Abstract
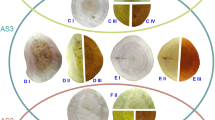
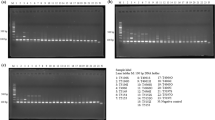
Illegal and unsustainable logging of forests have contributed to deforestation and a massive loss of biodiversity, and despite the efforts to regulate the trade of plants and combat the commercialization of illegal logging, the problem continues. DNA analysis of plant tissues is an effective method in species identification, and it is important to dedicate efforts to obtain and amplify DNA from dry wood or exposed to a sawmill process in which its origin is not clear. The aim of this research was to present a strategy to identify tree species from sawn timber and roundwood, based on the use of molecular identification tools. Wood samples of standing trees of Pinus pseudostrobus (PFCap1), round core samples (Ro) and plank wood (SaT) of Pinus pseudostrobus and Pinus devoniana were used. All the samples underwent a DNA extraction protocol, amplification, sequencing, and analysis in the NCBI database. The results showed an efficient extraction of the genetic material, revealing high purity ratios for the three sample groups (PFCap1, Ro and SaT) and quantitative data of the amplified DNA. The information obtained through sequencing showed identities of up to 100% homology (NCBI database). We can conclude that the standardized method developed was successful using the trnL and trnLF primers, managing to isolate, amplify, and sequence the DNA from sawn timber and roundwood, and to identify the age of the tree and the time elapsed between the felling of the tree and the sampling, which are directly related to the success of this method.



Similar content being viewed by others
References
Asif MJ, Cannon CH (2005) DNA extraction from processed wood: a case study for the identification of an endangered timber species (Gonystylus bancanus). Plant Mol Biol Rep 23(2):185–192. https://doi.org/10.1007/BF02772709
Benouadah N, Pranovich A, Aliouche D, Hemming J, Smeds A, Willför S (2018) Analysis of extractives from Pinus halepensis and Eucalyptus camaldulensis as predominant trees in Algeria. Holzforschung 72(2):97–104. https://doi.org/10.1515/hf-2017-0098
Carballo-Meilán A, Goodman AM, Baron MG, Gonzalez-Rodriguez J (2016) Application of chemometric analysis to infrared spectroscopy for the identification of wood origin. Cellulose 23(1):901–913. https://doi.org/10.1007/s10570-015-0848-z
de Souza KCA, de Melo DS, Galvão E, Mendonça RCCL, Leandro AS, de Souza SR (2017) DNA extraction and anatomic characterization in dried heartwood from Fabaceae species. Wood Res 62(1):13–26
Donaldson LA, Nanayakkara B, Radotić K, Djikanovic-Golubović D, Mitrović A, Pristov JB, Kalauzi A (2015) Xylem parenchyma cell walls lack a gravitropic response in conifer compression wood. Planta 242(6):1413–1424. https://doi.org/10.1007/s00425-015-2381-6
Dormontt EE, Boner M, Braun B, Breulmann G, Degen B, Espinoza E, Lee SL (2015) Forensic timber identification: It’s time to integrate disciplines to combat illegal logging. Biol Conserv 191:790–798. https://doi.org/10.1016/j.biocon.2015.06.038
Doyle J (1991) DNA protocols for plants. In Molecular techniques in taxonomy. JPW Young 57:283–293
Fatima T, Srivastava A, Hanur VS, Rao MS (2018) An effective wood DNA extraction protocol for three economic important timber species of India. American Journal of Plant Sciences 9(2):139–149. https://doi.org/10.4236/ajps.2018.92012
Funada R, Yamagishi Y, Begum S, Kudo K, Nabeshima E, Nugroho WD, Nakaba S (2016) Xylogenesis in trees: from cambial cell division to cell death. Secondary Xylem Biology. https://doi.org/10.1016/B978-0-12-802185-9.00002-4
Funda T, Fundova I, Gorzsás A, Fries A, Wu HX (2020) Predicting the chemical composition of juvenile and mature woods in Scots pine (Pinus sylvestris L.) using FTIR spectroscopy. Wood Sci Technol 54(2):289–311. https://doi.org/10.1007/s00226-020-01159-4
Gasson P (2011) How precise can wood identification be? Wood anatomy’s role in support of the legal timber trade, especially CITES. IAWA journal 32(2):137–154. https://doi.org/10.1163/22941932-90000049
Gómez-Zeledón J, Grasse W, Runge F, Land A, Spring O (2017) TaqMan qPCR pushes boundaries for the analysis of millennial wood. J Archaeol Sci 79:53–61. https://doi.org/10.1016/j.jas.2017.01.010
Hartvig I, Czako M, Kjaer ED, Nielsen LR, Theilade I (2015) The use of DNA barcoding in identification and conservation of rosewood (Dalbergia spp.). PLoS ONE. https://doi.org/10.1371/2Fjournal.pone.0138231
He T, Jiao L, Yu M, Guo J, Jiang X, Yin Y (2019) DNA barcoding authentication for the wood of eight endangered Dalbergia timber species using machine learning approaches. Holzforschung 73(3):277–285. https://doi.org/10.1515/hf-2018-0076
Higuchi T (1997) Biochemistry and molecular biology of wood. Springer Verlag, London
Horres R, Zizka G, Kahl G, Weising K (2000) Molecular phylogenetics of Bromeliaceae: evidence from trnL (UAA) intron sequences of the chloroplast genome. Plant Biol 2(3):306–315. https://doi.org/10.1055/s-2000-3700
Japelaghi RH, Haddad R, Garoosi GA (2011) Rapid and efficient isolation of high-quality nucleic acids from plant tissues rich in polyphenols and polysaccharides. Mol Biotechnol 49(2):129–137. https://doi.org/10.1007/s12033-011-9384-8
Jiao L, Yin Y, Xiao F, Sun Q, Song K, Jiang X (2012) Comparative analysis of two DNA extraction protocols from fresh and dried wood of Cunninghamia lanceolata (Taxodiaceae). IAWA Journal 33(4):441–456. https://doi.org/10.1163/22941932-90000106
Jiao L, Yu M, Wiedenhoeft AC, He T, Li J, Liu B, Yin Y (2018) DNA barcode authentication and library development for the wood of six commercial Pterocarpus species: the critical role of Xylarium specimens. Sci Rep 8(1):1–10. https://doi.org/10.1038/s41598-018-20381-6
Jiao L, He T, Dormontt EE, Zhang Y, Lowe AJ, Yin Y (2019) Applicability of chloroplast DNA barcodes for wood identification between Santalum album and its adulterants. Holzforschung 73(2):209–218. https://doi.org/10.1515/hf-2018-0047
Lowe AJ, Dormontt EE, Bowie MJ, Degen B, Gardner S, Thomas D, Sasaki N (2016) Opportunities for improved transparency in the timber trade through scientific verification. Bioscience 66(11):990–998. https://doi.org/10.1093/biosci/biw129
MacDicken K, Reams G, de Freitas J (2015) Introduction to the Changes in Global forest resources from 1990 to 2015. Forest Ecol Manag. https://doi.org/10.1016/2Fj.foreco.2015.06.018
Méndez-Cea B, Cobo-Simón I, Pérez-González A, García-García I, Linares JC, Rodríguez FJG (2019) DNA extraction and amplification from Pinaceae dry wood. Silvae Genet 68(1):55–57. https://doi.org/10.2478/sg-2019-0010
Miranda I, Sousa V, Ferreira J, Pereira H (2017) Chemical characterization and extractives composition of heartwood and sapwood from Quercus faginea. PLoS ONE 12(6):e0179268. https://doi.org/10.1371/journal.pone.0179268
Rachmayanti Y, Leinemann L, Gailing O, Finkeldey R (2006) Extraction, amplification and characterization of wood DNA from Dipterocarpaceae. Plant Mol Biol Rep 24(1):45–55. https://doi.org/10.1007/BF02914045
Rachmayanti Y, Leinemann L, Gailing O, Finkeldey R (2009) DNA from processed and unprocessed wood: factors influencing the isolation success. Forensic Sci Int 3(3):185–192. https://doi.org/10.1016/j.fsigen.2009.01.002
Ragheb HH, Elalfy MM, Ali FR (2019) Biochemicals and DNA degradation identifier markers of postmortem time interval in different causes of death. Vet Open A Open J 1(1):1–11
Soranzo N, Provan J, Powell W (1998) Characterization of microsatellite loci in Pinus sylvestris L. Mol Ecol 7:1260–1261
Staats M, Cuenca A, Richardson JE, Vrielink-van GR, Petersen G, Seberg O, Bakker FT (2011) DNA damage in plant herbarium tissue. PLoS ONE. https://doi.org/10.1371/journal.pone.0028448
Taberlet P, Gielly L, Pautou G, Bouvet J (1991) Universal primers for amplification of three non-coding regions of chloroplast DNA. Plant Mol Biol 17(5):1105–1109. https://doi.org/10.1007/BF00037152
Tanaka S, Ito M (2020) DNA barcoding for identification of agarwood source species using trnL-trnF and matK DNA sequences. J Nat Med 74(1):42–50. https://doi.org/10.1007/s11418-019-01338-z
Tang X, Zhao G, Ping L (2011) Wood identification with PCR targeting noncoding chloroplast DNA. Plant Mol Biol 77(6):609–617
Tiwari S, Tomar RS, Tripathi MK, Ahuja A (2017) Modified Protocol for Plant Genomic DNA Isolation. Indian Res J Genet & Biotech 9(4):478–485
Weimar H, Janzen N, Dieter M (2015) Market coverage of wood imports by the EU Timber Regulation. Thünen Working Paper. https://doi.org/10.3220/WP1440577266000
Yu M, Liu K, Zhou L, Zhao L, Liu S (2016) Testing three proposed DNA barcodes for the wood identification of Dalbergia odorifera T. Chen and Dalbergia tonkinensis Prain Holzforschung 70(2):127–136. https://doi.org/10.1515/hf-2014-0234
Acknowledgements
To the National Council of Science and Technology (CONACyT) and the Coordination of Scientific Research of our university (UMSNH), RVR†.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that no competing interests exist.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
About this article
Cite this article
Murillo-Sánchez, I.E., López-Albarrán, P., Santoyo-Pizano, G. et al. Molecular identification of timber species from sawn timber and roundwood. Conservation Genet Resour 13, 191–200 (2021). https://doi.org/10.1007/s12686-021-01193-9
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s12686-021-01193-9