Abstract
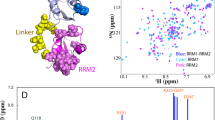
Transactive response DNA-binding protein of 43 kDa (TDP-43) is a 414-residue protein whose aberrant aggregation is implicated in neurodegenerative diseases, including amyotrophic lateral sclerosis (ALS) or frontotemporal lobar degeneration (FTLD). Intriguingly, TDP-43 has also been shown to functionally oligomerize to carry out physiological functions. TDP-43 also exists in mixed condensates or granules with other proteins (e.g. neuronal or stress granules), and its large C-terminal domain (CTD, residues 267–414) seems responsible for TDP-43 both homo- and heterotypic interactions underlying such diverse functional and pathological aggregation events. A myriad of distinct triggers may drive TDP-43 oligomerization, including interaction partners or changes in pH or salinity. In this Assignment Note, we report the complete backbone and a wealth of side chain chemical shift assignments for the CTD of TDP-43 at pH 4. The assignments presented here provide a solid starting point to study the aggregation pathway of TDP-43 at pH values below those considered physiological but relevant in pathological settings, and to contrast the aggregation behaviour under distinct conditions and in the presence of interacting partners.


Similar content being viewed by others
References
Afroz T, Hock EM, Ernst P et al (2017) Functional and dynamic polymerization of the ALS-linked protein TDP-43 antagonizes its pathologic aggregation. Nat Commun 8:45. https://doi.org/10.1038/s41467-017-00062-0
Asakawa K, Handa H, Kawakami K (2020) Optogenetic modulation of TDP-43 oligomerization accelerates ALS-related pathologies in the spinal motor neurons. Nat Commun 11:1004. https://doi.org/10.1038/s41467-020-14815-x
Babinchak WM, Haider R, Dumm BJ et al (2019) The role of liquid-liquid phase separation in aggregation of the TDP-43 low-complexity domain. J Biol Chem 294:6306–6317. https://doi.org/10.1074/jbc.RA118.007222
Barbieri EM, Shorter J (2019) TDP-43 shapeshifts to encipher FTD severity. Nat Neurosci 22:3–5. https://doi.org/10.1038/s41593-018-0299-6
Chen TC, Hsiao CL, Huang SJ, Huang JR (2016) The Nearest-neighbor effect on random-coil NMR chemical shifts demonstrated using a low-complexity amino-acid sequence. Protein Pept Lett 23:967–975. https://doi.org/10.2174/0929866523666160920100045
Chou CC, Zhang Y, Umoh ME et al (2018) TDP-43 pathology disrupts nuclear pore complexes and nucleocytoplasmic transport in ALS/FTD. Nat Neurosci 21:228–239. https://doi.org/10.1038/s41593-017-0047-3
Conicella AE, Zerze GH, Mittal J, Fawzi NL (2016) ALS mutations disrupt phase separation mediated by alpha-helical structure in the TDP-43 low-complexity C-terminal domain. Structure 24:1537–1549. https://doi.org/10.1016/j.str.2016.07.007
D’Ambrogio A, Buratti E, Stuani C, Corrado G, Romano M, Ayala YM, Baralle FE (2009) Functional mapping of the interaction between TDP-43 and hnRNP A2 in vivo. Nucleic Acids Res 37:4116–4126. https://doi.org/10.1093/nar/gkp342
Delaglio F, Grzesiek S, Vuister GW, Zhu G, Pfeifer J, Bax A (1995) NMRPipe: a multidimensional spectral processing system based on UNIX pipes. J Biomol NMR 6:277–293. https://doi.org/10.1007/BF00197809
Dewey CM, Cenik B, Sephton CF, Johnson BA, Herz J, Yu G (2012) TDP-43 aggregation in neurodegeneration: are stress granules the key? Brain Res 1462:16–25. https://doi.org/10.1016/j.brainres.2012.02.032
Gasset-Rosa F, Lu S, Yu H et al (2019) Cytoplasmic TDP-43 de-mixing independent of stress granules drives inhibition of nuclear import, loss of nuclear TDP-43, and cell death. Neuron 102:339-357.e337. https://doi.org/10.1016/j.neuron.2019.02.038
Kjaergaard M, Poulsen FM (2011) Sequence correction of random coil chemical shifts: correlation between neighbor correction factors and changes in the Ramachandran distribution. J Biomol NMR 50:157–165. https://doi.org/10.1007/s10858-011-9508-2
Kjaergaard M, Brander S, Poulsen FM (2011) Random coil chemical shift for intrinsically disordered proteins: effects of temperature and pH. J Biomol NMR 49:139–149. https://doi.org/10.1007/s10858-011-9472-x
Kroschwald S, Alberti S (2017) Gel or die: phase separation as a survival strategy. Cell 168:947–948. https://doi.org/10.1016/j.cell.2017.02.029
Laferrière F, Maniecka Z, Pérez-Berlanga M et al (2019) TDP-43 extracted from frontotemporal lobar degeneration subject brains displays distinct aggregate assemblies and neurotoxic effects reflecting disease progression rates. Nat Neurosci 22:65–77. https://doi.org/10.1038/s41593-018-0294-y
Lee W, Tonelli M, Markley JL (2015) NMRFAM-SPARKY: enhanced software for biomolecular NMR spectroscopy. Bioinformatics 31:1325–1327. https://doi.org/10.1093/bioinformatics/btu830
Li HR, Chiang WC, Chou PC, Wang WJ, Huang JR (2018) TAR DNA-binding protein 43 (TDP-43) liquid-liquid phase separation is mediated by just a few aromatic residues. J Biol Chem 293:6090–6098. https://doi.org/10.1074/jbc.AC117.001037
Lim L, Wei Y, Lu Y, Song J (2016) ALS-causing mutations significantly perturb the self-assembly and interaction with nucleic acid of the intrinsically disordered prion-like domain of TDP-43. PLoS Biol 14:e1002338. https://doi.org/10.1371/journal.pbio.1002338
Mompeán M, Romano V, Pantoja-Uceda D, Stuani C, Baralle FE, Buratti E, Laurents DV (2016) The TDP-43 N-terminal domain structure at high resolution. FEBS J 83:1242–1260. https://doi.org/10.1111/febs.13651
Nelson PT, Dickson DV, Trojanowski JQ et al (2019) Limbic-predominant age-related TDP-43 encephalopathy (LATE): consensus working group report. Brain 142:1503–1527. https://doi.org/10.1093/brain/awz099
Neumann M, Sampathu DM, Kwong LK et al (2006) Ubiquitinated TDP-43 in frontotemporal lobar degeneration and amyotrophic lateral sclerosis. Science 314:130–133. https://doi.org/10.1126/science.1134108
Riback JA, Katanski CD, Kear-Scott JL et al (2017) Stress-triggered phase separation Is an adaptive, evolutionarily tuned response. Cell 168:1028–1040. https://doi.org/10.1016/j.cell.2017.02.027
Schwarzinger S, Kroon GJ, Foss TR, Chung J, Wright PE, Dyson HJ (2001) Sequence-dependent correction of random coil NMR chemical shifts. J Am Chem Soc 123:2970–2978. https://doi.org/10.1021/ja003760i
Van Leeuwen W, Rabouille C (2019) Cellular stress leads to the formation of membraneless stress assemblies in eukaryotic cells. Traffic 20:623–638. https://doi.org/10.1111/tra.12669
Vogler TO, Wheeler JR, Nguyen ED et al (2018) TDP-43 and RNA form amyloid-like myo-granules in regenerating muscle. Nature 563:508–513. https://doi.org/10.1038/s41586-018-0665-2
Wang IF, Chang HY, Hou SC, Liou GG, Way TD, Shen CKJ (2012) The self-interaction of native TDP-43 C-terminus inhibits its degradation and contributes to early proteinopathies. Nat Commun 3:766. https://doi.org/10.1038/ncomms1766
Wang A, Conicella AE, Schmidt HB et al (2018) A single N-terminal phosphomimic disrupts TDP-43 polymerization, phase separation, and RNA splicing. EMBO J 37:e97452. https://doi.org/10.15252/embj.201797452
Zhang P, Fan B, Yang B et al (2019) Chronic optogenetic induction of stress granules is cytotoxic and reveals the evolution of ALS-FTD pathology. eLife 8:e39578. https://doi.org/10.7554/eLife.39578
Zhuo XF, Wang J, Zhang J, Jian LL, Hu HY, Lu JX (2020) Solid-state NMR reveals the structural transformation of the TDP-43 amyloidogenic region upon fibrillation. J Am Chem Soc 142:3412–3421. https://doi.org/10.1021/jacs.9b10736
Acknowledgements
This work has been funded by Grants CTQ2017-84371-P to D.P.-U. and SAF2016-76678-C2-2-R to D.V.L., from the Spanish MINECO; Grant MCB1412253 from the U. S. National Science Foundation to A.E.M; AriSLA (PathensTDP project) to E.B.; and Grant LCF/BQ/PR19/11700003 from La Caixa Foundation (ID 100010434) to M.M. NMR experiments were performed in the “Manuel Rico” NMR Laboratory (LMR) of the Spanish National Research Council (CSIC), a node of the Spanish Large-Scale National Facility (ICTS R-LRB).
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have not conflict of interest.
Additional information
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
About this article
Cite this article
Pantoja-Uceda, D., Stuani, C., Laurents, D.V. et al. NMR assignments for the C-terminal domain of human TDP-43. Biomol NMR Assign 15, 177–181 (2021). https://doi.org/10.1007/s12104-020-10002-7
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s12104-020-10002-7