Abstract
Range-limited endemic taxa are threatened by loss of genomic diversity, which can lead to extirpation and extinction. The imperiled and narrowly distributed edaphic endemic Carex scirpoidea subsp. convoluta (Cyperaceae), which is primarily limited to alvar soils on the northern shores of Lake Huron in North America, exhibits such risk. In contrast, the conspecific C. scirpoidea subsp. scirpoidea is widely distributed across a vast geographical range and occupies various arctic and alpine habitat types. Using ddRADseq, population genomic analyses of 11 North American populations of C. scirpoidea including five subsp. convoluta populations revealed similar-to-higher levels of genomic diversity in the latter compared to its more widely distributed conspecific subsp. scirpoidea. Dioecy and a nearly obligate outcrossing mode of reproduction have likely contributed to the maintenance of genomic diversity in subsp. convoluta, preventing inbreeding depression, and contributing to the evolutionary potential of populations of this taxon. Phylogenomic and Bayesian analyses of ddRADseq data revealed that subsp. convoluta is paraphyletic with Eastern North American subsp. scirpoidea populations, suggesting that it best be considered a locally adapted ecotype. Furthermore, Eastern North America populations of both taxa arose from populations that likely occupied periglacial refugia south of the Laurentide Ice Sheet, where they no longer occur. Herein, we identify conservation units among the sampled Eastern North American C. scirpoidea populations to aid management efforts, which we suggest should include habitat protection and replanting for habitat restoration purposes.






Similar content being viewed by others
Data availability
The datasets generated during and analyzed during the current study are available in the NCBI SRA repository, https://www.ncbi.nlm.nih.gov/bioproject/607130
References
Alexander PJ, Rajanikanth G, Bacon CD, Bailey CD (2007) Recovery of plant DNA using a reciprocating saw and silica-based columns. Mol Ecol Notes 7:5–9. https://doi.org/10.1111/j.1471-8286.2006.01549.x
Allendorf FW, Hohenlohe PA, Luikart G (2010) Genomics and the future of conservation genetics. Nat Rev Genet 11:697–709. https://doi.org/10.1038/nrg2844
Allessio Leck M, Schütz W (2005) Regeneration of Cyperaceae, with particular reference to seed ecology and seed banks. Perspect Plant Ecol Evol Syst 7:95–133. https://doi.org/10.1016/j.ppees.2005.05.001
Andrews S (2010) FastQC a quality-control tool for high-throughput sequence data. http://www.bioinformatics.babraham.ac.uk/projects/fastqc/. Accessed 6 Dec 2020
Baird NA, Etter PD, Atwood TS et al (2008) Rapid SNP discovery and genetic mapping using sequenced RAD markers. PLoS ONE 3:1–7. https://doi.org/10.1371/journal.pone.0003376
Bard NW (2018) Population genomics of a rare edaphic endemic. University of Colorado Denver
Bennett KD (1992) Holocene history of forest trees on the Bruce Peninsula, southern Ontario. Can J Bot 70:6–18. https://doi.org/10.1139/b92-002
Bradshaw AD (1991) The Croonian Lecture, 1991. Genostasis and the limits to evolution. Philos Trans R Soc Lond B 333:289–305. https://doi.org/10.1098/rstb.1991.0079
Brooker RW, Carlsson BÅ, Callaghan TV (2001) Carex bigelowii Torrey ex schweinitz (C. rigida Good., non Schrank; C. hyperborea Drejer). J Ecol 89:1072–1095. https://doi.org/10.1046/j.0022-0477.2001.00623.x
Brownell VR, Riley JR (2000) The alvars of Ontario: significant alvar natural areas in the Ontario Great Lakes region. Federation of Ontario Naturalists, Don Mills
Bruederle LP, Yarbrough SL, Fehlberg SD (2008) Allozyme variation in the genus Carex… 15 years later. In: Naczi RFC, Ford BA (eds) Sedges: uses, diversity, and systematics of the Cyperaceae. Missouri Botanical Garden Press, St Louis, MO, pp 187–196
Bucharova A, Bossdorf O, Hölzel N, Kollmann J, Prasse R, Durka W (2019) Mix and match: regional admixture provenancing strikes a balance among different seed-sourcing strategies for ecological restoration. Conserv Genet 20:7–17. https://doi.org/10.1007/s10592-018-1067-6
Catling PM (1995) The extent of confinement of Vascular Plants to Alvars in Southern Ontario. Can F Nat 109:172–181
Catling PM, Brownell VR (1995) Alvars of the Great Lakes Region. Can F Nat 109:143–171
Chong Z, Ruan J, Wu C (2012) Sequence analysis rainbow : an integrated tool for efficient clustering and assembling RAD-seq reads. Bioinformatics 28:2732–2737. https://doi.org/10.1093/bioinformatics/bts482
Clausen J (1951) Stages in the evolution of plant species. Cornell University Press, Ithaca, NY
Comer PJ, Cuthrell DL, Albert DA, Penskar MR (1997) Natural community abstract for limestone bedrock lakeshore. Michigan Natural Features Inventory, Lansing, MI
Crémieux L, Bischoff A, Müller-Schärer H, Steinger T (2010) Gene flow from foreign provenances into local plant populations: fitness consequences and implications for biodiversity restoration. Am J Bot 97:94–100. https://doi.org/10.3732/ajb.0900103
Danecek P, Auton A, Abecasis G et al (2011) The variant call format and VCFtools. Bioinformatics 27:2156–2158. https://doi.org/10.1093/bioinformatics/btr330
Darriba D, Taboada GL, Doallo R, Posada D (2015) jModelTest 2: more models, new heuristics and high-performance computing. Nat Methods 9:6–9. https://doi.org/10.1038/nmeth.2109
Davis MB, Shaw RG (2001) Range shifts and adaptive responses to quaternary climate change. Science 292:673–679. https://doi.org/10.1126/science.292.5517.673
DePrenger-Levine ME (2007) Genetic diversity in a rare North American endemic Carex scirpoidea ssp. convoluta. University of Colorado Denver
Dillenberger MS, Kadereit JW (2013) The phylogeny of the European high mountain genus Adenostyles (Asteraceae-Senecioneae) reveals that edaphic shifts coincide with dispersal events. Am J Bot 100:1171–1183. https://doi.org/10.3732/ajb.1300060
Dray S, Dufour A-B (2015) The ade4 package: implementing the duality diagram for ecologists. J Stat Softw 22:1–20. https://doi.org/10.18637/jss.v022.i04
Dunlop DA (1990) The biosystematics of Carex section Scirpinae (Cyperaceae). University of New Hampshire
Dunlop DA (2003) Carex Linnaeus sect. Scirpinae (Tuckerman) Kükenthal. In: Flora of North America, vol 23. http://www.efloras.org/florataxon.aspx?flora_xml:id=1&taxon_xml:id=302733
Dunlop DA, Crow GE (1999) The taxonomy of Carex section Scirpinae (Cyperaceae). Rhodora 101:163–199
Dyke AS (2005) Late quaternary vegetation history of northern North America based on pollen, macrofossil, and faunal remains. Geogr Phys Q 59:211–262. https://doi.org/10.7202/014755ar
Eaton DAR (2014) PyRAD: assembly of de novo RADseq loci for phylogenetic analyses. Bioinformatics 30:1844–1849. https://doi.org/10.1093/bioinformatics/btu121
Eaton DAR (2020) iPyrad: interactive assembly and analysis of RADseq data sets. Bioinformatics 36:2592–2594. https://doi.org/10.1093/bioinformatics/btz966
Ehlers J, Gibbard PL, Hughes PD (eds) (2011) Quaternary glaciations—extent and chronology, vol 15. Elsevier, Amsterdam
Ellstrand NC, Elam DR (1993) Population genetic consequences of small population size : implications for plant conservation. Annu Rev Ecol Syst 24:217–242
Evanno G, Regnaut S, Goudet J (2005) Detecting the number of clusters of individuals using the software STRUCTURE: a simulation study. Mol Ecol 14:2611–2620. https://doi.org/10.1111/j.1365-294X.2005.02553.x
Excoffier L, Lischer HEL (2010) Arlequin suite ver 3.5: a new series of programs to perform population genetics analyses under Linux and Windows. Mol Ecol Resour 10:564–567. https://doi.org/10.1111/j.1755-0998.2010.02847.x
Fant JB, Havens K, Keller JM et al (2014) The influence of contemporary and historic landscape features on the genetic structure of the sand dune endemic, Cirsium pitcheri (Asteraceae). Heredity (Edinb) 112:519–530. https://doi.org/10.1038/hdy.2013.134
Fischer J, Lindenmayer DB (2007) Landscape modification and habitat fragmentation: a synthesis. Glob Ecol Biogeogr 16:265–280. https://doi.org/10.1111/j.1466-8238.2006.00287.x
Foll M, Gaggiotti O (2008) A genome-scan method to identify selected loci appropriate for both dominant and codominant markers: a Bayesian perspective. Genetics 180:977–993. https://doi.org/10.1534/genetics.108.092221
Ford BA, Ross Mcqueen DA, Naczi RFC, Reznicek AA (1998) Allozyme variation and genetic relationships among species in the Carex willdenowii complex (Cyperaceae). Am J Bot 85:546–552. https://doi.org/10.2307/2446439
Frankham R (2005) Genetics and extinction. Biol Conserv 126:131–140. https://doi.org/10.1016/j.biocon.2005.05.002
Frankham R (2010) Where are we in conservation genetics and where do we need to go ? Conserv Genet. https://doi.org/10.1007/s10592-009-0010-2
Frankham R, Bradshaw CJA, Brook BW (2014) Genetics in conservation management: revised recommendations for the 50/500 rules, Red List criteria and population viability analyses. Biol Conserv 170:56–63. https://doi.org/10.1016/j.biocon.2013.12.036
Fraser DL, Bernatchez L (2001) Adaptive evolutionary conservation: towards a unified concept for defining conservation units. Mol Ecol 10:2741–2752
Funk WC, McKay JK, Hohenlohe PA, Allendorf FW (2012) Harnessing genomics for delineating conservation units. Trends Ecol Evol 27:489–496. https://doi.org/10.1016/j.tree.2012.05.012
Garrison E (2012) Vcflib: A C++ library for parsing and manipulating VCF files. https://www.https://github.com/vcflib/vcflib. Accessed 6 Dec 2020
Garrison E, Marth G (2012) Haplotype-based variant detection from short-read sequencing. arXiv:1207.3907[q-bio.GN]
GBIF (2018) Global Biodiversity Information Facility. https://www.gbif.org. Accessed 1 June 2018
Gómez-Fernández A, Alcocer I, Matesanz S (2016) Does higher connectivity lead to higher genetic diversity? Effects of habitat fragmentation on genetic variation and population structure in a gypsophile. Conserv Genet 17:631–641. https://doi.org/10.1007/s10592-016-0811-z
Goudet J (2005) hierfstat, a package for r to compute and test hierarchical F-statistics. Mol Ecol Notes 5:184–186. https://doi.org/10.1111/j.1471-8286.2004.00828.x
Guibert C, Civeyrel L, Linder P (2009) Male and female separation event trapped in a species tree. Taxon 58:172–180
Guindon S, Gascuel O (2003) A simple, fast and accurate method to estimate large phylogenies by maximum-likelihood. Syst Biol 52:696–704. https://doi.org/10.1080/10635150390235520
Hamilton JA, Eckert CG (2007) Population genetic consequences of geographic disjunction: a prairie plant isolated on Great Lakes alvars. Mol Ecol 16:1649–1660. https://doi.org/10.1111/j.1365-294X.2007.03241.x
Hamrick AJL, Godt MJW (1996) Effects of life history traits on genetic diversity in plant species. Philos Trans R Soc Lond 351:1291–1298. https://doi.org/10.1098/rstb.1996.0112
Hampe A, Petit R (2005) Conserving biodiversity under climate change: the rear edge matters. Ecol Lett 8:461–467. https://doi.org/10.1111/j.1461-0248.2005.00739.x
Handel SN (1976) Restricted pollen flow of two woodland herbs determined by neutron-activation analysis. Nature 260:422–423. https://doi.org/10.1038/260422a0
Hewitt GM (2000) The genetic legacy of the quarternary ice ages. Nature 405:907–913. https://doi.org/10.1038/35016000
Huson DH, Scornavacca C (2012) Dendroscope 3: an interactive tool for rooted phylogenetic trees and networks. Syst Biol 61:1061–1067. https://doi.org/10.1093/sysbio/sys062
IDigBio (2018) Integrated digitized biocollections portal. http://www.idigbio.org/portal. Accessed 1 June 2018
Jiménez-Mejías P, Fernández-Mazuecos M, Amat ME, Vargas P (2015) Narrow endemics in European mountains: high genetic diversity within the monospecific genus Pseudomisopates (Plantaginaceae) despite isolation since the late Pleistocene. J Biogeogr 42:1455–1468. https://doi.org/10.1111/jbi.12507
Jombart T (2008) adegenet: a R package for the multivariate analysis of genetic markers. Bioinformatics 24:1403–1405. https://doi.org/10.1093/bioinformatics/btn129
Jombart T, Ahmed I (2011) adegenet 1.3-1: new tools for the analysis of genome-wide SNP data. Bioinformatics 27:3070–3071. https://doi.org/10.1093/bioinformatics/btr521
Jost L (2008) GST and its relatives do not measure differentiation. Mol Ecol 17:4015–4026. https://doi.org/10.1111/j.1365-294X.2008.03887.x
Jost L, Archer F, Flanagan S et al (2018) Differentiation measures for conservation genetics. Evol Appl 11:1139–1148. https://doi.org/10.1111/eva.12590
Kozlov AM, Darriba D, Flouri T et al (2019) RAxML-NG: a fast, scalable and user-friendly tool for maximum likelihood phylogenetic inference. Bioinformatics 305:1–3. https://doi.org/10.1093/bioinformatics/btz305
Kruckeberg AR (1951) Intraspecific variability in the response of certain native plant species to serpentine soil. Am J Bot 38:408–419. https://doi.org/10.2307/2438248
Kruckeberg AR, Rabinowitz D (1985) Biological aspects of endemism in higher plants. Annu Rev Ecol Syst 16:447–479. https://doi.org/10.1146/annurev.es.16.110185.002311
Li H (2014) Toward better understanding of artifacts in variant calling from high-coverage samples. Bioinformatics 30:2843–2851. https://doi.org/10.1093/bioinformatics/btu356
Li H, Durbin R (2009) Fast and accurate short read alignment with Burrows-Wheeler transform. Bioinformatics 25:1754–1760. https://doi.org/10.1093/bioinformatics/btp324
Li P, Li M, Shi Y et al (2013) Phylogeography of North American herbaceous Smilax (Smilacaceae): combined AFLP and cpDNA data support a northern refugium in the Driftless Area. Am J Bot 100:801–814. https://doi.org/10.3732/ajb.1200250
Lischer HEL, Excoffier L (2012) PGDSpider: an automated data conversion tool for connecting population genetics and genomics programs. Bioinformatics 28:298–299. https://doi.org/10.1093/bioinformatics/btr642
Lowry DB (2012) Ecotypes and the controversy over stages in the formation of new species. Biol J Linnaean Soc 106:241–257. https://doi.org/10.1111/j.1095-8312.2012.01867.x
Ludwig B, Chadde S, Prout L (1996) Establishment record for Horseshoe Bay: research Natural Area within Hiawatha National Forest Mackinac County, Michigan
Martín-Bravo S, Jiménez-Mejias P, Villaverde T et al (2019) A tale of worldwide success: behind the scenes of Carex (Cyperaceae) biogeography and diversification. J Syst Evol 57:695–718. https://doi.org/10.1111/jse.12549
Meirmans PG, Van Tienderen PH (2004) GENOTYPE and GENODIVE: two programs for the analysis of genetic diversity of asexual organisms. Mol Ecol Notes 4:792–794. https://doi.org/10.1111/j.1471-8286.2004.00770.x
Modesto IS, Miguel C, Pina-Martins F et al (2014) Identifying signatures of natural selection in cork oak (Quercus suber L.) genes through SNP analysis. Tree Genet Genomes 10:1645–1660. https://doi.org/10.1007/s11295-014-0786-1
NatureServe (2019) NatureServe explorer: an online encyclopedia of life. In: NatureServe Arlingt. Virginia
Omernik JM (1995) Ecoregions: a framework for managing ecosystems. George Wright Forum 12:35–50
Parchman TL, Gompert Z, Mudge J et al (2012) Genome-wide association genetics of an adaptive trait in lodgepole pine. Mol Ecol 21:2991–3005. https://doi.org/10.1111/j.1365-294X.2012.05513.x
Pembrook JW (2014) Systematics of Carex section Scirpinae Tuck. (Cyperaceae), with insight to the origin of edaphic endemics. University of Colorado Denver
Peterson BK, Weber JN, Kay EH et al (2012) Double digest RADseq: an inexpensive method for de novo SNP discovery and genotyping in model and non-model species. PLoS ONE 7:1–11. https://doi.org/10.1371/journal.pone.0037135
Pina-Martins F, Silva DN, Fino J, Paulo OS (2017) Structure_threader: an improved method for automation and parallelization of programs structure, fastStructure and MavericK on multicore CPU systems. Mol Ecol Resour 17:e268–e274. https://doi.org/10.1111/1755-0998.12702
Pritchard JK, Stephens M, Donnelly P (2000) Inference of population structure using multilocus genotype data. Genetics 155:945–959
Puritz JB, Hollenbeck CM, Gold JR (2014) dDocent : a RADseq, variant-calling pipeline designed for population genomics of non-model organisms. PeerJ 2:1–14. https://doi.org/10.7717/peerj.431
QGIS (2019) QGIS Geographic Information System. Open Source Geospatial Foundation Project. http://qgis.org
Rajakaruna N (2004) The edaphic factor in the origin of plant species. Int Geol Rev 46:471–478. https://doi.org/10.2747/0020-6814.46.5.471
Rajakaruna N (2018) Lessons on evolution from the study of edaphic specialization. Bot Rev 84:39–78. https://doi.org/10.1007/s12229-017-9193-2
Reschke C, Reid R, Jones J et al (1999) Conserving Great Lakes Alvars. The Nature Conservancy Great Lakes Program, Chicago, IL
Reznicek AA, Voss EG, Walters BS (2011) Michigan Flora Online. University of Michigan. https://michiganflora.net/species.aspx?id=1045. Accessed 29 July 2020
Schaefer CA, Larson DW (1997) Vegetation, environmental characteristics and ideas on the maintenance of alvars on the Bruce Peninsula, Canada. J Veg Sci 8:797–810. https://doi.org/10.2307/3237024
SEINet (2018) SEINet Portal Network. www.swbiodiversity.org/seinet/index.php. Accessed 4 June 2018
Shackleford R (2003) Conservation assessment for Bulrush Sedge (Carex scirpoidea Michx.): subspecies scirpoidea and convoluta (Kükenth.) Dunlop. Milwaukee, Wisconsin
Stark KE, Lundholm JT, Larson DW (2004) Arrested development of soil on alvars of ontario, Canada: Implications for conservation and restoration. Nat Areas J 24:95–100
Stebbins GL, Major J (1965) Endemism and speciation in the California Flora. Ecol Monogr 35:1–35
Szövényi P, Arroyo K, Guggisberg A, Conti E (2009) Effects of Pleistocene glaciations on the genetic structure of Saxifraga florulenta (Saxifragaceae), a rare endemic of the Maritime Alps. Taxon 58:532–543. https://doi.org/10.1002/tax.582017
Tripp EA, Tsai YHE, Zhuang Y, Dexter KG (2017) RADseq dataset with 90% missing data fully resolves recent radiation of Petalidium (Acanthaceae) in the ultra-arid deserts of Namibia. Ecol Evol 7:7920–7936. https://doi.org/10.1002/ece3.3274
USDA; NRCS (2019) The plants database. In: Natl. Plant Data Team. http://plants.usda.gov
Van Auken OW (2000) Shrub invasions of North American semiarid grasslands. Annu Rev Ecol Syst 31:197–215. https://doi.org/10.1146/annurev.ecolsys.31.1.197
Vergiev S (2018) The impact of sea water immersion on the viability of psammophilous species Carex colchica and its capacity as dune stabilizer. Comptes Rendus L’Academie Bulg des Sci 71:648–654. https://doi.org/10.7546/CRABS.2018.05.09
Villaverde T, Escudero M, Martin-Bravo S et al (2017) Bipolar distributions in vascular plants: a review. Am J Bot 104:1680–1694. https://doi.org/10.3732/ajb.1700159
Waples RS (1991) Pacific salmon, Oncorhynchus spp., and the definition of “species” under the Endangered Species Act. Mar Fish Rev 53:11
Westergaard KB, Zemp N, Stenøien HK et al (2019) Population genomic evidence for plant glacial survival in Scandinavia. Mol Ecol 28:818–832. https://doi.org/10.1111/mec.14994
Whitlock MC (2011) G’ST and D do not replace FST. Mol Ecol 20:1083–1091. https://doi.org/10.1111/j.1365-294X.2010.04996.x
Willis SC, Hollenbeck CM, Puritz JB et al (2017) Haplotyping RAD loci: an efficient method to filter paralogs and account for physical linkage. Mol Ecol Resour 17:955–965. https://doi.org/10.1111/1755-0998.12647
Yarbrough SL (2000) Effects of dioecy on population genetic structure in Carex scirpoidea Michaux ssp. scirpoidea. University of Colorado Denver
Yu Z (2003) Late Quaternary dynamics of tundra and forest vegetation in the southern Niagara Escarpment, Canada. New Phytol 157:365–390. https://doi.org/10.1046/j.1469-8137.2003.00678.x
Zhou Y, Zhang L, Liu J et al (2014) Climatic adaptation and ecological divergence between two closely related pine species in Southeast China. Mol Ecol 23:3504–3522. https://doi.org/10.1111/mec.12830
Acknowledgements
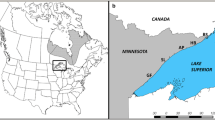
The authors would like to thank Kristine Westergaard, Jennifer Ramp Neale, Erin Tripp, Michael Wunder, Greg Ragland, Bradford S. Slaughter, and Anton A. Reznicek for providing expertise. Thanks to Audrey Spencer for preparing the distribution map for Fig. 1, and to The Nature Conservancy, the Ministry of Natural Resources and Forestry (Ontario), and United States Forest Service (Hiawatha National Forest) for providing collection permits.
Funding
Research funding was provided by the Department of Integrative Biology at the University of Colorado Denver, Botanical Society of America, and American Society of Plant Taxonomists.
Author information
Authors and Affiliations
Corresponding author
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Bard, N.W., Miller, C.S. & Bruederle, L.P. High genomic diversity maintained by populations of Carex scirpoidea subsp. convoluta, a paraphyletic Great Lakes ecotype. Conserv Genet 22, 169–185 (2021). https://doi.org/10.1007/s10592-020-01326-x
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10592-020-01326-x