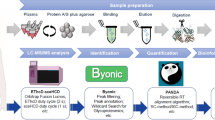
Abstract
The prevalence of oral squamous cell carcinoma (OSCC) is high in South and Southeast Asia regions. Most OSCC patients are detected at advanced stages low 5-year survival rates. Aberrant expression of glycosylated proteins was found to be associated with malignant transformation and cancer progression. Hence, identification of cancer-associated glycoproteins could be used as potential biomarkers that are beneficial for diagnosis or clinical management of patients. This study aims to identify the differentially expressed glycoproteins using lectin-based glycoproteomics approaches. Serum samples of 40 patients with OSCC, 10 patients with oral potentially malignant disorder (OPMD), and 10 healthy individuals as control group were subjected to two-dimensional gel electrophoresis (2-DE) coupled with lectin Concanavalin A and Jacalin that specifically bind to N- and O-glycosylated proteins, respectively. Five differentially expressed N- and O-glycoproteins with various potential glycosylation sites were identified, namely N-glycosylated α1-antitrypsin (AAT), α2-HS-glycoprotein (AHSG), apolipoprotein A-I (APOA1), and haptoglobin (HP); as well as O-glycosylated AHSG and clusterin (CLU). Among them, AAT and APOA1 were further validated using enzyme-linked immunosorbent assay (ELISA) (n = 120). It was found that AAT and APOA1 are significantly upregulated in OSCC and these glycoproteins are independent risk factors of OSCC. The clinical utility of AAT and APOA1 as potential biomarkers of OSCC is needed for further evaluation.




Similar content being viewed by others
References
Ferlay, J., Ervik, M., Lam, F., Colombet, M., Mery, L., Piñeros, M., et al.: Global cancer observatory: cancer today. Lyon, France, International Agency for Research on Cancer (2018)
Gupta, N., Gupta, R., Acharya, A.K., Patthi, B., Goud, V., Reddy, S., Garg, A., Singla, A.: Changing trends in oral cancer - a global scenario. Nepal J Epidemiol. 6, 613–619 (2016). https://doi.org/10.3126/nje.v6i4.17255
Speight, P.M., Khurram, S.A., Kujan, O.: Oral potentially malignant disorders: risk of progression to malignancy. Oral Surg. Oral Med. Oral Pathol. Oral Radiol. 125, 612–627 (2018). https://doi.org/10.1016/j.oooo.2017.12.011
Johnson, N.W., Jayasekara, P., Amarasinghe, A.A.: Squamous cell carcinoma and precursor lesions of the oral cavity: epidemiology and aetiology. Periodontol 2000. 57, 19–37 (2011). https://doi.org/10.1111/j.1600-0757.2011.00401.x
Warnakulasuriya, S.: Global epidemiology of oral and oropharyngeal cancer. Oral Oncol. 45, 309–316 (2009). https://doi.org/10.1016/j.oraloncology.2008.06.002
Le Campion, A., Ribeiro, C.M.B., Luiz, R.R., da Silva Junior, F.F., Barros, H.C.S., Dos Santos, K.C.B., et al.: Low survival rates of oral and oropharyngeal squamous cell carcinoma. Int J Dent. 2017, 5815493–5815497 (2017). https://doi.org/10.1155/2017/5815493
Montero, P.H., Patel, S.G.: Cancer of the oral cavity. Surg. Oncol. Clin. N. Am. 24, 491–508 (2015). https://doi.org/10.1016/j.soc.2015.03.006
Roth, Z., Yehezkel, G., Khalaila, I.: Identification and quantification of protein glycosylation. Int. J. Carbohydr. Chem. 2012, 1–10 (2012). https://doi.org/10.1155/2012/640923
Munkley, J., Elliott, D.J.: Hallmarks of glycosylation in cancer. Oncotarget. 7, 35478–35489 (2016). https://doi.org/10.18632/oncotarget.8155
Stowell, S.R., Ju, T., Cummings, R.D.: Protein glycosylation in cancer. Annu. Rev. Pathol. 10, 473–510 (2015). https://doi.org/10.1146/annurev-pathol-012414-040438
Kawahara, R., Ortega, F., Rosa-Fernandes, L., Guimaraes, V., Quina, D., Nahas, W., et al.: Distinct urinary glycoprotein signatures in prostate cancer patients. Oncotarget. 9, 33077–33097 (2018). https://doi.org/10.18632/oncotarget.26005
Pan, S., Brentnall, T.A., Chen, R.: Glycoproteins and glycoproteomics in pancreatic cancer. World J. Gastroenterol. 22, 9288–9299 (2016). https://doi.org/10.3748/wjg.v22.i42.9288
Song, E., Mechref, Y.: Defining glycoprotein cancer biomarkers by MS in conjunction with glycoprotein enrichment. Biomark. Med. 9, 835–844 (2015). https://doi.org/10.2217/bmm.15.55
Kailemia, M.J., Park, D., Lebrilla, C.B.: Glycans and glycoproteins as specific biomarkers for cancer. Anal. Bioanal. Chem. 409, 395–410 (2017). https://doi.org/10.1007/s00216-016-9880-6
Lin, W.L., Lin, Y.S., Shi, G.Y., Chang, C.F., Wu, H.L.: Lewisy promotes migration of oral cancer cells by glycosylation of epidermal growth factor receptor. PLoS One. 10, e0120162 (2015). https://doi.org/10.1371/journal.pone.0120162
Manoharan, S., Padmanabhan, M., Kolanjiappan, K., Ramachandran, C.R., Suresh, K.: Analysis of glycoconjugates in patients with oral squamous cell carcinoma. Clin. Chim. Acta. 339, 91–96 (2004). https://doi.org/10.1016/j.cccn.2003.09.006
Liu, G., Sengupta, P.K., Jamal, B., Yang, H.Y., Bouchie, M.P., Lindner, V., Varelas, X., Kukuruzinska, M.A.: N-glycosylation induces the CTHRC1 protein and drives oral cancer cell migration. J. Biol. Chem. 288, 20217–20227 (2013). https://doi.org/10.1074/jbc.M113.473785
Lin, M.C., Huang, M.J., Liu, C.H., Yang, T.L., Huang, M.C.: GALNT2 enhances migration and invasion of oral squamous cell carcinoma by regulating EGFR glycosylation and activity. Oral Oncol. 50, 478–484 (2014). https://doi.org/10.1016/j.oraloncology.2014.02.003
Hashim, O.H., Jayapalan, J.J., Lee, C.S.: Lectins: an effective tool for screening of potential cancer biomarkers. PeerJ. 5, e3784 (2017). https://doi.org/10.7717/peerj.3784
Chen, Y., Lim, B.K., Peh, S.C., Abdul-Rahman, P.S., Hashim, O.H.: Profiling of serum and tissue high abundance acute-phase proteins of patients with epithelial and germ line ovarian carcinoma. Proteome Sci. 6, 20 (2008). https://doi.org/10.1186/1477-5956-6-20
Pinho, S.S., Reis, C.A.: Glycosylation in cancer: mechanisms and clinical implications. Nat. Rev. Cancer. 15, 540–555 (2015). https://doi.org/10.1038/nrc3982
Chang, S.C., Lin, W.L., Chang, Y.F., Lee, C.T., Wu, J.S., Hsu, P.H., Chang, C.F.: Glycoproteomic identification of novel plasma biomarkers for oral cancer. J. Food Drug Anal. 27, 483–493 (2019). https://doi.org/10.1016/j.jfda.2018.12.008
Chen, J.T., Chen, C.H., Ku, K.L., Hsiao, M., Chiang, C.P., Hsu, T.L., Chen, M.H., Wong, C.H.: Glycoprotein B7-H3 overexpression and aberrant glycosylation in oral cancer and immune response. Proc. Natl. Acad. Sci. U. S. A. 112, 13057–13062 (2015). https://doi.org/10.1073/pnas.1516991112
Chen, Y.T., Chong, Y.M., Cheng, C.W., Ho, C.L., Tsai, H.W., Kasten, F.H., Chen, Y.L., Chang, C.F.: Identification of novel tumor markers for oral squamous cell carcinoma using glycoproteomic analysis. Clin. Chim. Acta. 420, 45–53 (2013). https://doi.org/10.1016/j.cca.2012.10.019
Guu, S.Y., Lin, T.H., Chang, S.C., Wang, R.J., Hung, L.Y., Fang, P.J., Tang, W.C., Yu, P., Chang, C.F.: Serum N-glycome characterization and anti-carbohydrate antibody profiling in oral squamous cell carcinoma patients. PLoS One. 12, e0178927 (2017). https://doi.org/10.1371/journal.pone.0178927
Saraswat, M., Makitie, A., Tohmola, T., Dickinson, A., Saraswat, S., Joenvaara, S., et al.: Tongue cancer patients can be distinguished from healthy controls by specific N-Glycopeptides found in serum. Proteomics Clin Appl. 12, e1800061 (2018). https://doi.org/10.1002/prca.201800061
Pan, S., Chen, R., Tamura, Y., Crispin, D.A., Lai, L.A., May, D.H., McIntosh, M.W., Goodlett, D.R., Brentnall, T.A.: Quantitative glycoproteomics analysis reveals changes in N-glycosylation level associated with pancreatic ductal adenocarcinoma. J. Proteome Res. 13, 1293–1306 (2014). https://doi.org/10.1021/pr4010184
Shah, P., Wang, X., Yang, W., Toghi Eshghi, S., Sun, S., Hoti, N., Chen, L., Yang, S., Pasay, J., Rubin, A., Zhang, H.: Integrated proteomic and Glycoproteomic analyses of prostate Cancer cells reveal glycoprotein alteration in protein abundance and glycosylation. Mol. Cell. Proteomics. 14, 2753–2763 (2015). https://doi.org/10.1074/mcp.M115.047928
Clerc, F., Reiding, K.R., Jansen, B.C., Kammeijer, G.S., Bondt, A., Wuhrer, M.: Human plasma protein N-glycosylation. Glycoconj. J. 33, 309–343 (2016). https://doi.org/10.1007/s10719-015-9626-2
Rodriguez-Pineiro, A.M., de la Cadena, M.P., Lopez-Saco, A., Rodriguez-Berrocal, F.J.: Differential expression of serum clusterin isoforms in colorectal cancer. Mol. Cell. Proteomics. 5, 1647–1657 (2006). https://doi.org/10.1074/mcp.M600143-MCP200
Sarrats, A., Saldova, R., Pla, E., Fort, E., Harvey, D.J., Struwe, W.B., de Llorens, R., Rudd, P.M., Peracaula, R.: Glycosylation of liver acute-phase proteins in pancreatic cancer and chronic pancreatitis. Proteomics Clin Appl. 4, 432–448 (2010). https://doi.org/10.1002/prca.200900150
Wu, J., Xie, X., Nie, S., Buckanovich, R.J., Lubman, D.M.: Altered expression of sialylated glycoproteins in ovarian cancer sera using lectin-based ELISA assay and quantitative glycoproteomics analysis. J. Proteome Res. 12, 3342–3352 (2013). https://doi.org/10.1021/pr400169n
Chaerkady, R., Thuluvath, P.J., Kim, M.S., Nalli, A., Vivekanandan, P., Simmers, J., Torbenson, M., Pandey, A.: O Labeling for a quantitative proteomic analysis of glycoproteins in hepatocellular carcinoma. Clin. Proteomics. 4, 137–155 (2008). https://doi.org/10.1007/s12014-008-9013-0
Qi, Y.J., Ward, D.G., Pang, C., Wang, Q.M., Wei, W., Ma, J., Zhang, J., Lou, Q., Shimwell, N.J., Martin, A., Wong, N., Chao, W.X., Wang, M., Ma, Y.F., Johnson, P.J.: Proteomic profiling of N-linked glycoproteins identifies ConA-binding procathepsin D as a novel serum biomarker for hepatocellular carcinoma. Proteomics. 14, 186–195 (2014). https://doi.org/10.1002/pmic.201300226
Liang, Y., Ma, T., Thakur, A., Yu, H., Gao, L., Shi, P., Li, X., Ren, H., Jia, L., Zhang, S., Li, Z., Chen, M.: Differentially expressed glycosylated patterns of alpha-1-antitrypsin as serum biomarkers for the diagnosis of lung cancer. Glycobiology. 25, 331–340 (2015). https://doi.org/10.1093/glycob/cwu115
Semaan, S.M., Wang, X., Marshall, A.G., Sang, Q.X.: Identification of potential glycoprotein biomarkers in estrogen receptor positive (ER+) and negative (ER-) human breast cancer tissues by LC-LTQ/FT-ICR mass spectrometry. J. Cancer. 3, 269–284 (2012). https://doi.org/10.7150/jca.4592
Yang, N., Feng, S., Shedden, K., Xie, X., Liu, Y., Rosser, C.J., Lubman, D.M., Goodison, S.: Urinary glycoprotein biomarker discovery for bladder cancer detection using LC/MS-MS and label-free quantification. Clin. Cancer Res. 17, 3349–3359 (2011). https://doi.org/10.1158/1078-0432.CCR-10-3121
Ercetin, E., Richtmann, S., Delgado, B.M., Gomez-Mariano, G., Wrenger, S., Korenbaum, E., et al.: Clinical significance of SERPINA1 gene and its encoded alpha1-antitrypsin protein in NSCLC. Cancers (Basel). 11, (2019). https://doi.org/10.3390/cancers11091306
Zhao, Z., Ma, J., Mao, Y., Dong, L., Li, S., Zhang, Y.: Silence of alpha1-antitrypsin inhibits migration and proliferation of triple negative breast cancer cells. Med Sci Monit. 24, 6851–6860 (2018). https://doi.org/10.12659/MSM.910665
Sun, Z., Yang, P.: Role of imbalance between neutrophil elastase and α1-antitrypsin in cancer development and progression. The lancet oncology. 5, 182–190 (2004). https://doi.org/10.1016/S1470-2045(04)01414-7
Ehlers, M.R.: Immune-modulating effects of alpha-1 antitrypsin. Biol. Chem. 395, 1187–1193 (2014). https://doi.org/10.1515/hsz-2014-0161
Csosz, E., Markus, B., Darula, Z., Medzihradszky, K.F., Nemes, J., Szabo, E., et al.: Salivary proteome profiling of oral squamous cell carcinoma in a Hungarian population. FEBS Open Bio. 8, 556–569 (2018). https://doi.org/10.1002/2211-5463.12391
Jessie, K., Jayapalan, J.J., Ong, K.C., Abdul Rahim, Z.H., Zain, R.M., Wong, K.T., Hashim, O.H.: Aberrant proteins in the saliva of patients with oral squamous cell carcinoma. Electrophoresis. 34, 2495–2502 (2013). https://doi.org/10.1002/elps.201300107
Kawahara, R., Bollinger, J.G., Rivera, C., Ribeiro, A.C., Brandao, T.B., Paes Leme, A.F., et al.: A targeted proteomic strategy for the measurement of oral cancer candidate biomarkers in human saliva. Proteomics. 16, 159–173 (2016). https://doi.org/10.1002/pmic.201500224
Guo, X., Hao, Y., Kamilijiang, M., Hasimu, A., Yuan, J., Wu, G., Reyimu, H., Kadeer, N., Abudula, A.: Potential predictive plasma biomarkers for cervical cancer by 2D-DIGE proteomics and ingenuity pathway analysis. Tumour Biol. 36, 1711–1720 (2015). https://doi.org/10.1007/s13277-014-2772-5
Hao, Y., Li, D., Xu, Y., Ouyang, J., Wang, Y., Zhang, Y., Li, B., Xie, L., Qin, G.: Investigation of lipid metabolism dysregulation and the effects on immune microenvironments in pan-cancer using multiple omics data. BMC Bioinformatics. 20, 195 (2019). https://doi.org/10.1186/s12859-019-2734-4
Lim, L.C., Looi, M.L., Zakaria, S.Z., Sagap, I., Rose, I.M., Chin, S.F., et al.: Identification of differentially expressed proteins in the serum of colorectal Cancer patients using 2D-DIGE proteomics analysis. Pathol Oncol Res. 22, 169–177 (2016). https://doi.org/10.1007/s12253-015-9991-y
Ren, L., Yi, J., Li, W., Zheng, X., Liu, J., Wang, J., du, G.: Apolipoproteins and cancer. Cancer Med. 8, 7032–7043 (2019). https://doi.org/10.1002/cam4.2587
Bunkenborg, J., Pilch, B.J., Podtelejnikov, A.V., Wisniewski, J.R.: Screening for N-glycosylated proteins by liquid chromatography mass spectrometry. Proteomics. 4, 454–465 (2004). https://doi.org/10.1002/pmic.200300556
Campos, D., Freitas, D., Gomes, J., Magalhaes, A., Steentoft, C., Gomes, C., et al.: Probing the O-glycoproteome of gastric cancer cell lines for biomarker discovery. Mol. Cell. Proteomics. 14, 1616–1629 (2015). https://doi.org/10.1074/mcp.M114.046862
Camisasca, D.R., da Ros, G.L., Soares, M.R., Sandim, V., Nogueira, F.C., Garcia, C.H., et al.: A proteomic approach to compare saliva from individuals with and without oral leukoplakia. J. Proteome. 151, 43–52 (2017). https://doi.org/10.1016/j.jprot.2016.07.029
Lo, W.Y., Lai, C.C., Hua, C.H., Tsai, M.H., Huang, S.Y., Tsai, C.H., Tsai, F.J.: S100A8 is identified as a biomarker of HPV18-infected oral squamous cell carcinomas by suppression subtraction hybridization, clinical proteomics analysis, and immunohistochemistry staining. J. Proteome Res. 6, 2143–2151 (2007). https://doi.org/10.1021/pr060551+
Liao, K.A., Tsay, Y.G., Huang, L.C., Huang, H.Y., Li, C.F., Wu, T.F.: Search for the tumor-associated proteins of oral squamous cell carcinoma collected in Taiwan using proteomics strategy. J. Proteome Res. 10, 2347–2358 (2011). https://doi.org/10.1021/pr101146w
Tung, C.L., Lin, S.T., Chou, H.C., Chen, Y.W., Lin, H.C., Tung, C.L., Huang, K.J., Chen, Y.J., Lee, Y.R., Chan, H.L.: Proteomics-based identification of plasma biomarkers in oral squamous cell carcinoma. J. Pharm. Biomed. Anal. 75, 7–17 (2013). https://doi.org/10.1016/j.jpba.2012.11.017
Acknowledgements
The authors would like to thank the Oral Cancer Research & Coordinating Centre, University of Malaya (OCRCC-UM) for providing serum samples and data from the Malaysian Oral Cancer Database and Tissue Bank System (MOCDTBS).
Funding
This work was supported by the University of Malaya Postgraduate Research Grant (PPP) PG326–2016A and University of Malaya (UM) High Impact Research (HIR) MoE Grants UM.C/625/1/HIR/MOE/DENT/09 and UM.C/625/1/HIR/MOHE/MED/16/5 from the Ministry of Education Malaysia.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no conflicts of interest.
Ethical approval
This article does not contain any studies with human participants or animals performed by any of the authors.
Additional information
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Rights and permissions
About this article
Cite this article
Wong, YL., Anand, R., Yuen, K.M. et al. Identification of potential glycoprotein biomarkers in oral squamous cell carcinoma using sweet strategies. Glycoconj J 38, 1–11 (2021). https://doi.org/10.1007/s10719-021-09973-z
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10719-021-09973-z