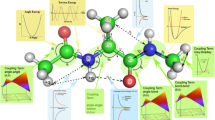
Abstract
Many molecular simulation methods use force fields to help model and simulate molecules and their behavior in various environments. Force fields are sets of functions and parameters used to calculate the potential energy of a chemical system as a function of the atomic coordinates. Despite the widespread use of force fields, their inadequacies are often thought to contribute to systematic errors in molecular simulations. Furthermore, different force fields tend to give varying results on the same systems with the same simulation settings. Here, we present a pipeline for comparing the geometries of small molecule conformers. We aimed to identify molecules or chemistries that are particularly informative for future force field development because they display inconsistencies between force fields. We applied our pipeline to a subset of the eMolecules database, and highlighted molecules that appear to be parameterized inconsistently across different force fields. We then identified over-represented functional groups in these molecule sets. The molecules and moieties identified by this pipeline may be particularly helpful for future force field parameterization.




Similar content being viewed by others
Abbreviations
- FF:
-
Force field
- QM:
-
Quantum mechanical
- TFD:
-
Torsion fingerprint deviation
- RMSD:
-
Root-mean-square deviation
- MMFF:
-
Merck molecular force field
- GAFF:
-
General AMBER force field
- SMIRNOFF:
-
SMIRKS native open force field
References
Bayly C, McKay D, Truchon J (2010) An Informal AMBER Small Molecule Force Field: Parm@ Frosst. http://www.ccl.net/cca/data/parm_at_Frosst/
Chodera J, Qiu Y, Boothroyd S, Wang L-P, Mobley D (2019) The Open Force Field 1.0 small molecule force field, our first optimized force field (codename “Parsley”)
Dauber-Osguthorpe P, Hagler AT (2018) Biomolecular force fields: where have we been, where are we now, where do we need to go and how do we get there? J Comput Aided Mol Des. https://doi.org/10.1007/s10822-018-0111-4
Eastman P, Swails J, Chodera JD, McGibbon RT, Zhao Y, Beauchamp KA, Wang L-P, Simmonett AC, Harrigan MP, Stern CD, Wiewiora RP, Brooks BR, Pande VS (2017) OpenMM 7: rapid development of high performance algorithms for molecular dynamics. PLOS Comput Biol 13(7):e1005659
eMolecules (2015) eMolecules Database Download. https://www.emolecules.com/info/plus/download-database
Fennell CJ, Wymer KL, Mobley DL (2014) A fixed-charge model for alcohol polarization in the condensed phase, and its role in small molecule hydration. J Phys Chem B 118(24):6438–6446 Publisher: American Chemical Society
Hagler AT (2018) Force field development phase II: relaxation of physics-based criteria... or inclusion of more rigorous physics into the representation of molecular energetics. J Comput Aided Mol Des 33:205–264
Haider N (2020) Checkmol/Matchmol Homepage. http://merian.pch.univie.ac.at/~nhaider/cheminf/cmmm
Haider N (2010) Functionality pattern matching as an efficient complementary structure/reaction search tool: an open-source approach. Molecules 15(8):5079–5092
Halgren TA (1992) The representation of van der Waals (vdW) interactions in molecular mechanics force fields: Potential form, combination rules, and vdW parameters. J Am Chem Soc 114(20):7827–7843
Halgren TA (1996a) Merck molecular force field. I. Basis, form, scope, parameterization, and performance of MMFF94. J Comput Chem 17(5–6):490–519
Halgren TA (1996b) Merck molecular force field. II. MMFF94 van der Waals and electrostatic parameters for intermolecular interactions. J Comput Chem 17(5–6):520–552
Halgren TA (1996c) Merck molecular force field. III. Molecular geometries and vibrational frequencies for MMFF94. J Comput Chem 17(5–6):553–586
Halgren TA (1996d) Merck molecular force field. V. Extension of MMFF94 using experimental data, additional computational data, and empirical rules. J Comput Chem 17(5–6):616–641
Halgren TA (1999) MMFF VI. MMFF94s option for energy minimization studies. J Comput Chem 20(7):720–729
Halgren TA, Nachbar RB (1996) Merck molecular force field. IV. conformational energies and geometries for MMFF94. J Comput Chem 17(5–6):587–615
Harder E, Damm W, Maple J, Wu C, Reboul M, Xiang JY, Wang L, Lupyan D, Dahlgren MK, Knight JL, Kaus JW, Cerutti DS, Krilov G, Jorgensen WL, Abel R, Friesner RA (2016) OPLS3: a force field providing broad coverage of drug-like small molecules and proteins. J Chem Theory Comput 12(1):281–296
Hawkins PCD, Skillman AG, Warren GL, Ellingson BA, Stahl MT (2010) Conformer generation with OMEGA: algorithm and validation using high quality structures from the protein databank and Cambridge structural database. J Chem Inf Model 50(4):572–584
Jakalian A, Bush BL, Jack DB, Bayly CI (2000) Fast, efficient generation of high-quality atomic charges. AM1-BCC model: I. Method. J Comput Chem 21(2):132–146
Jakalian A, Jack DB, Bayly CI (2002) Fast, efficient generation of high-quality atomic charges. AM1-BCC model: II. Parameterization and validation. J Comput Chem 23(16):1623–1641
Jang H (2020) Update on Parsley minor releases (openff-1.1.0, 1.2.0)
Jang H, Maat J, Qiu Y, Smith DG, Boothroyd S, Wagner J, Bannan CC, Gokey T, Lim VT, Lucas X, Tjanaka B, Shirts MR, Gilson MK, Chodera JD, Bayly CI, Mobley DL, Wang L-P (2020) Openforcefield/openforcefields: Version 1.2.0 “Parsley” Update. Zenodo
Jorgensen WL, Chandrasekhar J, Madura JD, Impey RW, Klein ML (1983) Comparison of simple potential functions for simulating liquid water. J Chem Phys 79(2):926–935
Lim VT, Hahn DF, Tresadern G, Bayly CI, Mobley D (2020) Benchmark assessment of molecular geometries and energies from small molecule force fields. chemRxiv
Maat J (2020) Training dataset selection
Mobley DL, Bannan CC, Rizzi A, Bayly CI, Chodera JD, Lim VT, Lim NM, Beauchamp KA, Shirts MR, Gilson MK, Eastman PK (2018a) Open Force Field Consortium: escaping atom types using direct chemical perception with SMIRNOFF v0.1. bioRxiv, p. 286542
Mobley DL, Bannan CC, Rizzi A, Bayly CI, Chodera JD, Lim VT, Lim NM, Beauchamp KA, Slochower DR, Shirts MR, Gilson MK, Eastman PK (2018b) Escaping atom types in force fields using direct chemical perception. J Chem Theory Comput 14:11
Nash SG, Nocedal J (1991) A numerical study of the limited memory BFGS method and the truncated-Newton method for large scale optimization. SIAM J Optim 1(3):358–372
Qiu Y, Smith DGA, Boothroyd S, Wagner J, Bannan CC,Gokey T, Jang H, Lim VT, Stern CD, Rizzi A, Lucas X,Tjanaka B, Shirts MR, Gilson MK, Chodera JD, BaylyCI, Mobley DL, Wang L-P (2019) Introducing the firstoptimized Open Force Field 1.0.0 (codename ”Parsley”)
Roos K, Wu C, Damm W, Reboul M, Stevenson JM, Lu C, Dahlgren MK, Mondal S, Chen W, Wang L, Abel R, Friesner RA, Harder ED (2019) OPLS3e: extending force field coverage for drug-like small molecules. J Chem Theory Comput 15(3):1863–1874
Schulz-Gasch T, Schärfer C, Guba W, Rarey M (2012a) TFD: Torsion Fingerprints as a new measure to compare small molecule conformations. J Chem Inf Model 52(6):1499–1512
Schulz-Gasch T, Schärfer C, Guba W, Rarey M (2012b) TFD: Torsion Fingerprints as a new measure to compare small molecule conformations. J Chem Inf Model 52(6):1499–1512
Sellers BD, James NC, Gobbi A (2017) A comparison of quantum and molecular mechanical methods to estimate strain energy in drug like fragments. J Chem Inf Model 57(6):1265–1275
Smith DGA, Altarawy D, Burns LA, Welborn M, Naden LN, Ward L, Ellis S, Pritchard BP, Crawford TD (2020) The MolSSI QCArchive project: an open-source platform to compute, organize, and share quantum chemistry data. WIREs Comput Mol Sci. https://doi.org/10.1002/wcms.1491
ToolKit Szybki (2015) Version 1.9.0. OpenEye Scientific Software Inc., Santa Fe
Vanommeslaeghe K, MacKerell AD (2012) Automation of the CHARMM General Force Field (CGenFF) I: bond perception and atom typing. J Chem Inf Model 52(12):3144–3154
Vanommeslaeghe K, Raman EP, MacKerell AD (2012) Automation of the CHARMM General Force Field (CGenFF) II: assignment of bonded parameters and partial atomic charges. J Chem Inf Model 52(12):3155–3168
Wagner J (2020) Openforcefield/openforcefields: Version 1.1.0 “Parsley” Update. Zenodo
Wang J (2017) A snapshot of GAFF2 development
Wang J, Wolf RM, Caldwell JW, Kollman PA, Case DA (2004) Development and testing of a general amber force field. J Comput Chem 25(9):1157–1174
Wang J, Wang W, Kollman PA, Case DA (2006) Automatic atom type and bond type perception in molecular mechanical calculations. J Mol Graph Model 25(2):247–260
Wlodek S, Skillman A, Nicholls A (2010) Ligand entropy in gas-phase, upon solvation and protein complexation. fast estimation with quasi-Newton Hessian. J Chem Theory Comput 6(7):2140–2152
Acknowledgements
JNE appreciates financial support from the National Institute of Health (R01GM108889). VTL appreciates funding from the National Science Foundation Graduate Research Fellowship Program. CCB appreciates financial support from The Molecular Sciences Software Institute under NSF Grant ACI-154758. DLM appreciates financial support from the National Institutes of Health (R01GM108889 and R01GM132386) and the National Science Foundation (CHE 1352608). The contents of this paper are solely the responsibility of the authors and do not necessarily represent the official views of the NIH.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Code and data availability
We provide the code used in this project in our GitHub repository (https://github.com/mobleylab/off-ffcompare and with a DOI at https://dx.doi.org/10.5281/zenodo.3995606). Additionally, at https://dx.doi.org/10.5281/zenodo.3995059 we provide a supporting data package. This includes a .csv file which has TanimotoCombo and TFD scores, SMILES strings, and eMolecules identifiers for all 2,698,456 molecules analyzed. Additionally, we provide optimized geometries of 265,847 molecules with four or more difference flags. An archived copy of the GitHub repository is provided in the electronic Supporting Information associated with this paper.
Disclaimers
The content is solely the responsibility of the authors and does not necessarily represent the official views of the National Institutes of Health.
Disclosures
DLM is a member of the Scientific Advisory Board of OpenEye Scientific Software and an Open Science Fellow with Silicon Therapeutics.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
About this article
Cite this article
Ehrman, J.N., Lim, V.T., Bannan, C.C. et al. Improving small molecule force fields by identifying and characterizing small molecules with inconsistent parameters. J Comput Aided Mol Des 35, 271–284 (2021). https://doi.org/10.1007/s10822-020-00367-1
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10822-020-00367-1