Abstract
Purpose
Integrins, transmembrane receptors that mediate cell-extracellular matrix and cell-cell interactions, have been linked to several cancer-associated features. A less explored function of integrins in cancer is their role in leukocyte homing and activation. Understanding their relationship with immune cell infiltrates and immune checkpoints is an area of interest in cancer research.
Methods
The expression of 33 different integrins was evaluated in relation with breast cancer patient outcome using transcriptomic data (Affymetrix dataset, exploratory cohort) and the METABRIC study (validation cohort). The TIMER online tool was used to assess the association of the identified integrin genes with immune cell infiltration, and the TCGA and METABRIC studies to assess correlations between integrin gene expression and genomic signatures of immune activation.
Results
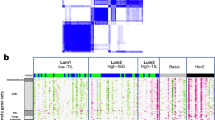
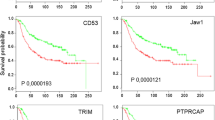
We identified 7 genes coding for integrin α and β subunits, i.e., ITGA4, ITGB2, ITGAX, ITGB7, ITGAM, ITGAL and ITGA8, which predict a favorable prognosis in Basal-like and HER2+ breast cancers. Their expression positively correlated with the presence of immune cell infiltrates within the tumor (dendritic cells, CD4+ T-cells, neutrophils, CD8+ T-cells and B-cells), with markers of T-cell activation and antigen presentation, and with gene signatures of immune surveillance (cytotoxic T lymphocyte activation and IFN gamma signature). By contrast, we found that genes coding for integrins that predicted a detrimental outcome (IBSP, ITGB3BP, ITGB6, ITGB1 and ITGAV) were not associated with any of these parameters.
Conclusions
We identified an integrin signature composed of 7 genes with potential to recognize immune infiltrated and activated Basal-like and HER2+ breast cancers with a favorable prognosis.







Similar content being viewed by others
Data availability
Data are available upon reasonable request.
Abbreviations
- APC:
-
antigen presenting cell
- CI:
-
confidence interval
- CTL:
-
cytotoxic T lymphocyte
- DC:
-
dendritic cells
- ECM:
-
extracellular matrix
- FDR:
-
false discovery rate
- HR:
-
hazard ratio
- ICAM:
-
intercellular adhesion molecule
- IFN:
-
Interferon
- KM:
-
Kaplan-Meier
- LAD:
-
leukocyte adhesion deficiency
- LFA:
-
leukocyte function-associated antigen 1
- NK:
-
natural killer
- OS:
-
overall survival
- P:
-
p-value
- PD-1:
-
programmed cell death protein-1
- PD-L1:
-
programmed cell death protein ligand-1
- RFS:
-
relapse-free survival
- TGF-β:
-
transforming growth factor β
- TIMER:
-
Tumor Immune Estimation Resource
- VCAM-1:
-
vascular intercellular adhesion molecule 1
- VLA-4:
-
very late antigen 4
References
H. Hamidi, J. Ivaska, Every step of the way: integrins in cancer progression and metastasis. Nat. Rev. Cancer 18, 533–548 (2018)
S. Cabodi, P. Di Stefano, M. del Pilar Camacho Leal, A. Tinnirello, B. Bisaro, V. Morello, L. Damiano, S. Aramu, D. Repetto, G. Tornillo, P. Defilippi, Integrins and signal transduction. Adv. Exp. Med. Biol. 674, 43–54 (2010)
B. Luo, C.V. Carman, T.A. Springer, Structural basis of integrin regulation and signaling. Annu. Rev. Immunol. 25, 619–647 (2007)
Alday-Parejo, Stupp, Rüegg, Are integrins still practicable targets for anti-cancer therapy? Cancers 11, 978 (2019)
J.S. Desgrosellier, D.A. Cheresh, Integrins in cancer: biological implications and therapeutic opportunities. Nat. Rev. Cancer 10, 9–22 (2010)
G. Sökeland, U. Schumacher, The functional role of integrins during intra- and extravasation within the metastatic cascade. Mol. Cancer 18, 12 (2019)
L. Seguin, J.S. Desgrosellier, S.M. Weis, D.A. Cheresh, Integrins and cancer: regulators of cancer stemness, metastasis, and drug resistance. Trends Cell Biol. 25, 234–240 (2015)
Brown, Marshall, Integrin-mediated TGFβ activation modulates the tumour microenvironment. Cancers 11, 1221 (2019)
H. Harjunpää, M. Llort Asens, C. Guenther, S.C. Fagerholm, Cell adhesion molecules and their roles and regulation in the immune and tumor microenvironment. Front. Immunol. 10, 1078 (2019)
C.L. Abram, C. A. Lowell, The ins and outs of leukocyte integrin signaling. Annu. Rev. Immunol. 27, 339–362 (2009)
Y. Zhang, H. Wang, Integrin signalling and function in immune cells. Immunology 135, 268–275 (2012)
N. Harbeck, F. Penault-Llorca, J. Cortes, M. Gnant, N. Houssami, P. Poortmans, K. Ruddy, J. Tsang, F. Cardoso, Breast cancer. Nat. Rev. Dis. Prim. 5, 66 (2019)
G. Stelzer, N. Rosen, I. Plaschkes, S. Zimmerman, M. Twik, S. Fishilevich, T.I. Stein, R. Nudel, I. Lieder, Y. Mazor, S. Kaplan, D. Dahary, D. Warshawsky, Y. Guan-Golan, A. Kohn, N. Rappaport, M. Safran, D. Lancet, The genecards suite: from gene data mining to disease genome sequence analyses. Curr. Protoc. Bioinform. 54, 1.30.1–1.30.33 (2016)
GeneCards – the human gene database [Internet]. [cited 15 Jun 2020]. Available: http://www.genecards.org/
E.A. Bruford, B. Braschi, P. Denny, T.E.M. Jones, R.L. Seal, S. Tweedie, Guidelines for human gene nomenclature. Nat. Genet. 52, 754–758 (2020)
HUGO Gene Nomenclature Committee webpage [Internet]. [cited 15 Jun 2020]. Available: https://www.genenames.org
Z. Mihály, B. Győrffy, Improving pathological assessment of breast cancer by employing array-based transcriptome analysis. Microarrays 2, 228–242 (2013)
B. Györffy, A. Lanczky, A.C. Eklund, C. Denkert, J. Budczies, Q. Li, Z. Szallasi, An online survival analysis tool to rapidly assess the effect of 22,277 genes on breast cancer prognosis using microarray data of 1,809 patients. Breast Cancer Res. Treat. 123, 725–731 (2010)
Kaplan-Meier Plotter [Internet]. [cited 15 Jun 2020]. Available: https://kmplot.com
P.-K. Raj-Kumar, J. Liu, J. A. Hooke, A. J. Kovatich, L. Kvecher, C. D. Shriver, H. Hu, PCA-PAM50 improves consistency between breast cancer intrinsic and clinical subtyping reclassifying a subset of luminal A tumors as luminal B. Sci. Rep. 9, 7956 (2019)
A.K. Falck, A. Röme, M. Fernö, H. Olsson, G. Chebil, P.O. Bendahl, L. Rydén, St Gallen molecular subtypes in screening-detected and symptomatic breast cancer in a prospective cohort with long-term follow-up. Br. J. Surg. 103, 513–523 (2016)
T. Li, J. Fan, B. Wang, N. Traugh, Q. Chen, J.S. Liu, B. Li, X.S. Liu, A web server for comprehensive analysis of tumor-infiltrating immune cells. Cancer Res. 77, e108–e110 (2017)
TIMER: Tumor IMmune Estimation Resource [Internet]. [cited 18 Jun 2020]. Available: http://timer.cistrome.org/
M. Ayers, J. Lunceford, M. Nebozhyn, E. Murphy, A. Loboda, D.R. Kaufman, A. Albright, J.D. Cheng, S.P. Kang, V. Shankaran, S.A. Piha-Paul, J. Yearley, T.Y. Seiwert, A. Ribas, T.K. McClanahan, IFN-γ–related mRNA profile predicts clinical response to PD-1 blockade. J. Clin. Invest. 127, 2930–2940 (2017)
M.S. Rooney, S.A. Shukla, C.J. Wu, G. Getz, Hacohen, molecular and genetic properties of tumors associated with local immune cytolytic activity. Cell 160, 48–61 (2015)
P. Jiang, S. Gu, D. Pan, J. Fu, A. Sahu, X. Hu, Z. Li, N. Traugh, X. Bu, B. Li, J. Liu, G.J. Freeman, M.A. Brown, K.W. Wucherpfennig, X.S. Liu, Signatures of T cell dysfunction and exclusion predict cancer immunotherapy response. Nat. Med. 24, 1550–1558 (2018)
B.D. Lehmann, J.A. Bauer, X. Chen, M.E. Sanders, A.B. Chakravarthy, Y. Shyr, J.A. Pietenpol, Identification of human triple-negative breast cancer subtypes and preclinical models for selection of targeted therapies. J. Clin. Invest. 121, 2750–2767 (2011)
A.R. Cortazar, V. Torrano, N. Martín-Martín, A. Caro-Maldonado, L. Camacho, I. Hermanova, E. Guruceaga, L.F. Lorenzo-Martín, R. Caloto, R.R. Gomis, I. Apaolaza, V. Quesada, J. Trka, A. Gomez-Muñoz, S. Vincent, X.R. Bustelo, F.J. Planes, A.M. Aransay, A. Carracedo, CANCERTOOL: A visualization and representation interface to exploit cancer datasets. Cancer Res. 78, 6320–6328 (2018)
CANCERTOOL [Internet]. [cited 18 Jun 2020]. Available: http://web.bioinformatics.cicbiogune.es/CANCERTOOL/index.html
A. Alcaraz-Sanabria, M. Baliu-Piqué, C. Saiz-Ladera, K. Rojas, A. Manzano, G. Marquina, A. Casado, F.J. Cimas, P. Pérez-Segura, A. Pandiella, B. Gyorffy, A. Ocana, Genomic signatures of immune activation predict outcome in advanced stages of ovarian cancer and basal-like breast tumors. Front. Oncol. 9, 1–10 (2020)
J. Pérez-Pena, J. Tibor Fekete, R. Páez, M. Baliu-Piqué, J. García-Saenz, V. García-Barberán, A. Manzano, P. Pérez-Segura, A. Esparis-Ogando, A. Pandiella, B. Gyorffy, A. Ocana, A transcriptomic immunologic signature predicts favorable outcome in neoadjuvant chemotherapy treated triple negative breast tumors. Front. Immunol. 10, 1–9 (2019)
B.D. Lehmann, B. Jovanović, X. Chen, M.V. Estrada, K.N. Johnson, Y. Shyr, H.L. Moses, M.E. Sanders, J.A. Pietenpol, Refinement of triple-negative breast cancer molecular subtypes: implications for neoadjuvant chemotherapy selection. PLoS One 11, e0157368 (2016)
A. Ribas, J.D. Wolchok, Cancer immunotherapy using checkpoint blockade. Science 359, 1350–1355 (2018)
S. Gettinger, L. Horn, D. Jackman, D. Spigel, S. Antonia, M. Hellmann, J. Powderly, R. Heist, L.V. Sequist, D.C. Smith, P. Leming, W.J. Geese, D. Yoon, A. Li, J. Brahmer, Five-year follow-up of nivolumab in previously treated advanced non–small-cell lung cancer: Results from the CA209-003 study. J. Clin. Oncol. 36, 1675–1684 (2018)
P. Schmid, S. Adams, H.S. Rugo, A. Schneeweiss, C.H. Barrios, H. Iwata, V. Dieras, R. Hegg, S.A. Im, G. Shaw, V. Wright, L. Henschel, S.Y. Molinero, R. Chui, A. Funke, E.P. Husain, S. Winer, Loi, L.A. Emens, Atezolizumab and nab-paclitaxel in advanced triple-negative breast cancer. N. Engl. J. Med. 379, 2108–2121 (2018)
P. Bonaventura, T. Shekarian, V. Alcazer, J. Valladeau-Guilemond, S. Valsesia-Wittmann, S. Amigorena, C. Caux, S. Depil, Cold tumors: a therapeutic challenge for immunotherapy. Front. Immunol. 10, 1–10 (2019)
G.T. Gibney, L.M. Weiner, M.B. Atkins, Predictive biomarkers for checkpoint inhibitor-based immunotherapy. Lancet Oncol. 17, e542–e551 (2016)
A. Ocana, A. Pandiella, Targeting oncogenic vulnerabilities in triple negative breast cancer: Biological bases and ongoing clinical studies. Oncotarget 8, 22218–22234 (2017)
A. Marra, G. Viale, G. Curigliano, Recent advances in triple negative breast cancer: The immunotherapy era. BMC Med. 17, 1–9 (2019)
Planes-Laine, Rochigneux, Bertucci, Chrétien, Viens, Sabatier, Gonçalves, PD-1/PD-L1 Targeting in breast cancer: the first clinical evidences are emerging. A literature review. Cancers 11, 1033 (2019)
R. Sackstein, T. Schatton, S.R. Barthel, T-lymphocyte homing: an underappreciated yet critical hurdle for successful cancer immunotherapy. Lab. Invest. 97, 669–697 (2017)
P. Alcaide, S. Auerbach, F.W. Luscinskas, Neutrophil recruitment under shear flow: it’s all about endothelial cell rings and gaps. Microcirculation 16, 43–57 (2009)
B.L. Walling, M. Kim, LFA-1 in T cell migration and differentiation. Front. Immunol. 9, 952 (2018)
T.K. Kishimoto, N. Hollander, T.M. Roberts, D.C. Anderson, T.A. Springer, Heterogeneous mutations in the β subunit common to the LFA-1, Mac-1, and p150,95 glycoproteins cause leukocyte adhesion deficiency. Cell 50, 193–202 (1987)
M. Baiula, S. Spampinato, L. Gentilucci, A. Tolomelli, Novel ligands targeting α4β1 integrin: therapeutic applications and perspectives. Front. Chem. 7, 1–12 (2019)
M. Vicente-Manzanares, F. Sánchez-Madrid, Targeting the integrin interactome in human disease. Curr. Opin. Cell Biol. 55, 17–23 (2018)
M.M. Morrison, Leukocyte beta2-integrins; Genes and disease. J. Genet. Syndr. Gene Ther. 4, 154 (2013)
I. Dotan, M. Allez, S. Danese, M. Keir, S. Tole, J. McBride, The role of integrins in the pathogenesis of inflammatory bowel disease: Approved and investigational anti-integrin therapies. Med. Res. Rev. 40, 245–262 (2020)
J.Z. Kechagia, J. Ivaska, P. Roca-Cusachs, Integrins as biomechanical sensors of the microenvironment. Nat. Rev. Mol. Cell Biol. 20, 457–473 (2019)
K.M. Moore, G.J. Thomas, S.W. Duffy, J. Warwick, R. Gabe, P. Chou, I.O. Ellis, A.R. Green, S. Haider, K. Brouilette, A. Saha, S. Vallath, R. Bowen, C. Chelala, D. Eccles, W.J. Tapper, A.M. Thompson, P. Quinlan, L. Jordan, C. Gillett, A. Brentnall, S. Violette, P.H. Weinreb, J. Kendrew, S.T. Barry, I.R. Hart, J.L. Jones, J.F. Marshall, Therapeutic targeting of integrin αvβ6 in breast cancer. J. Natl. Cancer Inst. 106, dju169 (2014)
S. Giampieri, C. Manning, S. Hooper, L. Jones, C.S. Hill, E. Sahai, Localized and reversible TGFβ signalling switches breast cancer cells from cohesive to single cell motility. Nat. Cell Biol. 11, 1287–1296 (2009)
M.D. Allen, G.J. Thomas, S. Clark, M.M. Dawoud, S. Vallath, S.J. Payne, J.J. Gomm, S.A. Dreger, S. Dickinson, D.R. Edwards, C.J. Pennington, I. Sestak, J. Cuzick, J.F. Marshall, I.R. Hart, J.L. Jones, Altered microenvironment promotes progression of preinvasive breast cancer: myoepithelial expression of v 6 integrin in DCIS Identifies high-risk patients and predicts recurrence. Clin. Cancer Res. 20, 344–357 (2014)
Funding
This work was supported by the Instituto de Salud Carlos III (PI19/00808), ACEPAIN, Diputación de Albacete, CIBERONC and CRIS Cancer Foundation (to A. Ocaña) and the Ministry of Economy and Competitiveness of Spain (BFU2015-71371-R), the Instituto de Salud Carlos III through the Spanish Cancer Centers Network Program (RD12/0036/0003) and CIBERONC, the scientific foundation of the AECC and the CRIS Foundation (to A. Pandiella). Our laboratories receive support from the European Community through the regional development funding program (FEDER). BG was supported by a 2018 − 2.1.17-TET-KR-00001 grant and by the Higher Education Institutional Excellence Programme (2020 − 4.1.1.-TKP2020) of the Ministry for Innovation and Technology of Hungary, within the framework of the Bionic thematic programme of the Semmelweis University.
Author information
Authors and Affiliations
Contributions
K.R., M.B.P. and A.O. conceived and designed the study. K.R., M.B.P., A.M., C.S.L and B.G. searched the data and performed the analyses. M.B.P., A.O., K.R., A.M., C.S.L, V.G.B, F.C., P.P.S, A.P. and B.G wrote the manuscript. All authors reviewed, included modifications and approved the final version of the manuscript.
Corresponding author
Ethics declarations
Conflict of interest
A.O. receives funding from Entrechem, travel expenses from Merck and advisory board fees from Daiichi Sankyo. P.P.S. receives funding from Merck and MSD. A.P. receives consultancy fees from Daiichi Sankyo.
Ethics approval and consent to participate
This study does not involve human participants or experimental animal models.
Consent for publication
This manuscript does not contain personal and/or medical information on identifiable living individuals.
Additional information
Publisher’s Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
ESM 1
(DOCX 2.36 MB)
Rights and permissions
About this article
Cite this article
Rojas, K., Baliu-Piqué, M., Manzano, A. et al. In silico transcriptomic mapping of integrins and immune activation in Basal-like and HER2+ breast cancer. Cell Oncol. 44, 569–580 (2021). https://doi.org/10.1007/s13402-020-00583-9
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s13402-020-00583-9