Abstract
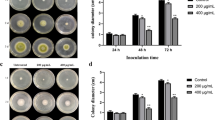
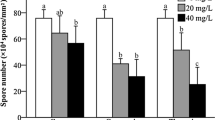
Antimicrobial peptides (AMPs) are biologically active molecules that can eradicate bacteria by destroying the bacterial membrane structure, causing the bacteria to rupture. However, little is known about the extent and effect of AMPs on filamentous fungi. In this study, we synthesized small molecular polypeptides by an inexpensive heat conjugation approach and examined their effects on the growth of Aspergillus flavus and its secondary metabolism. The antimicrobial agents significantly inhibited aflatoxin production, conidiation, and sclerotia formation in A. flavus. Furthermore, we found that the expression of aflatoxin structural genes was significantly inhibited, and the intracellular reactive oxygen species (ROS) level was reduced. Additionally, the antimicrobial agents can change membrane permeability. Overall, our results demonstrated that antimicrobial agents, safe to mammalian cells, have an obvious impact on aflatoxin production, which indicated that antimicrobial agents may be adopted as a new generation of potential agents for controlling aflatoxin contamination.









Similar content being viewed by others
Data availability
All data generated or analyzed during this study are included in this manuscript.
Change history
26 April 2021
A Correction to this paper has been published: https://doi.org/10.1007/s42770-021-00501-7
References
Rodrigues I, Handl J, Binder EM (2011) Mycotoxin occurrence in commodities, feeds and feed ingredients sourced in the Middle East and Africa. Food Addit Contam Part B Surveill 4:168–179. https://doi.org/10.1080/19393210.2011.589034
Luz C, Calpe J, Saladino F, Luciano FB, Fernandez-Franzón M, Mañes J, Meca G (2018) Antimicrobial packaging based on ɛ-polylysine bioactive film for the control of mycotoxigenic fungi in vitro and in bread. J Food Process Preserv 42:e13370. https://doi.org/10.1111/jfpp.13370
Liu Y, Chang CCH, Marsh GM, Wu F (2012) Population attributable risk of aflatoxin-related liver cancer: systematic review and meta-analysis. Eur J Cancer 48:2125–2136. https://doi.org/10.1016/j.ejca.2012.02.009
Iimura K, Furukawa T, Yamamoto T, Negishi L, Suzuki M, Sakuda S. The mode of action of cyclo(L-Ala-L-Pro) in inhibiting aflatoxin production of Aspergillus flavus. Toxins (Basel). 2017; 9: E219. doi:https://doi.org/10.3390/toxins9070219
Perlin DS, Rautemaa-Richardson R, Alastruey-Izquierdo A (2017) The global problem of antifungal resistance: prevalence, mechanisms, and management. Lancet Infect Dis 17:e383–e392. https://doi.org/10.1016/S1473-3099(17)30316-X
Rončević T, Vukičević D, Ilić N, Krce L, Gajski G, Tonkić M, Goić-Barišić I, Zoranić L, Sonavane Y, Benincasa M, Juretić D, Maravić A, Tossi A (2018) Antibacterial activity affected by the conformational flexibility in glycine-lysine based α-helical antimicrobial peptides. J Med Chem 61:2924–2936. https://doi.org/10.1021/acs.jmedchem.7b01831
Zasloff M (2002) Antimicrobial peptides of multicellular organisms. Nature. 415:389–395. https://doi.org/10.1038/415389a
Shagaghi N, Palombo EA, Clayton AHA, Bhave M (2018) Antimicrobial peptides: biochemical determinants of activity and biophysical techniques of elucidating their functionality. World J Microbiol Biotechnol 34:62. https://doi.org/10.1007/s11274-018-2444-5
León-Calvijo MA, Leal-Castro AL, Almanzar-Reina GA, Rosas-Pérez JE, García-Castañeda JE, Rivera-Monroy ZJ (2015) Antibacterial activity of synthetic peptides derived from lactoferricin against Escherichia coli ATCC 25922 and Enterococcus Faecalis ATCC 29212. Biomed Res Int 2015:453826–453828. https://doi.org/10.1155/2015/453826
Lai PK, Kaznessis YN (2018) Insights into membrane translocation of protegrin antimicrobial peptides by multistep molecular dynamics simulations. ACS Omega 3:6056–6065. https://doi.org/10.1021/acsomega.8b00483
den Hertog AL, van Marle J, van Veen HA, Van't Hof W, Bolscher JG, Veerman EC, Nieuw Amerongen AV (2005) Candidacidal effects of two antimicrobial peptides: histatin 5 causes small membrane defects, but LL-37 causes massive disruption of the cell membrane. Biochem J 388:689–695. https://doi.org/10.1042/bj20042099
Benincasa M, Scocchi M, Pacor S, Tossi A, Nobili D, Basaglia G, Busetti M, Gennaro R (2006) Fungicidal activity of five cathelicidin peptides against clinically isolated yeasts. J Antimicrob Chemother 58:950–959. https://doi.org/10.1093/jac/dkl382
Nawrot R, Barylski J, Nowicki G, Nowicki G, Broniarczyk J, Buchwald W, Goździcka-Józefiak A (2014) Plant antimicrobial peptides. Folia Microbiol (Praha) 59:181–196. https://doi.org/10.1007/s12223-013-0280-4
Lin CH, Chang MW, Chen CY (2014) A potent antimicrobial peptide derived from the protein LsGRP1 of Lilium. Phytopathology. 104:340–346. https://doi.org/10.1094/phyto-09-13-0252-r
De Lucca AJ, Bland JM, Grimm C, Jacks TJ, Cary JW, Jaynes JM, Cleveland TE, Walsh TJ (1998) Fungicidal properties, sterol binding, and proteolytic resistance of the synthetic peptide D4E1. Can J Microbiol 44:514–520. https://doi.org/10.1139/w98-032
Rajasekaran K, Stromberg KD, Cary JW, Cleveland TE (2001) Broad-spectrum antimicrobial activity in vitro of the synthetic peptide D4E1. J Agric Food Chem 49:2799–2803. https://doi.org/10.1021/jf010154d
DeGray G, Rajasekaran K, Smith F, Sanford J, Daniell H (2001) Expression of an antimicrobial peptide via the chloroplast genome to control phytopathogenic bacteria and fungi. Plant Physiol 127:852–862. https://doi.org/10.1104/pp.010233
Rajasekaran K, Sayler RJ, Sickler CM, Majumdar R, Jaynes JM, Cary JW (2018) Control of Aspergillus flavus growth and aflatoxin production in transgenic maize kernels expressing a tachyplesin-derived synthetic peptide, AGM182. Plant Sci 270:150–156. https://doi.org/10.1016/j.plantsci.2018.02.006
Zeitler B, Herrera Diaz A, Dangel A, Thellmann M, Meyer H, Sattler M, Lindermayr C (2013) De-novo design of antimicrobial peptides for plant protection. PLoS One 8:e71687. https://doi.org/10.1371/journal.pone.0071687
Holaskova E, Galuszka P, Frebort I, Oz MT (2014) Antimicrobial peptide production and plant-based expression systems for medical and agricultural biotechnology. Biotechnol Adv 33:1005–1023. https://doi.org/10.1016/j.biotechadv.2015.03.007
Tang M, Zhou YC, Gao JY, Peng JL, Wang Y, Zhao QR, Liao LH, Wang K, Pan MJ, Xing M, Pan W, Dai DL, Fu M, Yu L, Zhang CQ, Wang YC, Zhang Y, Xu L, Li J, Bao X, Piao WX, Lin SH, Lu KB, Zhang XL, Cao WG, Yang K, He ZM, Weng SP, Liu QY, He JG (2017) Heat conjugation of antibacterial agents from amino acids and plant oil. Sci Rep 7:10852. https://doi.org/10.1038/s41598-017-11451-2
Yeaman MR, Yount NY (2003) Mechanisms of antimicrobial peptide action and resistance. Pharmacol Rev 55:27–55. https://doi.org/10.1124/pr.55.1.2
Jayashree T, Subramanyam C (2000) Oxidative stress as a prerequisite for aflatoxin production by Aspergillus parasiticus. Free Radic Biol Med 29:981–985. https://doi.org/10.1016/S0891-5849(00)00398-1
Reverberi M, Zjalic S, Ricelli A, Punelli F, Camera E, Fabbri C, Picardo M, Fanelli C, Fabbri AA (2008) Modulation of antioxidant defense in Aspergillus parasiticus is involved in aflatoxin biosynthesis: a role for the ApyapA gene. Eukaryot Cell 7:988–1000. https://doi.org/10.1128/EC.00228-07
Grintzalis K, Vernardis SI, Klapa MI, Georgiou CD (2014) Role of oxidative stress in sclerotial differentiation and aflatoxin B1 biosynthesis in Aspergillus flavus. Appl Environ Microbiol 80:5561–5571. https://doi.org/10.1128/AEM.01282-14
Zhao X, Zhi QQ, Li JY, Keller NP, He ZM (2018) The antioxidant gallic acid inhibits aflatoxin formation in Aspergillus flavus by modulating transcription factors Farb and CreA. Toxins (Basel) 10:E270. https://doi.org/10.3390/toxins10070270
Wu MY, Mead ME, Kim SC, Rokas A, Yu JH (2017) WetA bridges cellular and chemical development in Aspergillus flavus. PLoS One 12:e0179571. https://doi.org/10.1371/journal.pone.0179571
Rushing BR, Selim MI (2019) Aflatoxin B1: a review on metabolism, toxicity, occurrence in food, occupational exposure, and detoxification methods. Food Chem Toxicol 124:81–100. https://doi.org/10.1016/j.fct.2018.11.047
Delattin N, De Brucker K, De Cremer K, Cammue BPA, Thevissen K (2017) Antimicrobial peptides as a strategy to combat fungal biofilms. Curr Top Med Chem 17:604–612. https://doi.org/10.2174/1568026616666160713142228
Amare MG, Keller NP (2014) Molecular mechanisms of Aspergillus flavus secondary metabolism and development. Fungal Genet 66:11–18. https://doi.org/10.1016/j.fgb.2014.02.008
Linz JE, Hong SY, Roze LV (2013) Oxidative stress-related transcription factors in the regulation of secondary metabolism. Toxins (Basel) 5:683–702. https://doi.org/10.3390/toxins5040683
Roze LV, Laivenieks M, Hong SY, Wee J, Wong SS, Vanos B, Awad D, Ehrlich KC, Linz JE (2015) Aflatoxin biosynthesis is a novel source of reactive oxygen species—a potential redox signal to initiate resistance to oxidative stress? Toxins (Basel) 7:1411–1430. https://doi.org/10.3390/toxins7051411
Medina A, Schmidt-Heydt M, Rodríguez A, Parra R, Geisen R, Magan N (2015) Impacts of environmental stress on growth, secondary metabolite biosynthetic gene clusters and metabolite production of xerotolerant/xerophilic fungi. Curr Genet 61:325–334. https://doi.org/10.1007/s00294-014-0455-9
Rautenbach M, Troskie AM, Vosloo JA (2016) Antifungal peptides: to be or not to be membrane active. Biochimie. 130:132–145. https://doi.org/10.1016/j.biochi.2016.05.013
Wang KR, Dang W, Xie JQ, Zhu RR, Sun MY, Jia FJ, Zhao YY, An XP, Qiu S, Li XY, Ma ZL, Yan WJ, Wang R (1848) Antimicrobial peptide protonectin disturbs the membrane integrity and induces ROS production in yeast cells. Biochim Biophys Acta Biomembr 2015:2365–2373. https://doi.org/10.1016/j.bbamem.2015.07.008
Kovač K, Šarkanj B, Klapec T, Borišev I, Kovač M, Nevistić A, Strelec I (2018) Antiaflatoxigenic effect of fullerene C60 nanoparticles at environmentally plausible concentrations. AMB Express 8:14. https://doi.org/10.1186/s13568-018-0544-0
Kovač M, Šubarić D, Bulaić M, Kovač T, Šarkanj B (2018) Yesterday masked, today modified; what do mycotoxins bring next? Arh Hig Rada Toksikol 69:196–214. https://doi.org/10.2478/aiht-2018-69-3108
Mohale S, Magan N, Medina A (2013) Comparison of growth, nutritional utilisation patterns, and niche overlap indices of toxigenic and atoxigenic Aspergillus flavus strains. Fungal Biol 117:650–659. https://doi.org/10.1016/j.funbio.2013.07.002
Lee T-H, Hall KN, Aguilar M-I (2015) Antimicrobial peptide structure and mechanism of action: a focus on the role of membrane structure. Curr Top Med Chem 16:25–39. https://doi.org/10.2174/1568026615666150703121700
Amaike S, Keller NP (2011) Aspergillus flavus. Annu Rev Phytopathol 49:107–133. https://doi.org/10.1146/annurev-phyto-072910-095221
Bayram Ö, Braus GH (2012) Coordination of secondary metabolism and development in fungi: the velvet family of regulatory proteins. FEMS Microbiol Rev 36:1–24. https://doi.org/10.1111/j.1574-6976.2011.00285.x
Zhao X, Spraker JE, Bok JW, Velk T, Hel ZM, Keller NP (2017) A cellular fusion cascade regulated by laea is required for sclerotial development in Aspergillus flavus. Front Microbiol 8:1925. https://doi.org/10.3389/fmicb.2017.01925
Liu J, Sun LH, Zhang NY, Zhang JC, Guo J, Li C, Rajput SA, Qi DS (2016) Effects of nutrients in substrates of different grains on Aflatoxin B1 production by Aspergillus flavus. Biomed Res Int 2016:7232858–7232810. https://doi.org/10.1155/2016/7232858
Prosser JI, Trinci APJ (2009) A model for hyphal growth and branching. J Gen Microbiol 111:153–164. https://doi.org/10.1099/00221287-111-1-153
Balmant W, Sugai-Guérios MH, Coradin JH, Krieger N, Furigo A, Mitchell DA (2015) A model for growth of a single fungal hypha based on well-mixed tanks in series: simulation of nutrient and vesicle transport in aerial reproductive hyphae. PLoS One 10:e0120307. https://doi.org/10.1371/journal.pone.0120307
Pontecorvo G, Roper JA, Chemmons LM, Macdonald KD, Bufton AWJ (1953) The genetics of Aspergillus nidulans. Adv Genet 5:141–238. https://doi.org/10.1016/S0065-2660(08)60408-3
Chang P-K, Scharfenstein LL, Mack B, Ehrlich KC (2012) Deletion of the Aspergillus flavus orthologue of A. nidulans fluG reduces conidiation and promotes production of sclerotia but does not abolish aflatoxin biosynthesis. Appl Environ Microbiol 78:7557–7563. https://doi.org/10.1128/aem.01241-12
Lin JQ, Zhao XX, Wang CC, Xie Y, Li GH, He ZM (2013) 5-Azacytidine inhibits aflatoxin biosynthesis in Aspergillus flavus. Ann Microbiol 63:763–769. https://doi.org/10.1007/s13213-012-0531-7
Leiter É, Szappanos H, Oberparleiter C, Kaiserer L, Csernoch L, Pusztahelyi T, Emri T, Pócsi I, Salvenmoser W, Marx F (2005) Antifungal protein PAF severely affects the integrity of the plasma membrane of Aspergillus nidulans and induces an apoptosis-like phenotype. Antimicrob Agents Chemother 49:2445–2453. https://doi.org/10.1128/AAC.49.6.2445-2453.2005
Crespo-Sempere A, Selma-Lázaro C, Palumbo JD, González-Candelas L, Martínez-Culebras PV (2016) Effect of oxidant stressors and phenolic antioxidants on the ochratoxigenic fungus Aspergillus carbonarius. J Sci Food Agric 96:169–177. https://doi.org/10.1002/jsfa.7077
Funding
This work was supported in part by the Science and Technology Planning Project of Guangdong Province, China (Grant No. 2016B020204001), the National Natural Science Foundation of China (Grant Nos. 31870031 and 31470198), and the Opening Fund of Jiangsu Key Laboratory for Food Quality and Safety-State Key Laboratory Cultivation Base (028074911709). This work was also supported by a grant from Guangzhou Science and Technology Program (201804010328) and Science and Technology Transformation Program of Sun Yat-sen University of China (33000-18843234).
Author information
Authors and Affiliations
Contributions
Li J and He ZM conceived and designed the experiment. Li J, Zhi QQ, Zhang J, Yuan XY, Wan YL, and Jia LH performed the experiments and analyzed the data. Li J wrote the manuscript. Zhi QQ, Liu QY, and He ZM revised the manuscript. Shi JR and He ZM provided the funding for the study.
Corresponding authors
Ethics declarations
Conflict of interest
On behalf of all authors, the corresponding author states that there is no conflict of interest.
Ethics approval
Not applicable.
Consent to participate
All the authors listed have approved the manuscript.
Consent for publication
All the authors listed have approved the publication.
Code availability
Not applicable.
Additional information
Responsible Editor: Mariza Landgraf
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
About this article
Cite this article
Li, J., Zhi, QQ., Zhang, J. et al. Synthetic antimicrobial agents inhibit aflatoxin production. Braz J Microbiol 52, 821–835 (2021). https://doi.org/10.1007/s42770-021-00423-4
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s42770-021-00423-4