Abstract
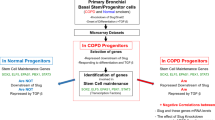
Slug/Snail2 belongs to the Epithelial-Mesenchymal Transition (EMT)-inducing transcription factors involved in development and diseases. Slug is expressed in adult stem/progenitor cells of several epithelia, making it unique among these transcription factors. To investigate Slug role in human bronchial epithelium progenitors, we studied primary bronchial basal/progenitor cells in an air-liquid interface culture system that allows regenerating a bronchial epithelium. To identify Slug downstream genes we knocked down Slug in basal/progenitor cells from normal subjects and subjects with COPD, a respiratory disease presenting anomalies in the bronchial epithelium and high levels of TGF-β in the lungs. We show that normal and COPD bronchial basal/progenitors, even when treated with TGF-β, express both epithelial and mesenchymal markers, and that the epithelial marker E-cadherin is not a target of Slug and, moreover, positively correlates with Slug. We reveal that Slug downstream genes responding to both differentiation and TGF-β are different in normal and COPD progenitors, with in particular a set of proliferation-related genes that are among the genes repressed downstream of Slug in normal but not COPD. In COPD progenitors at the onset of differentiation in presence of TGF-β,we show that there is positive correlations between the effect of differentiation and TGF-β on proliferation-related genes and on Slug protein, and that their expression levels are higher than in normal cells. As well, the expression of Smad3 and β-Catenin, two molecules from TGF-βsignaling pathways, are higher in COPD progenitors, and our results indicate that proliferation-related genes and Slug protein are increased by different TGF-β-induced mechanisms.
Graphical abstract







Similar content being viewed by others
Data Availability
The data analyzed in this publication have been deposited in the NCBI’s Gene.
Expression Omnibus (GEO) and will be accessible through GEO Series Accession.
Number: GSE122957 and GSE123129.
References
Nieto, M. A., Huang, R. Y., Jackson, R. A., & Thiery, J. P. (2016). Emt: 2016. Cell, 166(1), 21–45.
Nassour, M., Idoux-Gillet, Y., Selmi, A., Côme, C., Faraldo, M. L. M., Deugnier, M. A., & Savagner, P. (2012). Slug controls stem/progenitor cell growth dynamics during mammary gland morphogenesis. PLoS One, 7(12), e53498.
Mistry, D. S., Chen, Y., Wang, Y., Zhang, K., & Sen, G. L. (2014). SNAI2 controls the undifferentiated state of human epidermal progenitor cells. Stem Cells, 32(12), 3209–3218.
Tesei, A., Zoli, W., Arienti, C., Storci, G., Granato, A. M., Pasquinelli, G., Valente, S., Orrico, C., Rosetti, M., Vannini, I., Dubini, A., Dell’Amore, D., Amadori, D., & Bonafè, M. (2009). Isolation of stem/progenitor cells from normal lung tissue of adult humans. Cell Proliferation, 42(3), 298–308.
Hackett, N. R., Shaykhiev, R., Walters, M. S., Wang, R., Zwick, R. K., Ferris, B., Witover, B., Salit, J., & Crystal, R. G. (2011). The human airway epithelial basal cell transcriptome. PLoS One, 6(5), e18378.
Rock, J. R., Onaitis, M. W., Rawlins, E. L., Lu, Y., Clark, C. P., Xue, Y., Randell, S. H., & Hogan, B. L. M. (2009). Basal cells as stem cells of the mouse trachea and human airway epithelium. Proceedings of the National Academy of Sciences of the United States of America, 106(31), 12771–12775.
Rock, J. R., Randell, S. H., & Hogan, B. L. (2010). Airway basal stem cells: A perspective on their roles in epithelial homeostasis and remodeling. Disease Models & Mechanisms, 3(9–10), 545–556.
Fulcher, M. L., Gabriel, S., Burns, K. A., Yankaskas, J. R., & Randell, S. H. (2005). Well-differentiated human airway epithelial cell cultures. Methods in Molecular Medicine, 107, 183–206.
Gohy, S. T., Hupin, C., Fregimilicka, C., Detry, B. R., Bouzin, C., Gaide Chevronay, H., Lecocq, M., Weynand, B., Ladjemi, M. Z., Pierreux, C. E., Birembaut, P., Polette, M., & Pilette, C. (2015). Imprinting of the COPD airway epithelium for dedifferentiation and mesenchymal transition. The European Respiratory Journal, 45(5), 1258–1272.
Mertens, T. C. J., Karmouty-Quintana, H., Taube, C., & Hiemstra, P. S. (2017). Use of airway epithelial cell culture to unravel the pathogenesis and study treatment in obstructive airway diseases. Pulmonary Pharmacology & Therapeutics, 45, 101–113.
Rigden, H. M., Alias, A., Havelock, T., O'Donnell, R., Djukanovic, R., Davies, D. E., & Wilson, S. J. (2016). Squamous metaplasia is increased in the bronchial epithelium of smokers with chronic obstructive pulmonary disease. PLoS One, 11(5), e0156009.
Sohal, S. S., & Walters, E. H. (2013). Role of epithelial mesenchymal transition (EMT) in chronic obstructive pulmonary disease (COPD). Respiratory Research, 14, 120.
Steiling, K., van den Berge, M., Hijazi, K., Florido, R., Campbell, J., Liu, G., Xiao, J., Zhang, X., Duclos, G., Drizik, E., Si, H., Perdomo, C., Dumont, C., Coxson, H. O., Alekseyev, Y. O., Sin, D., Pare, P., Hogg, J. C., McWilliams, A., Hiemstra, P. S., Sterk, P. J., Timens, W., Chang, J. T., Sebastiani, P., O’Connor, G. T., Bild, A. H., Postma, D. S., Lam, S., Spira, A., & Lenburg, M. E. (2013). A dynamic bronchial airway gene expression signature of chronic obstructive pulmonary disease and lung function impairment. American Journal of Respiratory and Critical Care Medicine, 187(9), 933–942.
Randell, S. H. (2006). Airway epithelial stem cells and the pathophysiology of chronic obstructive pulmonary disease. Proceedings of the American Thoracic Society, 3(8), 718–725.
Powell, H. A., Iyen-Omofoman, B., Baldwin, D. R., Hubbard, R. B., & Tata, L. J. (2013). Chronic obstructive pulmonary disease and risk of lung cancer: The importance of smoking and timing of diagnosis. Journal of Thoracic Oncology, 8(1), 6–11.
Young, R. P., Hopkins, R. J., Christmas, T., Black, P. N., Metcalf, P., & Gamble, G. D. (2009). COPD prevalence is increased in lung cancer, independent of age, sex and smoking history. The European Respiratory Journal, 34(2), 380–386.
Mahmood, M. Q., Sohal, S. S., Shukla, S. D., Ward, C., Hardikar, A., Noor, W. D., Muller, H. K., Knight, D. A., & Walters, E. H. (2015). Epithelial mesenchymal transition in smokers: Large versus small airways and relation to airflow obstruction. International Journal of Chronic Obstructive Pulmonary Disease, 10, 1515–1524.
Mahmood, M. Q., Reid, D., Ward, C., Muller, H. K., Knight, D. A., Sohal, S. S., & Walters, E. H. (2017). Transforming growth factor (TGF) beta1 and Smad signalling pathways: A likely key to EMT-associated COPD pathogenesis. Respirology, 22(1), 133–140.
Slabakova, E., Pernicova, Z., Slavickova, E., Starsichova, A., Kozubik, A., & Soucek, K. (2011). TGF-beta1-induced EMT of non-transformed prostate hyperplasia cells is characterized by early induction of SNAI2/Slug. Prostate, 71(12), 1332–1343.
Xu, J., Lamouille, S., & Derynck, R. (2009). TGF-beta-induced epithelial to mesenchymal transition. Cell Research, 19(2), 156–172.
Yumoto, K., Thomas, P. S., Lane, J., Matsuzaki, K., Inagaki, M., Ninomiya-Tsuji, J., Scott, G. J., Ray, M. K., Ishii, M., Maxson, R., Mishina, Y., & Kaartinen, V. (2013). TGF-beta-activated kinase 1 (Tak1) mediates agonist-induced Smad activation and linker region phosphorylation in embryonic craniofacial neural crest-derived cells. The Journal of Biological Chemistry, 288(19), 13467–13480.
Bryant, D. M., Datta, A., Rodriguez-Fraticelli, A. E., Peranen, J., Martin-Belmonte, F., & Mostov, K. E. (2010). A molecular network for de novo generation of the apical surface and lumen. Nature Cell Biology, 12(11), 1035–1045.
Leroy, P., & Mostov, K. E. (2007). Slug is required for cell survival during partial epithelial-mesenchymal transition of HGF-induced tubulogenesis. Molecular Biology of the Cell, 18(5), 1943–1952.
Troncale, S., Barbet, A., Coulibaly, L., Henry, E., He, B., Barillot, E., Dubois, T., Hupé, P., & de Koning, L. (2012). NormaCurve: A SuperCurve-based method that simultaneously quantifies and normalizes reverse phase protein array data. PLoS One, 7(6), e38686.
Maubant, S., Tahtouh, T., Brisson, A., Maire, V., Némati, F., Tesson, B., Ye, M., Rigaill, G., Noizet, M., Dumont, A., Gentien, D., Marty-Prouvost, B., de Koning, L., Mahmood, S. F., Decaudin, D., Cruzalegui, F., Tucker, G. C., Roman-Roman, S., & Dubois, T. (2018). LRP5 regulates the expression of STK40, a new potential target in triple-negative breast cancers. Oncotarget, 9(32), 22586–22604.
de la Grange, P., Dutertre, M., Correa, M., & Auboeuf, D. (2007). A new advance in alternative splicing databases: From catalogue to detailed analysis of regulation of expression and function of human alternative splicing variants. BMC Bioinformatics, 8, 180.
de la Grange, P., Dutertre, M., Martin, N., & Auboeuf, D. (2005). FAST DB: A website resource for the study of the expression regulation of human gene products. Nucleic Acids Research, 33(13), 4276–4284.
de la Grange, P., Gratadou, L., Delord, M., Dutertre, M., & Auboeuf, D. (2010). Splicing factor and exon profiling across human tissues. Nucleic Acids Research, 38(9), 2825–2838.
Kanehisa, M., Goto, S., Sato, Y., Furumichi, M., & Tanabe, M. (2012). KEGG for integration and interpretation of large-scale molecular data sets. Nucleic Acids Research, 40(Database issue), D109–D114.
Haw, R., & Stein, L. (2012). Using the reactome database. Current Protocols in Bioinformatics. Chapter 8:Unit8.7. https://doi.org/10.1002/0471250953.bi0807s38.
da Huang, W., Sherman, B. T., & Lempicki, R. A. (2009). Systematic and integrative analysis of large gene lists using DAVID bioinformatics resources. Nature Protocols, 4(1), 44–57.
Akbani, R., Becker, K. F., Carragher, N., Goldstein, T., de Koning, L., Korf, U., Liotta, L., Mills, G. B., Nishizuka, S. S., Pawlak, M., Petricoin III, E. F., Pollard, H. B., Serrels, B., & Zhu, J. (2014). Realizing the promise of reverse phase protein arrays for clinical, translational, and basic research: A workshop report: The RPPA (reverse phase protein Array) society. Molecular & Cellular Proteomics, 13(7), 1625–1643.
Coussens, M. J., Corman, C., Fischer, A. L., Sago, J., & Swarthout, J. (2011). MISSION LentiPlex pooled shRNA library screening in mammalian cells. Journal of Visualized Experiments, 58.
Bolos, V., Peinado, H., Perez-Moreno, M. A., Fraga, M. F., Esteller, M., & Cano, A. (2003). The transcription factor Slug represses E-cadherin expression and induces epithelial to mesenchymal transitions: A comparison with snail and E47 repressors. Journal of Cell Science, 116(Pt 3), 499–511.
Savagner, P. (2001). Leaving the neighborhood: Molecular mechanisms involved during epithelial-mesenchymal transition. Bioessays, 23(10), 912–923.
Jurikova, M., Danihel, L., Polak, S., & Varga, I. (2016). Ki67, PCNA, and MCM proteins: Markers of proliferation in the diagnosis of breast cancer. Acta Histochemica, 118(5), 544–552.
Logan, C. Y., & Nusse, R. (2004). The Wnt signaling pathway in development and disease. Annual Review of Cell and Developmental Biology, 20, 781–810.
Jolly, M. K., Tripathi, S. C., Jia, D., Mooney, S. M., Celiktas, M., Hanash, S. M., Mani, S. A., Pienta, K. J., Ben-Jacob, E., & Levine, H. (2016). Stability of the hybrid epithelial/mesenchymal phenotype. Oncotarget, 7(19), 27067–27084.
Ye, G. D., Sun, G. B., Jiao, P., Chen, C., Liu, Q. F., Huang, X. L., Zhang, R., Cai, W. Y., Li, S. N., Wu, J. F., Liu, Y. J., Wu, R. S., Xie, Y. Y., Chan, E. C., Liou, Y. C., & Li, B. A. (2016). OVOL2, an inhibitor of WNT signaling, reduces invasive activities of human and mouse Cancer cells and is Down-regulated in human colorectal tumors. Gastroenterology, 150(3), 659–671 e616.
Sterneck, E., Poria, D. K., & Balamurugan, K. (2020). Slug and E-cadherin: Stealth accomplices? Frontiers in Molecular Biosciences, 7, 138.
Mahmood, M. Q., Walters, E. H., Shukla, S. D., et al. (2017). Beta-catenin, twist and snail: Transcriptional regulation of EMT in smokers and COPD, and relation to airflow obstruction. Scientific Reports, 7(1), 10832.
Sohal, S. S., Reid, D., Soltani, A., Ward, C., Weston, S., Muller, H. K., Wood-Baker, R., & Walters, E. H. (2011). Evaluation of epithelial mesenchymal transition in patients with chronic obstructive pulmonary disease. Respiratory Research, 12, 130.
Sun, Y., Shao, L., Bai, H., Wang, Z. Z., & Wu, W. S. (2010). Slug deficiency enhances self-renewal of hematopoietic stem cells during hematopoietic regeneration. Blood, 115(9), 1709–1717.
Turner, F. E., Broad, S., Khanim, F. L., Jeanes, A., Talma, S., Hughes, S., Tselepis, C., & Hotchin, N. A. (2006). Slug regulates integrin expression and cell proliferation in human epidermal keratinocytes. The Journal of Biological Chemistry, 281(30), 21321–21331.
Wang, W. L., Huang, H. C., Kao, S. H., Hsu, Y. C., Wang, Y. T., Li, K. C., Chen, Y. J., Yu, S. L., Wang, S. P., Hsiao, T. H., Yang, P. C., & Hong, T. M. (2015). Slug is temporally regulated by cyclin E in cell cycle and controls genome stability. Oncogene, 34(9), 1116–1125.
Bhat-Nakshatri, P., Appaiah, H., Ballas, C., Pick-Franke, P., Goulet Jr., R., Badve, S., Srour, E. F., & Nakshatri, H. (2010). SLUG/SNAI2 and tumor necrosis factor generate breast cells with CD44+/CD24- phenotype. BMC Cancer, 10, 411.
Phillips, S., Prat, A., Sedic, M., Proia, T., Wronski, A., Mazumdar, S., Skibinski, A., Shirley, S. H., Perou, C. M., Gill, G., Gupta, P. B., & Kuperwasser, C. (2014). Cell-state transitions regulated by SLUG are critical for tissue regeneration and tumor initiation. Stem Cell Reports, 2(5), 633–647.
Guo, X., Ramirez, A., Waddell, D. S., Li, Z., Liu, X., & Wang, X. F. (2008). Axin and GSK3- control Smad3 protein stability and modulate TGF - signaling. Genes & Development, 22(1), 106–120.
Kim, J. Y., Kim, Y. M., Yang, C. H., Cho, S. K., Lee, J. W., & Cho, M. (2012). Functional regulation of Slug/Snail2 is dependant on GSK-3b-mediated phosphorylation. The FEBS Journal, 279(16), 2929–2939.
Liu, C., Li, Y., Semenov, M., Han, C., Baeg, G. H., Tan, Y., Zhang, Z., Lin, X., & He, X. (2002). Control of beta-catenin phosphorylation/degradation by a dual mechanism. Cell, 108(6), 837–847.
Derynck, R., Akhurst, R. J., & Balmain, A. (2001). TGF-beta signaling in tumor suppression and cancer progression. Nature Genetics, 29(2), 117–129.
Fynan, T. M., & Reiss, M. (1993). Resistance to inhibition of cell growth by transforming growth factor-beta and its role in oncogenesis. Critical Reviews in Oncogenesis, 4(5), 493–540.
Acknowledgements
We thank the Pulmonary Department, the Pathology Department and the Thoracic Surgery Department at Bichat-Claude Bernard University Hospital (Paris, France) and INSERM UMR 1152 for providing lung tissues and isolating the cells. We thank Audrey Rapinat and David Gentien at the Genomics platform, Curie Institute (Paris, France) for Affymetrix GeneChip hybridization. We thank Dusko Ilic and Pierre Savagner for critical reading of the manuscript and helpful discussions.
Funding
PL is supported by the French National Center for Scientific Research (CNRS). This work was supported by a donation from Association Science et Technologie (Groupe Servier) to PL and by funding from French National Institute for Medical Research (INSERM).
Author information
Authors and Affiliations
Contributions
Study conception and design: PL; Conceived and Designed Experiments CBB, PL.
Conducted experiments, analyzed and interpreted data/: CBB, CC, A J, BO, AC, PdlG, LdK, PL; Writing - draft preparation, review and editing: PdlG, LdK, PL; Supervision: PdlG, LdK, PL; Funding acquisition: PL.
Corresponding author
Ethics declarations
Competing Interests
The authors declare that they have no competing interests.
Ethical Approval
This study was performed in line with the principles of the Declaration of Helsinki. Approval was obtained from the ethics committee of Paris Nord, IRB 00006477 Paris 7 University, France.
Consent to Participate
Informed consent was obtained from all individual participants included in the study.
Consent to Publish
Not applicable
Additional information
Publisher’s Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Fig. S1
Bronchial epithelial progenitors express Slug in their nuclei. Primary bronchial epithelial basal cells from normal (a) or COPD (b) subjects were grown on filters and analyzed undifferentiated at confluence by fluorescent immunocytochemistry. Cells were fixed and labeled simultaneously with progenitor cell marker p63 (white) and Slug (green) antibodies and with Hoechst as a marker of nuclei (blue). Bars are 20 μm (PPTX 1836 kb)
Fig. S2
Bronchial epithelial progenitors co-express epithelial and mesenchymal markers. Primary bronchial epithelial basal cells, normal and COPD, were grown on filters at confluence and either analyzed undifferentiated (a) or changed to ALI culture to induce differentiation, without TGF-β or in presence of 1 ng/ml of TGF-β and analyzed at day 6 of ALI culture for mRNA expression (b). RNA were extracted from normal and COPD cells and analyzed by RT-qPCR to determine the mRNA levels of KRT5 gene, an epithelial cytoskeletal marker and ACTA 2 and Vim genes, mesenchymal cytoskeletal markers.. GAPDH was used to normalize cDNA amounts between samples and results were calculated as a ratio on GAPDH. Data are for n ≥ 11 and experiments were done at least in duplicate. (a) Results shown are log2 (ratio on GAPDH) and are presented as a scatter plot with the mean ±SD. ns: non significant. (b) Results are presented as the fold-change induced by TGF-β on mRNA expression with mean ±SEM, and compare normal and COPD cells. Statistical significance is at P value < 1.00E-03 *** as indicated, ns is non significan (PPTX 129 kb)
Fig. S3
Slug and E-cad/CDH1 mRNA levels in normal and COPD bronchial epithelial progenitors in presence of TGF-β. Primary bronchial epithelial basal cells, normal and COPD, were grown on filters and at confluence changed to ALI culture to induce differentiation in presence of 1 ng/ml of TGF-β. Cells were analyzed at day 6 of ALI culture for Slug (a) and E-cad/CDH1 (b) mRNA expression. RNA were extracted from normal and COPD cells and analyzed by RT-qPCR. GAPDH was used to normalize cDNA amounts between samples and results were calculated as a ratio on GAPDH. Data shown represent the mean for n ≥ 11, and experiments were done at least in duplicate. Statistical significance is at P value < 5.00E-02. ns: non significant (PPTX 87 kb)
Fig. S4
Bronchial epithelial progenitors express Slug in their nuclei and co-express E-cadherin with or without TGF-β. Normal (a, b) or COPD (c, d) primary bronchial epithelial basal cells were grown on filters and at confluence changed to ALI culture to induce differentiation, without TGF-β (a, c) or in presence of 1 ng/ml of TGF-β (b, d). Cells were analyzed at day 6 of ALI culture by fluorescent immunocytochemistry. Cells were fixed and labeled simultaneously with Slug (green) and E-cad (red) antibodies and with Hoechst as a marker of nuclei (blue). Bars are 20 μm (PPTX 3593 kb)
Fig. S5
Effect of Slug knockdown on Slug mRNA and protein levels in normal and COPD bronchial epithelial progenitors. Normal and COPD bronchial epithelial basal/progenitor cells were transduced in transwell inserts with shRNA lentiviral particles corresponding to either a SNAI2/Slug specific sequence or control non-targeting sequences. At day 4 post-transduction, cells were changed to ALI conditions and at day 6 post-transduction cells were analyzed for mRNA or for protein expression. RNA or proteins lysates were prepared from normal and COPD cells and analyzed respectively by RT-qPCR (a) and by Western blot (b). For RT-qPCR analysis, GAPDH was used to normalize cDNA amounts between samples and results were calculated as a ratio on GAPDH. Results are presented as the fold-change induced by shRNA with SNAI2/Slug specific sequence on Slug mRNA expression (a) or Slug protein expression (b) with mean ±SEM. Data are for n ≥ 4. Statistical significance is at P value < 5.00E-02 *, < 1.00E-02 ** or < 1.00E-03 *** as indicated (PPTX 119 kb)
Fig. S6
Comparison of TGF-β effect on the expression of proliferation-related genes between normal and COPD bronchial epithelial progenitors. Primary bronchial epithelial basal cells, normal and COPD, were grown on filters and analyzed undifferentiated at confluence or grown on filters and at confluence changed to ALI culture to induce differentiation, without TGF-β or in presence of 1 ng/ml of TGF-β. Cells were analyzed at day 6 of ALI culture for mRNA expression. RNA were prepared from normal and COPD cells and analyzed respectively by RT-qPCR. GAPDH was used to normalize cDNA amounts between samples and results were calculated as a ratio on GAPDH. Data shown are for n ≥ 11. Results are presented as the fold-change induced by TGF-β on mRNA expression with mean ±SEM. Statistical significance is at P value < 1.00E-03 *** as indicated (PPTX 270 kb)
Table S1
(DOCX 286 kb)
ESM 1
(DOCX 130 kb)
Rights and permissions
About this article
Cite this article
Ben Brahim, C., Courageux, C., Jolly, A. et al. Proliferation Genes Repressed by TGF-β Are Downstream of Slug/Snail2 in Normal Bronchial Epithelial Progenitors and Are Deregulated in COPD. Stem Cell Rev and Rep 17, 703–718 (2021). https://doi.org/10.1007/s12015-021-10123-z
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s12015-021-10123-z