonellis KJ, Scherf U, Speed TP (2003) Exploration, normalization, and summaries of high density oligonucleotide array probe level data. Biostatistics 4:249–264. https://doi.org/10.1093/biostatistics/4.2.249
Lama K, Peer R, Shlizerman L, Meir S, Doron-Faigenboim A, Sadka A, Aharoni A, Flaishman MA (2020) Tissue-specific organic acid metabolism in reproductive and non-reproductive parts of the fig fruit is partially induced by pollination. Physiol Plant 168:133–147. https://doi.org/10.1111/ppl.12941
Lan H, Carson R, Provart NJ, Bonner A (2007) Combining classifiers to predict gene function in Arabidopsis thaliana using large-scale gene expression measurements. BMC Bioinform 8:358. https://doi.org/10.1186/1471-2105-8-358
Li SJ, Liu XJ, Xie XL, Sun Cd, Grierson D, Xr Y, Ks C (2015) CrMYB73, a PH-like gene, contributes to citric acid accumulation in citrus fruit. Sci Hortic 197:212–217. https://doi.org/10.1016/j.scienta.2015.09.037
Li LJ, Tan WS, Li W, Zhu YB, Cheng YS, Ni H (2019) Citrus taste modification potentials by genetic engineering. Int J Mol Sci 20:6194. https://doi.org/10.3390/ijms20246194
Lorenzon R, Mariotti-Ferrandiz E, Aheng C, Ribet C, Toumi F, Pitoiset F, Chaara W, Derian N, Johanet C, Drakos I et al (2018) Clinical and multi-omics cross-phenotyping of patients with autoimmune and autoinflammatory diseases: the observational transimmunom protocol. BMJ Open 8:e021037. https://doi.org/10.1136/bmjopen-2017-021037
Lu X, Cao X, Li F, Li J, Xiong J, Long G, Cao S, Xie S (2016) Comparative transcriptome analysis reveals a global insight into molecular processes regulating citrate accumulation in sweet orange (Citrus sinensis). Physiol Plant 158:463–482. https://doi.org/10.1111/ppl.12484
Ma B, Yuan Y, Gao M, Xing L, Li C, Li M, Ma F (2018) Genome-wide identification, classification, molecular evolution and expression analysis of malate dehydrogenases in apple. Int J Mol Sci 19:3312. https://doi.org/10.3390/ijms19113312
Metsalu T, Vilo J (2015) ClustVis: a web tool for visualizing clustering of multivariate data using principal component analysis and heatmap. Nucleic Acids Res 43:W566–W570. https://doi.org/10.1093/NAR/GKV468
Paczkowska M, Barenboim J, Sintupisut N, Fox NS, Zhu H, Abd-Rabbo D, Mee AW, Boutros P (2020) Integrative pathway enrichment analysis of multivariate omics data. Nat Commun 11:735. https://doi.org/10.1038/s41467-019-13983-9
Pan Z, Li Y, Deng X, Xiao S (2014) Non-targeted metabolomic analysis of orange (Citrus sinensis [L.] Osbeck) wild type and bud mutant fruits by direct analysis in real-time and HPLC-electrospray mass spectrometry. Metabolomics 10:508–523. https://doi.org/10.1007/s11306-013-0597-7
Piryonesi SM, El-Diraby TE (2020) Role of data analytics in infrastructure asset management: overcoming data size and quality problems. J Transp Eng B Pave 146:04020022. https://doi.org/10.1061/JPEODX.0000175
Pittman JK (2012) Multiple transport pathways for mediating intracellular pH homeostasis: the contribution of H+/ion exchangers. Front Plant Sci 3:11. https://doi.org/10.3389/fpls.2012.00011
Provost F, Fawcett T (2013) Data science for business: what you need to know about data mining and data-analytic thinking, 1st edn. O’Reilly Media, Sebastopol, Calif.
Sadka A, Dahan E, Or E, Roose ML, Marsh KB, Cohen L (2001) Comparative analysis of mitochondrial citrate synthase gene structure, transcript level and enzymatic activity in acidless and acid-containing Citrus varieties. Funct Plant Biol 28:383–390. https://doi.org/10.1071/PP00136
Shaik R, Ramakrishna W (2014) Machine learning approaches distinguish multiple stress conditions using stress-responsive genes and identify candidate genes for broad resistance in rice. Plant Physiol 164:481–495. https://doi.org/10.1104/pp.113.225862
Song J, Du L, Li L, Kalt W, Palmer LC, Fillmore S, Zhang Y, Zhang Z, Li X (2015) Quantitative changes in proteins responsible for flavonoid and anthocyanin biosynthesis in strawberry fruit at different ripening stages: a targeted quantitative proteomic investigation employing multiple reaction monitoring. J Proteomics 122:1–10. https://doi.org/10.1016/j.jprot.2015.03.017
Strazzer P, Spelt CE, Li S, Bliek M, Federici CT, Roose ML, Koes R, Quattrocchio FM (2019) Hyperacidification of citrus fruits by a vacuolar proton-pumping P-ATPase complex. Nat Commun 10:1–11. https://doi.org/10.1038/s41467-019-08516-3
Terol J, Soler G, Talon M et al (2010) The aconitate hydratase family from Citrus. BMC Plant Biol 10:222. https://doi.org/10.1186/1471-2229-10-222
Van de Poel B, Bulens I, Oppermann Y, Hertog ML, Nicolai BM, Sauter M, Geeraerd AH (2013) S-adenosyl-l-methionine usage during climacteric ripening of tomato in relation to ethylene and polyamine biosynthesis and transmethylation capacity. Physiol Plant 148:176–188. https://doi.org/10.1111/j.1399-3054.2012.01703.x
Zhang L, Li H, Gao L, Qi Y, Fu W, Li X, Zhou X, Gao Q, Gao Z, Jia H (2017) Acyl-CoA oxidase 1 is involved in γ-decalactone release from peach (Prunus persica) fruit. Plant Cell Rep 36:829–842. https://doi.org/10.1007/s00299-017-2113-4
Zinati Z, Alemzadeh A, KayvanJoo AH (2016) Computational approaches for classification and prediction of P-type ATPase substrate specificity in Arabidopsis. Physiol Mol Boil Plants 22:163–174. https://doi.org/10.1007/s12298-016-0351-5
Acknowledgements
We would like to greatly thank Department of Agroecology of Agriculture and Natural Resources of Darab for supporting this research.
Author information
Authors and Affiliations
Contributions
ZZ designed the research, scientifically analyzed the results, wrote and edited the manuscript. SS wrote and edited the manuscript. HA designed the research and edited the manuscript. AT preformed the analysis and edited the manuscript. All of the authors approved the final version of the manuscript.
Corresponding author
Ethics declarations
Conflict of interest
The authors declare no conflict of interest.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Communicated by Heakeun Yun, Ph.D.
Supplementary Information
Below is the link to the Supplementary Information.
Rights and permissions
About this article
Cite this article
Zinati, Z., Sazegari, S., Amin, H. et al. Mining transcriptome data to identify genes and pathways related to lemon taste using supervised and unsupervised data learning methods. Hortic. Environ. Biotechnol. 62, 593–603 (2021). https://doi.org/10.1007/s13580-021-00337-y
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s13580-021-00337-y