Abstract
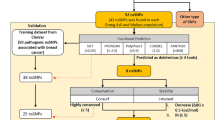
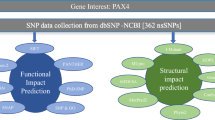
Researches have revealed that functional non-synonymous Single Nucleotide Polymorphism (nsSNPs) present in the Zinc-finger with UFM1-Specific Peptidase domain protein (ZUFSP) may be involved in genetic instability and carcinogenesis. For the first time, we employed in-silico approach using predictive tools to identify and validate potential nsSNPs that could be pathogenic. Our result revealed that 8 nsSNPs (rs 112738382, rs 140094037, rs 201652589, rs 201847265, rs 202076827, rs 373634906, rs 375114528, rs 772591104) are pathogenic after being subjected to rigorous filtering process. The structural impact of the nsSNPs on ZUFSP structure indicated that the nsSNPs affect the stability of the protein by lowering ZUFSP protein stability. Furthermore, conservation analysis showed that rs 201652589, rs 140094037, rs 201847265, and rs 772591104 were highly conserved. Interestingly, the protein–protein affinity between ZUFSP and Ubiquitin was altered rs 201652589, rs 140094037, rs 201847265, and rs 772591104 had a binding affinity of − 0.46, − 0.83, − 1.62, and − 1.12 kcal/mol respectively. Our study has been able to identify potential nsSNPs that could be used as genetic biomarkers for some diseases arising as a result of aberration in the ZUFSP structure, however, being a predictive study, the identified nsSNPs need to be experimentally investigated.








Similar content being viewed by others
References
Baker RT, Tobias JW, Varshavsky A (1992) Ubiquitin-specific proteases of Saccharomyces cerevisiae. Cloning of UBP2 and UBP3, and functional analysis of the UBP gene family. J Biol Chem 267:23364–23375
Matsui SI, Sandberg AA, Negoro S et al (1982) Isopeptidase: a novel eukaryotic enzyme that cleaves isopeptide bonds. Proc Natl Acad Sci USA. https://doi.org/10.1073/pnas.79.5.1535
Poondla N, Chandrasekaran AP, Kim KS, Ramakrishna S (2019) Deubiquitinating enzymes as cancer biomarkers: new therapeutic opportunities? BMB Rep. https://doi.org/10.5483/BMBRep.2019.52.3.048
Poondla N, Chandrasekaran AP, Kim KS, Ramakrishna S (2019) Deubiquitinating enzymes as cancer biomarkers: new therapeutic opportunities? BMB Rep 52:181–189
Mevissen TET, Komander D (2017) Mechanisms of deubiquitinase specificity and regulation. Annu Rev Biochem. https://doi.org/10.1146/annurev-biochem-061516-044916
Nijman SMB, Luna-Vargas MPA, Velds A et al (2005) A genomic and functional inventory of deubiquitinating enzymes. Cell. https://doi.org/10.1016/j.cell.2005.11.007
Kwasna D, Abdul Rehman SA, Natarajan J et al (2018) Discovery and characterization of ZUFSP/ZUP1, a distinct deubiquitinase class important for genome stability. Mol Cell. https://doi.org/10.1016/j.molcel.2018.02.023
Hanpude P, Bhattacharya S, Dey AK, Maiti TK (2015) Deubiquitinating enzymes in cellular signaling and disease regulation. IUBMB Life. https://doi.org/10.1002/iub.1402
D’Arcy P, Wang X, Linder S (2015) Deubiquitinase inhibition as a cancer therapeutic strategy. Pharmacol Ther. https://doi.org/10.1016/j.pharmthera.2014.11.002
Jemal A, Bray F, Ferlay J (1999) Global cancer statistics: 2011. CA Cancer J Clin. https://doi.org/10.3322/caac.20107.Available
Landegren U, Nilsson M, Kwok PY (1998) Reading bits of genetic information: methods for single-nucleotide polymorphism analysis. Genome Res 8:769–776
Ke X, Taylor MS, Cardon LR (2008) Singleton SNPs in the human genome and implications for genome-wide association studies. Eur J Hum Genet 16:506–515. https://doi.org/10.1038/sj.ejhg.5201987
Shastry BS (2002) SNP alleles in human disease and evolution. J Hum Genet 47:561–566
Dakal TC, Kala D, Dhiman G et al (2017) Predicting the functional consequences of non-synonymous single nucleotide polymorphisms in IL8 gene. Sci Rep. https://doi.org/10.1038/s41598-017-06575-4
Sheryl ST et al (2001) dbSNP: the NCBI database of genetic variation. Nucleic Acids Res 29(1):308–311
Abdelraheem NE, El-tayeb GM, Osman LO, Abedlrhman SA (2016) A comprehensive in silico analysis of the functional and structural impact of non-synonymous single nucleotide polymorphisms in the human KRAS gene. Am J Bioinform Res 6:32–55. https://doi.org/10.5923/j.bioinformatics.20160602.02
Vaser R, Adusumalli S, Leng SN et al (2016) SIFT missense predictions for genomes. Nat Protoc 11:1–9. https://doi.org/10.1038/nprot.2015.123
Adzhubei IA, Schmidt S, Peshkin L et al (2010) A method and server for predicting damaging missense mutations. Nat Methods 7:248–249
Hecht M, Bromberg Y, Rost B (2015) Better prediction of functional effects for sequence variants. BMC Genom. https://doi.org/10.1186/1471-2164-16-S8-S1
Bromberg Y, Overton J, Vaisse C et al (2009) In silico mutagenesis: a case study of the melanocortin 4 receptor. FASEB J 23:3059–3069. https://doi.org/10.1096/fj.08-127530
Yachdav G, Hecht M, Pasmanik-Chor M et al (2014) HeatMapViewer: interactive display of 2D data in biology. F1000Research. https://doi.org/10.12688/f1000research.3-48.v1
Choi Y, Sims GE, Murphy S et al (2012) Predicting the functional effect of amino acid substitutions and indels. PLoS ONE 7:e46688. https://doi.org/10.1371/journal.pone.0046688
Capriotti E, Fariselli P, Rossi I, Casadio R (2008) A three-state prediction of single point mutations on protein stability changes. BMC Bioinform 9(Suppl 2):S6. https://doi.org/10.1186/1471-2105-9-S2-S6
Emidio C, Fariselli P (2017) PhD-SNPg: a webserver and lightweight tool for scoring single nucleotide variants. Nucleic Acids Res 45(W1): W247–W252
Ferrer-Costa C, Gelpí JL, Zamakola L, Parraga I, De La Cruz X, Orozco M (2005) PMUT: a web-based tool for the annotation of pathological mutations on proteins. Bioinformatics 21:3176–3178
Calabrese R, Capriotti E, Fariselli P et al (2009) Functional annotations improve the predictive score of human disease-related mutations in proteins. Hum Mutat 30:1237–1244. https://doi.org/10.1002/humu.21047
Emidio C, Fariselli P, Casadio R (2005) I-Mutant2.0: predicting stability changes upon mutation from the protein sequence or structure. Nucleic Acids Res 33(2):W306–W310. https://doi.org/10.1093/nar/gki375.
Cheng J, Randall A, Baldi P (2006) Prediction of protein stability changes for single-site mutations using support vector machines. Prot Struct Funct Genet 62:1125–1132. https://doi.org/10.1002/prot.20810
Liang-Tsung Huang, Gromiha MM, Ho S (2007) iPTREE-STAB: interpretable decision tree based method for predicting protein stability changes upon mutations. Bioinformatics 23(10):1292–1293. https://doi.org/10.1093/bioinformatics/btm100
Emidio C, Fariselli P, Casadio R (2005) I-Mutant2.0: predicting stability changes upon mutation from the protein sequence or structure. Nucleic Acids Res 33(2):W306–W310. https://doi.org/10.1093/nar/gki375.
Witvliet DK, et al. (2016) ELASPIC web-server: proteome-wide structure-based prediction of mutation effects on protein stability and binding affinity. Bioinformatics 32(10):1589–1591. https://doi.org/10.1093/bioinformatics/btw031
Pejaver V, Hsu WL, Xin F et al (2014) The structural and functional signatures of proteins that undergo multiple events of post-translational modification. Prot Sci 23:1077–1093. https://doi.org/10.1002/pro.2494
Petersen B, Petersen T. N., Andersen P., et al (2009) A generic method for assignment of reliability scores applied to solvent accessibility predictions. BMC Struct Biol, 9:51, 1-10. https://doi.org/10.1186/1472-6807-9-51
Rodrigues CHM, Myung Y, Pires DEV, Ascher DB (2019) MCSM-PPI2: predicting the effects of mutations on protein-protein interactions. Nucleic Acids Res 47:W338–W344. https://doi.org/10.1093/nar/gkz383
Case DA, Cheatham TE, Darden T et al (2005) The Amber biomolecular simulation programs. J Comput Chem 26:1668–1688. https://doi.org/10.1002/jcc.20290
Wang J, Wolf RM, Caldwell JW et al (2004) Development and testing of a general Amber force field. J Comput Chem 25:1157–1174. https://doi.org/10.1002/jcc.20035
Berendsen HJC, Postma JPM, Van Gunsteren WF et al (1984) Molecular dynamics with coupling to an external bath. J Chem Phys 81:3684–3690. https://doi.org/10.1063/1.448118
Ryckaert JP, Ciccotti G, Berendsen HJC (1977) Numerical integration of the Cartesian equations of motion of a system with constraints: molecular dynamics of n-alkanes. J Comput Phys 23:327–341. https://doi.org/10.1016/0021-9991(77)90098-5
Roe DR, Cheatham TE (2013) PTRAJ and CPPTRAJ: software for processing and analysis of molecular dynamics trajectory data. J Chem Theory Comput 9:3084–3095. https://doi.org/10.1021/ct400341p
Soremekun OS, Soliman MES (2019) From genomic variation to protein aberration: mutational analysis of single nucleotide polymorphism present in ULBP6 gene and implication in immune response. Comput Biol Med. https://doi.org/10.1016/j.compbiomed.2019.103354
Nailwal M, Chauhan JB (2017) In silico analysis of non-synonymous single nucleotide polymorphisms in human DAZL gene associated with male infertility. Syst Biol Reprod Med. https://doi.org/10.1080/19396368.2017.1305466
Suresh PS, Venkatesh T, Rajan T (2012) Single nucleotide polymorphisms in genes that are common targets of luteotropin and luteolysin in primate corpus luteum: computational exploration. Gene 511:353–357. https://doi.org/10.1016/j.gene.2012.09.076
Brender JR, Zhang Y (2015) Predicting the effect of mutations on protein-protein binding interactions through structure-based interface profiles. PLOS Comput Biol 11:e1004494. https://doi.org/10.1371/journal.pcbi.1004494
Wang LL, Li Y, Zhou SF (2009) A bioinformatics approach for the phenotype prediction of nonsynonymous single nucleotide polymorphisms in human cytochromes P450E. Drug Metab Dispos 37:977–991. https://doi.org/10.1124/dmd.108.026047
Acknowledgements
The authors appreciate the financial and infrastructural support of College of Health Sciences, UKZN and also acknowledge the Centre for High Performance Computing (CHPC, www.chpc.ac.za), Cape Town for provision of computational resource.
Funding
This research was not funded by any organization for the submitted work.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Ethical Approval
This research is an in-silico research and needed no ethical Approval or consent to participate.
Research Involving Human and Animal Rights
This research did not involve human or animal subjects.
Consent for Publication
The authors grant the publishers to publish the content of this research.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Ajadi, M.B., Soremekun, O.S., Adewumi, A.T. et al. Functional Analysis of Single Nucleotide Polymorphism in ZUFSP Protein and Implication in Pathogenesis. Protein J 40, 28–40 (2021). https://doi.org/10.1007/s10930-021-09962-z
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10930-021-09962-z