Abstract
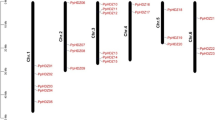
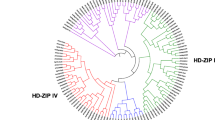
The Homeodomain leucine zipper (HD-Zip) transcription factor family is one of the most important transcription factor families in the plant kingdom and it plays pivotal roles in plant development, growth as well as response to all kinds of stresses. However, the HD-Zip gene family has not been characterized in pineapple yet. Pineapple is one of the most economically important tropical fruit in the world. Here, we identified a total of 33 HD-Zip genes in pineapple by a genome-wide study. These 33 genes were divided into 4 subgroups and variably located on 14 chromosomes and one scaffold_1724. The gene structure and conserved motifs of these genes relatively share similar characteristics in a subfamily. We found at least two abiotic stress responding cis-elements in the promoter region of AcoHDzip genes, the light-responsive element is the most abundant cis-element that can be found in all the genes. Besides, transcriptome analysis showed the expression profile of 33 genes in different developmental stages. We identified the nine genes with higher expression in gynoecium and two genes with highly expressed in ovules and flowers respectively. Our studies will provide valuable information about the HD-Zip transcription factor family in pineapple, which would be useful for further functional studies in plant growth, plant development, and stress response.






Similar content being viewed by others
Abbreviations
- ACO:
-
Ananas comosus
- AT:
-
Arabidopsis thaliana
- OS:
-
Oryza sativa
- HD-Zip:
-
Homeodomain leucine zipper
- TFs:
-
Transcription Factors
References
Adamantia, Agalou, Sigit, Purwantomo, Elin, Övernäs, Henrik, Johannesson, Xiaoyi, Zhu et al (2008) A genome-wide survey of HD-Zip genes in rice and analysis of drought-responsive family members. Plant Molecular Biology
Agalou A, Purwantomo S, Vern SE, Johannesson H, Zhu X, Estiati A, Kam RJD, Engstr M P, Slamet-Loedin IH, Zhu Z, J PMB et al (2008a) A genome-wide survey of HD-Zip genes in rice and analysis of drought-responsive family members. 66 (1–2):87–103
Agalou A, Purwantomo S, Övernäs E, Johannesson H, Zhu X, Estiati A, de Kam RJ, Engström P, Slamet-Loedin IH, Zhu Z, Wang M, Xiong L, Meijer AH, Ouwerkerk PBF et al (2008b) A genome-wide survey of HD-Zip genes in rice and analysis of drought-responsive family members. Plant Mol Biol 66(1):87–103. https://doi.org/10.1007/s11103-007-9255-7
Aoyama T, Dong CH, Wu Y, Carabelli M, Sessa G, Ruberti I, Morelli G, Chua NH et al (1995) Ectopic expression of the Arabidopsis transcriptional activator Athb-1 alters leaf cell fate in tobacco. Plant Cell 7(11):1773–1785. https://doi.org/10.1105/tpc.7.11.1773
Ariel FD, Manavella PA, Dezar CA, Chan RL et al (2007a) The true story of the HD-Zip family. Trends Plant Sci 12(9):419–426. https://doi.org/10.1016/j.tplants.2007.08.003
Ariel FD, Manavella PA, Dezar CA, Chan RL et al (2007b) The true story of the HD-Zip family. Trends in Plant ence 12(9):419–426
Bailey TL, Boden M, Buske FA, Frith M, Grant CE, Clementi L, Ren J, Li WW, Noble WS et al (2009) MEME SUITE: tools for motif discovery and searching. Nucleic acids research 37 (Web Server issue):W202–208. https://doi.org/10.1093/nar/gkp335
Baima S, Nobili F, Sessa G, Lucchetti S, Morelli G et al (1995) The expression of the Athb-8 homeobox gene is restricted to provascular cells in Arabidopsis thaliana. Development 121(12):4171–4182
Baima S, Possenti M, Matteucci A, Wisman E, Morelli G et al (2001) The Arabidopsis ATHB-8 HD-Zip Protein Acts as a Differentiation-Promoting Transcription Factor of the Vascular Meristems. Plant Physiol 126(2):643–655
Barter RL, Yu B (2018) Superheat: An R package for creating beautiful and extendable heatmaps for visualizing complex data. Journal of computational and graphical statistics : a joint publication of American Statistical Association, Institute of Mathematical Statistics, Interface Foundation of North America 27(4):910–922. https://doi.org/10.1080/10618600.2018.1473780
Berardini TZ, Reiser L, Li D, Mezheritsky Y, Muller R, Strait E, Huala E et al (2015) The Arabidopsis information resource: Making and mining the "gold standard" annotated reference plant genome. Genesis (New York, NY : 2000) 53 (8):474–485. https://doi.org/10.1002/dvg.22877
Chen H, Hu B, Zhao L, Shi D, Qin Y et al (2019) Differential Expression Analysis of Reference Genes in Pineapple (Ananas comosus L.) during Reproductive Development and Response to Abiotic Stress, Hormonal Stimuli. Tropical Plant Biology
Dengler KN (2002) Cell cycling frequency and expression of the homeobox gene ATHB-8 during leaf vein development in Arabidopsis. Planta 216(2):212–219
Edgar RC (2004) MUSCLE: Multiple Sequence Alignment with High Accuracy and High Throughput. Nucleic Acids Res 32(5):1792–1797
Elisabeth G, Alexandre G, Christine H, Ivan I, Appel RD, Amos B et al (2003) ExPASy: the proteomics server for in-depth protein knowledge and analysis. Nucleic Acids Res 13:13
Gould KS, Lamotte O, Klinguer A, Pugin A, Wendehenne D et al (2010) Nitric oxide production in tobacco leaf cells: a generalized stress response? Plant, Cell Environ 26(11):1851–1862
Guo AY, Zhu QH, Chen X, Luo JC et al (2007) GSDS: A gene structure display server. Hereditas 29(8):1023–1026
Henriksson E (2005) Homeodomain Leucine Zipper Class I Genes in Arabidopsis. Expression Patterns and Phylogenetic Relationships. Plant Physiology 139 (1):509–518
Huey RB (2002) Plants Versus Animals: Do They Deal with Stress in Different Ways? Integrative & Comparative Biology
Ivica L, Peer B (2017) 20 years of the SMART protein domain annotation resource. Nucleic Acids Res D1:D1
Javelle, Marie, Klein-Cosson, Catherine, Vernoud, Vanessa, Boltz, Véronique, Maher, Chris et al (2011) Genome-Wide Characterization of the HD-ZIP IV Transcription Factor Family in Maize: Preferential Expression in the Epidermis. Plant Physiology
John C, Harris, Maria, Hrmova, Sergiy, Lopato, Peter, Langridge et al (2011) Modulation of plant growth by HD-Zip class I and II transcription factors in response to environmental stimuli. New Phytologist
Kawahara R, Komamine A, Fukuda H et al (1995) Isolation and characterization of homeobox-containing genes of carrot. Plant Mol Biol 27(1):155–164
Lam-Tung N, Schmidt HA, Arndt VH, Quang MB et al (2014) IQ-TREE: A Fast and Effective Stochastic Algorithm for Estimating Maximum-Likelihood Phylogenies. Mol Biol Evol 1:1
Li Z, Zhang C, Guo Y, Niu W, Wang Y, Xu Y et al (2017) Evolution and expression analysis reveal the potential role of the HD-Zip gene family in regulation of embryo abortion in grapes (Vitis vinifera L.). Bmc Genomics 18 (1):1–16
Lulu W, Yi L, Xingyue J, Liping L, Xiaozhuan D, Yanhui L, Lihua Z, Ping Z, Xiaomei W, Yeqiang L, Lin D, QIn Y et al (2020) Floral transcriptomes reveal gene networks in pineapple floral growth and fruit development. Communications Biology Sep 10;3(1):500.3(1):500. https://doi.org/10.1038/s42003-020-01235-2
Ming R (2016) Sequencing of the Pineapple Genome as a Model CAM species. Revista Do Instituto De Pesquisas E Estudos 72(5):687–693
Morelli G, Ruberti I (2002) Light and shade in the photocontrol of Arabidopsis growth. Trends Plant Sci 7(9):399–404
Nakamura M, Katsumata H, Abe M, Yabe N, Komeda Y, Yamamoto KT, Takahashi T et al (2006a) Characterization of the Class IV Homeodomain-Leucine Zipper Gene Family in Arabidopsis. Plant Physiol 141(4):1363–1375
Nakamura M, Katsumata H, Abe M, Yabe N, Takahashi T et al (2006b) Characterization of the class IV homeodomain-leucine zipper (HD-Zip IV) gene family in Arabidopsis. Plant Physiol 141(4):1363–1375
Ohashi Y, Oka A, Rodrigues-Pousada R, Possenti M, Ruberti I, Morelli G, Aoyama T et al (2003) Modulation of Phospholipid Signaling by GLABRA2 in Root-Hair Pattern Formation. Ence 300(5624):1427–1430
Olsson A, Engstrm P, Sderman E et al (2004a) The homeobox genes ATHB12 and ATHB7 encode potential regulators of growth in response to water deficit in Arabidopsis. Plant Mol Biol 55(5):663–677
Olsson A, Engstrm P, Sderman E et al (2004b) The homeobox genes ATHB12 and ATHB7encode potential regulators of growth in response to water deficit in Arabidopsis. Plant Mol Biol 55(5):663–677
Otsuga D, Deguzman B, Prigge MJ, Drews GN, Clark SE et al (2010) REVOLUTA regulates meristem initiation at lateral positions. Plant Journal 25(2):223–236
Prigge MJ, Clark SEJE, Development et al (2010) Evolution of the class III HD-Zip gene family in land plants. 8 (4):350–361
Priyadarshani SVGN, Cai H, Zhou Q, Liu Y, Cheng Y, Xiong J, Patson D, Zhao H, Qin Y et al (2019) An Efficient Agrobacterium Mediated Transformation of Pineapple with GFP-Tagged Protein Allows Easy, Non-Destructive Screening of Transgenic Pineapple Plants. Biomolecules 9. https://doi.org/10.3390/biom9100617
Ré DA, Capella M, Bonaventure G, Chan RL et al (2014) Arabidopsis AtHB7 and AtHB12 evolved divergently to fine tune processes associated with growth and responses to water stress. Bmc Plant Biology
Roy S (2015) Function of MYB domain transcription factors in abiotic stress and epigenetic control of stress response in plant genome. Plant Signaling & Behavior 11 (1):-
Sara E-G, Jaina M, Alex B, R ES, Aurélien L, C PS, Matloob Q, J RL, A SG, Alfredo S, et al (2018) The Pfam protein families database in 2019. Nucleic Acids Res D 1:D1
Sawa S, Ohgishi M, Goda H, Higuchi K, Shimada Y et al (2002) The HAT2 gene, a member of the HD-Zip gene family, isolated as an auxin inducible gene by DNA microarray screening, affects auxin response in Arabidopsis. Plant Journal 32(6):1011–1022
Schena M, Davis RW (1992) HD-Zip proteins: members of an Arabidopsis homeodomain protein superfamily. Proceedings of the National Academy of Ences of the United States of America 89(9):3894–3898
Shen W, Li H, Teng R, Wang Y, Wang W, Zhuang J et al (2018) Genomic and transcriptomic analyses of HD-Zip family transcription factors and their responses to abiotic stress in tea plant (Camellia sinensis). Genomics 111:S0888754318302623-
Schrick K, Bruno M, Khosla A, Cox PN, Marlatt SA, Roque RA, Nguyen HC, He C, Snyder MP, Singh D et al (2014) Shared functions of plant and mammalian StAR-related lipid transfer (START) domains in modulating transcription factor activity. BMC Biol 12(1):70
Son O, Cho HY, Kim MR, Lee H, Lee MS, Song E, Park JH, Nam KH, Chun JY, Kim HJ et al (2004) Induction of a homeodomain-leucine zipper gene by auxin is inhibited by cytokinin in Arabidopsis roots. Biochem Biophys Res Commun 326(1):203–209
Vollbrecht E, Veit B, Sinha N, Hake S et al (1991) The developmental gene Knotted-1 is a member of a maize homeobox gene family. Nature 350(6315):241–243
Voorrips RE (2002) MapChart: Software for the Graphical Presentation of Linkage Maps and QTLs. J Hered 1:1
Wang Y, Henriksson E, S?Derman E, Henriksson KN, Sundberg E, Engstr?M PJDB et al (2004) The Arabidopsis homeobox gene, ATHB16, regulates leaf development and the sensitivity to photoperiod in Arabidopsis. 264 (1):228–239
Wellik DM (2011) Hox genes and kidney development. Pediatric Nephrology 26(9):1559–1565
Xu H, Yu Q, Shi Y, Hua X, Tang H, Yang L, Ming R, Zhang J et al (2018) PGD: Pineapple Genomics Database. Horticulture Research 5(1):66. https://doi.org/10.1038/s41438-018-0078-2
Yang JY (2002) OSTF1: A HD-GL2 Family Homeobox Gene is Developmentally Regulated During Early Embryogenesis in Rice. Plant Cell Physiol 43(6):628–638
Zhang F, Zhang F, Huang L, Cruz CV, Ali J, Xu J, Zhou Y, Li Z et al (2016) Overlap between Signaling Pathways Responsive to Xanthomonas oryzae pv. oryzae Infection and Drought Stress in Rice Introgression Line Revealed by RNA-Seq. Journal of Plant Growth Regulation 35 (2):345–356
Zhang XM, Yao WJ, Zhao K, Jiang TB, Zhou BR et al (2017) Bioinformatics and Response to Salt Stress Analysis of the HD-Zip Transcription Factor Family in ×. Bulletin of Botanical Research
Funding
This work was supported by NSFC (U1605212; 31970333) and a Guangxi Distinguished Experts Fellowship to Y.Q.
Author information
Authors and Affiliations
Contributions
Q.Z. and L.L. performed the experiment, and manuscript writing; H.C. performed the experiment, data analyse;Z.l., M.H.W., T.L. helped to design the figure; H.Z. and S. V. G. N. P. revised and edited the manuscript; Q.Y. Fund acquisition and revising manuscript.
Corresponding author
Ethics declarations
Conflict of Interest
The authors declare no conflict of interest.
Additional information
Communicated by: Ray Ming.
Publisher’s Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
12042_2020_9279_MOESM1_ESM.docx
Supplementary file1 Table S1. Characterization of HD-Zip gene superfamily in pineapple. Table S2. The primers of the selected genes of HD-Zip family for RT-qPCR (DOCX 19 KB)
Rights and permissions
About this article
Cite this article
Zhou, Q., Liu, L., Cheng, H. et al. Genome-wide Identification and Expression Pattern Analysis of the HD-Zip Transcription Factor Family in Pineapple (Ananas Comosus). Tropical Plant Biol. 14, 120–131 (2021). https://doi.org/10.1007/s12042-020-09279-8
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s12042-020-09279-8