Abstract
Purpose
Understanding the microbial linkages among the soils, plants, and animals is crucial for maintaining the balance of the grazed grassland ecosystem. However, previous studies always focused on the biotopes of soil, phyllosphere, and feces separately and little has been known about microbial distributions and migrations among these biotopes. In this study, systematic surveys about the overlap and differentiation among various microbiotas of biotopes and how the environmental filter on microorganisms served for the ecosystem were conducted.
Materials and methods
Soil, phyllosphere, and feces samples were collected from grazed and ungrazed grassland in Inner Mongolia, China. High-throughput sequencing and qPCR were employed to obtain the genome data. The results were analyzed by various statistical tools such as genomic analysis and machine learning classification to describe the disparities and linkages among the microbiotas of three biotopes.
Results and discussion
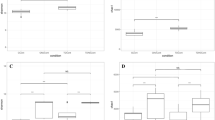
Our findings revealed that the biotopes drove the distinct microbial community assemblages with various richness, β-diversity and composition. The substantial overlaps between soil and phyllosphere in fungi, bacteria and archaea indicated that soil played the role of the microbial source for the phyllosphere. Communal OTUs could be found between phyllosphere and feces. Nevertheless, Ascomycota were the only microorganisms migrating among all the biotopes massively. After the long-term impact of feces via grazing, the soil and phyllosphere microbiota also altered significantly.
Conclusions
Biotopes drive the discrepancy of microbiota distribution among the soil, phyllosphere, and feces. Soil could potentially perform as the microbial reservoir for the phyllosphere. The phyllosphere plays the role of bridge to link soil and feces biotope through its fluctuated condition. However, there was only fungal migration running through the ecosystem to link all the biotopes. These findings promoted our understanding of the biotope contribution to microbial migration and improved the knowledge of microbial linkages in the grazed grassland ecosystem.






Similar content being viewed by others
References
Adrian L, Szewzyk U, Wecke J, Gorisch H (2000) Bacterial dehalorespiration with chlorinated benzenes. Nature 408:580–583. https://doi.org/10.1038/35046063
Andrés P, Moore JC, Simpson RT, Selby G, Cotrufo F, Denef K, Haddix ML, Shaw EA, de Tomasel CM, Molowny-Horas R, Wall DH (2016) Soil food web stability in response to grazing in a semi-arid prairie: The importance of soil textural heterogeneity. Soil Biol Biochem 97:131–143. https://doi.org/10.1016/j.soilbio.2016.02.014
Bardgett RD, van der Putten WH (2014) Belowground biodiversity and ecosystem functioning. Nature 515:505–511. https://doi.org/10.1038/nature13855
Bastian M, Heymann S, Jacomy M (2009) Gephi: An Open Source Software for Exploring and Manipulating Networks, International Aaai Conference on Weblogs & Social Media; Third International Aaai Conference on Weblogs & Social. Media:361–362
Bates ST, Berg-Lyons D, Caporaso JG, Walters WA, Knight R, Fierer N (2011) Examining the global distribution of dominant archaeal populations in soil. ISME J 5:908–917. https://doi.org/10.1038/ismej.2010.171
Beattie GA, Lindow SE (1995) The secret life of foliar bacterial pathogens on leaves. Annu Rev Phytopathol 33:145–172. https://doi.org/10.1146/annurev.py.33.090195.001045
Benjamini Y, Krieger AM, Yekutieli D (2006) Adaptive linear step-up procedures that control the false discovery rate. Biometrika 93:491–507. https://doi.org/10.1093/biomet/93.3.491
Bokulich NA, Mills DA (2013) Improved selection of internal transcribed spacer-specific primers enables quantitative, ultra-high-throughput profiling of fungal communities. Appl Environ Microbiol 79:2519–2526. https://doi.org/10.1128/AEM.03870-12
Brauer SL, Cadillo-Quiroz H, Yashiro E, Yavitt JB, Zinder SH (2006) Isolation of a novel acidiphilic methanogen from an acidic peat bog. Nature 442:192–194. https://doi.org/10.1038/nature04810
Brochier-Armanet C, Gribaldo S, Forterre P (2012) Spotlight on the Thaumarchaeota. ISME J 6:227–230. https://doi.org/10.1038/ismej.2011.145
Brookes PC, Powlson DS, Jenkinson DS (1982) Measurement of microbial biomass phosphorus in Soil. Soil Biol Biochem 14:319–329. https://doi.org/10.1016/0038-0717(82)90001-3
Bunge M, Adrian L, Kraus A, Opel M, Lorenz WG, Andreesen JR, Gorisch H, Lechner U (2003) Reductive dehalogenation of chlorinated dioxins by an anaerobic bacterium. Nature 421:357–360. https://doi.org/10.1038/nature01237
Cafarchia C, Camarda A, Romito D, Campolo M, Quaglia NC, Tullio D, Otranto D (2006) Occurrence of yeasts in cloacae of migratory birds. Mycopathologia 161:229–234. https://doi.org/10.1007/s11046-005-0194-z
Cai Y, Zheng Y, Bodelier PL, Conrad R, Jia Z (2016) Conventional methanotrophs are responsible for atmospheric methane oxidation in paddy soils. Nat Commun 7:11728. https://doi.org/10.1038/ncomms11728
Cai Y, Chang SX, Cheng Y (2017) Greenhouse gas emissions from excreta patches of grazing animals and their mitigation strategies. Earth Sci Rev 171:44–57. https://doi.org/10.1016/j.earscirev.2017.05.013
Copeland JK, Yuan L, Layeghifard M, Wang PW, Guttman DS (2015) Seasonal community succession of the phyllosphere microbiome. Mol Plant Microbe Interact 28:274–285. https://doi.org/10.1094/MPMI-10-14-0331-FI
De Vries FT, Liiri ME, Bjørnlund L, Bowker MA, Christensen S, Setälä HM, Bardgett RD (2012) Land use alters the resistance and resilience of soil food webs to drought. Nat Clim Change 2:276–280. https://doi.org/10.1038/nclimate1368
Delmotte N, Knief C, Chaffron S, Innerebner G, Roschitzki B, Schlapbach R, von Mering C, Vorholt JA (2009) Community proteogenomics reveals insights into the physiology of phyllosphere bacteria. P Natl Acad Sci USA 106:16428–16433. https://doi.org/10.1073/pnas.0905240106
Dennis RLH, Dapporto L, Dover JW (2014) Ten years of the resource-based habitat paradigm: the biotope-habitat issue and implications for conserving butterfly diversity. J Insect Biodiv 2:1–32. https://doi.org/10.12976/jib/2014.2.8
Dixon P (2003) VEGAN, a package of R functions for community ecology. J Veg Sci 14:927–930. https://doi.org/10.1658/1100-9233(2003)014[0927:Vaporf]2.0.Co;2
Egidi E, Delgado-Baquerizo M, Plett JM, Wang J, Eldridge DJ, Bardgett RD, Maestre FT, Singh BK (2019) A few Ascomycota taxa dominate soil fungal communities worldwide. Nat Commun 10:2369. https://doi.org/10.1038/s41467-019-10373-z
Erkel C, Kube M, Reinhardt R, Liesack W (2006) Genome of Rice Cluster I archaea - the key methane producers in the rice rhizosphere. Science 313:370–372. https://doi.org/10.1126/science.1127062
Faith JJ, McNulty NP, Rey FE, Gordon JI (2011) Predicting a human gut microbiota's response to diet in gnotobiotic mice. Science 333:101–104. https://doi.org/10.1126/science.1206025
Fierer N, Leff JW, Adams BJ, Nielsen UN, Bates ST, Lauber CL, Owens S, Gilbert JA, Wall DH, Caporaso JG (2012) Cross-biome metagenomic analyses of soil microbial communities and their functional attributes. Proc Natl Acad Sci USA 109:21390–21395. https://doi.org/10.1073/pnas.1215210110
Fierer N (2017) Embracing the unknown: disentangling the complexities of the soil microbiome. Nat Rev Microbiol 15:579–590. https://doi.org/10.1038/nrmicro.2017.87
Fischbach MA, Sonnenburg JL (2011) Eating for two: how metabolism establishes interspecies interactions in the gut. Cell Host Microbe 10:336–347. https://doi.org/10.1016/j.chom.2011.10.002
Giller KE, Witter E, McGrath SP (2009) Heavy metals and soil microbes. Soil Biol Biochem 41:2031–2037. https://doi.org/10.1016/j.soilbio.2009.04.026
Gourion B, Rossignol M, Vorholt JA (2006) A proteomic study of Methylobacterium extorquens reveals a response regulator essential for epiphytic growth. P Natl Acad Sci USA 103:13186–13191. https://doi.org/10.1073/pnas.0603530103
Gupta V, Kapur PK, Kumar D (2017) Modeling and measuring attributes influencing DevOps implementation in an enterprise using structural equation modeling. Inform Software Tech 92:75–91. https://doi.org/10.1016/j.infsof.2017.07.010
Hamilton N (2017) ggtern: An Extension to 'ggplot2', for the Creation of Ternary Diagrams. Anesthesiology 64:72–86. https://doi.org/10.18637/jss.v087.c03
Harrell FE (2008) Hmisc: Harrell Miscellaneous. 3:4–4
Horton MW, Bodenhausen N, Beilsmith K, Meng D, Muegge BD, Subramanian S, Vetter MM, Vilhjalmsson BJ, Nordborg M, Gordon JI, Bergelson J (2014) Genome-wide association study of Arabidopsis thaliana leaf microbial community. Nat Commun 5:5320. https://doi.org/10.1038/ncomms6320
Idnurm A, Bahn YS, Nielsen K, Lin X, Fraser JA, Heitman J (2005) Deciphering the model pathogenic fungus Cryptococcus neoformans. Nat Rev Microbiol 3:753–764. https://doi.org/10.1038/nrmicro1245
Jia T, Wang R, Fan X, Chai B (2018) A comparative study of fungal community structure, diversity and richness between the soil and the phyllosphere of native grass species in a copper tailings dam in Shanxi Province, China. Appl Sci 8:1297. https://doi.org/10.3390/app8081297
Jiao S, Xu Y, Zhang J, Lu Y (2019) Environmental filtering drives distinct continental atlases of soil archaea between dryland and wetland agricultural ecosystems. Microbiome 7:15. https://doi.org/10.1186/s40168-019-0630-9
Jost DI, Joergensen RG, Sundrum A (2012) Effect of cattle faeces with different microbial biomass content on soil properties, gaseous emissions and plant growth. Biol Fert Soil 49:61–70. https://doi.org/10.1007/s00374-012-0697-y
Knief C, Ramette A, Frances L, Alonso-Blanco C, Vorholt JA (2010) Site and plant species are important determinants of the Methylobacterium community composition in the plant phyllosphere. ISME J 4:719–728. https://doi.org/10.1038/ismej.2010.9
Kobori H (2009) Current trends in conservation education in Japan. Biol Conserv 142:1950–1957. https://doi.org/10.1016/j.biocon.2009.04.017
Konneke M, Bernhard AE, de la Torre JR, Walker CB, Waterbury JB, Stahl DA (2005) Isolation of an autotrophic ammonia-oxidizing marine archaeon. Nature 437:543–546. https://doi.org/10.1038/nature03911
Kunova A, Bonaldi M, Saracchi M, Pizzatti C, Chen X, Cortesi P (2016) Selection of Streptomyces against soil borne fungal pathogens by a standardized dual culture assay and evaluation of their effects on seed germination and plant growth. BMC Microbiol 16:272. https://doi.org/10.1186/s12866-016-0886-1
Laforest-Lapointe I, Messier C, Kembel SW (2016) Tree phyllosphere bacterial communities: exploring the magnitude of intra- and inter-individual variation among host species. PeerJ 4:e2367. https://doi.org/10.7717/peerj.2367
Lajoie G, Maglione R, Kembel SW (2020) Adaptive matching between phyllosphere bacteria and their tree hosts in a neotropical forest. Microbiome 8:70. https://doi.org/10.1186/s40168-020-00844-7
Levy-Booth DJ, Prescott CE, Grayston SJ (2014) Microbial functional genes involved in nitrogen fixation, nitrification and denitrification in forest ecosystems. Soil Biol Biochem 75:11–25. https://doi.org/10.1016/j.soilbio.2014.03.021
Lin X, Wang S, Ma X, Xu G, Luo C, Li Y, Jiang G, Xie Z (2009) Fluxes of CO2, CH4, and N2O in an alpine meadow affected by yak excreta on the Qinghai-Tibetan plateau during summer grazing periods. Soil Biol Biochem 41:718–725. https://doi.org/10.1016/j.soilbio.2009.01.007
Lindow SE, Leveau JHJ (2002) Phyllosphere microbiology. Curr Opin Biotech 13:238–243. https://doi.org/10.1016/s0958-1669(02)00313-0
Lindow SE, Brandl MT (2003) Microbiology of the phyllosphere. Appl Environ Microbiol 69:1875–1883. https://doi.org/10.1128/aem.69.4.1875-1883.2003
Liu S, Zheng R, Guo X, Wang X, Chen L, Hou Y (2018) Effects of yak excreta on soil organic carbon mineralization and microbial communities in alpine wetlands of southwest of China. J Soil Sediment 19:1490–1498. https://doi.org/10.1007/s11368-018-2149-2
Lu YH, Conrad R (2005) In situ stable isotope probing of methanogenic archaea in the rice rhizosphere. Science 309:1088–1090. https://doi.org/10.1126/science.1113435
Luo F, Zhong JX, Yang YF, Scheuermann RH, Zhou JZ (2006) Application of random matrix theory to biological networks. Phys Lett A 357:420–423. https://doi.org/10.1016/j.physleta.2006.04.076
Mahecha MD, Reichstein M, Carvalhais N, Lasslop G, Lange H, Seneviratne SI, Vargas R, Ammann C, Arain MA, Cescatti A, Janssens IA, Migliavacca M, Montagnani L, Richardson AD (2010) Global Convergence in the Temperature Sensitivity of Respiration at Ecosystem Level. Science 329:838–840. https://doi.org/10.1126/science.1189587
Maignien L, DeForce EA, Chafee ME, Eren AM, Simmons SL (2014) Ecological succession and stochastic variation in the assembly of Arabidopsis thaliana phyllosphere communities. MBio 5:e00682–e00613. https://doi.org/10.1128/mBio.00682-13
Mardani A, Streimikiene D, Zavadskas EK, Cavallaro F, Nilashi M, Jusoh A, Zare H (2017) Application of structural equation modeling (SEM) to solve environmental sustainability problems: a comprehensive review and meta-analysis. Sustainability 9:1814. https://doi.org/10.3390/su9101814
Melotto M, Underwood W, Koczan J, Nomura K, He SY (2006) Plant stomata function in innate immunity against bacterial invasion. Cell 126:969–980. https://doi.org/10.1016/j.cell.2006.06.054
Monier JM, Lindow SE (2003) Differential survival of solitary and aggregated bacterial cells promotes aggregate formation on leaf surfaces. P Natl Acad Sci USA 100:15977–15982. https://doi.org/10.1073/pnas.2436560100
Mori H, Maruyama F, Kato H, Toyoda A, Dozono A, Ohtsubo Y, Nagata Y, Fujiyama A, Tsuda M, Kurokawa K (2014) Design and experimental application of a novel non-degenerate universal primer set that amplifies prokaryotic 16S rRNA genes with a low possibility to amplify eukaryotic rRNA genes. DNA Res 21:217–227. https://doi.org/10.1093/dnares/dst052
Nielsen K, De Obaldia AL, Heitman J (2007) Cryptococcus neoformans mates on pigeon guano: implications for the realized ecological niche and globalization. Eukaryot Cell 6:949–959. https://doi.org/10.1128/EC.00097-07
Olenin S, Ducrotoy JP (2006) The concept of biotope in marine ecology and coastal management. Mar Pollut Bull 53:20–29. https://doi.org/10.1016/j.marpolbul.2006.01.003
Peay KG, Belisle M, Fukami T (2012) Phylogenetic relatedness predicts priority effects in nectar yeast communities. Proc Biol Sci 279:749–758. https://doi.org/10.1098/rspb.2011.1230
Perazzolli M, Antonielli L, Storari M, Puopolo G, Pancher M, Giovannini O, Pindo M, Pertot I (2014) Resilience of the natural phyllosphere microbiota of the grapevine to chemical and biological pesticides. Appl Environ Microbiol 80:3585–3596. https://doi.org/10.1128/AEM.00415-14
Pires AC, Cleary DF, Almeida A, Cunha A, Dealtry S, Mendonca-Hagler LC, Smalla K, Gomes NC (2012) Denaturing gradient gel electrophoresis and barcoded pyrosequencing reveal unprecedented archaeal diversity in mangrove sediment and rhizosphere samples. Appl Environ Microbiol 78:5520–5528. https://doi.org/10.1128/AEM.00386-12
Rastogi G, Coaker GL, Leveau JH (2013) New insights into the structure and function of phyllosphere microbiota through high-throughput molecular approaches. FEMS Microbiol Lett 348:1–10. https://doi.org/10.1111/1574-6968.12225
Rinnan R, Baath E (2009) Differential utilization of carbon substrates by bacteria and fungi in tundra soil. Appl Environ Microbiol 75:3611–3620. https://doi.org/10.1128/AEM.02865-08
Rinta-Kanto JM, Sinkko H, Rajala T, Al-Soud WA, Sorensen SJ, Tamminen MV, Timonen S (2016) Natural decay process affects the abundance and community structure of Bacteria and Archaea in Picea abies logs. FEMS Microbiol Ecol 92:fiw087. https://doi.org/10.1093/femsec/fiw087
Roccarina D, Lauritano EC, Gabrielli M, Franceschi F, Ojetti V, Gasbarrini A (2010) The role of methane in intestinal diseases. Am J Gastroenterol 105:1250–1256. https://doi.org/10.1038/ajg.2009.744
Rodiek J, Thomas JW (2007) Wildlife habitat planning and landscape architecture. Landscape Research 5:12–16. https://doi.org/10.1080/01426398008705949
Romero-Perez GA, Ominski KH, McAllister TA, Krause DO (2011) Effect of environmental factors and influence of rumen and hindgut biogeography on bacterial communities in steers. Appl Environ Microbiol 77:258–268. https://doi.org/10.1128/AEM.01289-09
Rothschild D et al (2018) Environment dominates over host genetics in shaping human gut microbiota. Nature 555:210–218. https://doi.org/10.1038/nature25973
Rousk J, Bååth E (2007) Fungal biomass production and turnover in soil estimated using the acetate-in-ergosterol technique. Soil Biol Biochem 39:2173–2177. https://doi.org/10.1016/j.soilbio.2007.03.023
Saggar S, Bolan NS, Bhandral R, Hedley CB, Luo J (2010) A review of emissions of methane, ammonia, and nitrous oxide from animal excreta deposition and farm effluent application in grazed pastures. New Zeal J Agr Res 47:513–544. https://doi.org/10.1080/00288233.2004.9513618
Schimel J, Balser TC, Wallenstein M (2007) Microbial stress-response physiology and its implications for ecosystem function. Ecology 88:1386–1394. https://doi.org/10.1890/06-0219
Sonnenburg ED, Zheng H, Joglekar P, Higginbottom SK, Firbank SJ, Bolam DN, Sonnenburg JL (2010) Specificity of polysaccharide use in intestinal bacteroides species determines diet-induced microbiota alterations. Cell 141:1241–1252. https://doi.org/10.1016/j.cell.2010.05.005
Stams AJ, Plugge CM (2009) Electron transfer in syntrophic communities of anaerobic bacteria and archaea. Nat Rev Microbiol 7:568–577. https://doi.org/10.1038/nrmicro2166
Sun S, Li S, Avera BN, Strahm BD, Badgley BD (2017) Soil bacterial and fungal communities show distinct recovery patterns during forest ecosystem restoration. Appl Environ Microbiol 83:e00966–e00917. https://doi.org/10.1128/AEM.00966-17
Taffner J, Cernava T, Erlacher A, Berg G (2019) Novel insights into plant-associated archaea and their functioning in arugula (Eruca sativa Mill.). J Adv Res 19:39–48. https://doi.org/10.1016/j.jare.2019.04.008
Vacher C, Hampe A, Porté AJ, Sauer U, Compant S, Morris CE (2016) The Phyllosphere: Microbial Jungle at the Plant–Climate Interface. Annu Rev Ecol Evol System 47:1–24. https://doi.org/10.1146/annurev-ecolsys-121415-032238
van de Pol JA, van Best N, Mbakwa CA, Thijs C, Savelkoul PH, Arts IC, Hornef MW, Mommers M, Penders J (2017) Gut colonization by methanogenic archaea is associated with organic dairy consumption in children. Front Microbiol 8:355. https://doi.org/10.3389/fmicb.2017.00355
Van Horn DJ, Okie JG, Buelow HN, Gooseff MN, Barrett JE, Takacs-Vesbach CD (2014) Soil microbial responses to increased moisture and organic resources along a salinity gradient in a polar desert. Appl Environ Microbiol 80:3034–3043. https://doi.org/10.1128/AEM.03414-13
Ventura M, Canchaya C, Tauch A, Chandra G, Fitzgerald GF, Chater KF, van Sinderen D (2007) Genomics of Actinobacteria: tracing the evolutionary history of an ancient phylum. Microbiol Mol Biol Rev 71:495–548. https://doi.org/10.1128/MMBR.00005-07
Vorholt JA (2012) Microbial life in the phyllosphere. Nat Rev Microbiol 10:828–840. https://doi.org/10.1038/nrmicro2910
Wang Q, Garrity GM, Tiedje JM, Cole JR (2007) Naive Bayesian classifier for rapid assignment of rRNA sequences into the new bacterial taxonomy. Appl Environ Microbiol 73:5261–5267. https://doi.org/10.1128/AEM.00062-07
Wardle DA, Bardgett RD, Klironomos JN, Setala H, van der Putten WH, Wall DH (2004) Ecological linkages between aboveground and belowground biota. Science 304:1629–1633. https://doi.org/10.1126/science.1094875
Whipps JM, Hand P, Pink D, Bending GD (2008) Phyllosphere microbiology with special reference to diversity and plant genotype. J Appl Microbiol 105:1744–1755. https://doi.org/10.1111/j.1365-2672.2008.03906.x
Wickham H (2009) ggplot2: elegant graphics for data analysis. Springer Science & Business Media, New York, pp 1–212. https://doi.org/10.1007/978-0-387-98141-3
Wilson M, Lindow SE (1994a) Coexistence among epiphytic bacterial-populations mediated through nutritional resource partitioning. Appl Environ Microb 60:4468–4477
Wilson M, Lindow SE (1994b) Ecological Similarity and Coexistence of Epiphytic Ice-Nucleating (Ice(+)) Pseudomonas-Syringae Strains and a Non-Ice-Nucleating (Ice(−)) Biological-Control Agent. Appl Environ Microb 60:3128–3137
Wu GD, Chen J, Hoffmann C, Bittinger K, Chen YY, Keilbaugh SA, Bewtra M, Knights D, Walters WA, Knight R, Sinha R, Gilroy E, Gupta K, Baldassano R, Nessel L, Li H, Bushman FD, Lewis JD (2011) Linking long-term dietary patterns with gut microbial enterotypes. Science 334:105–108. https://doi.org/10.1126/science.1208344
Yelle DJ, Ralph J, Lu F, Hammel KE (2008) Evidence for cleavage of lignin by a brown rot basidiomycete. Environ Microbiol 10:1844–1849. https://doi.org/10.1111/j.1462-2920.2008.01605.x
Yvon-Durocher G, Caffrey JM, Cescatti A, Dossena M, del Giorgio P, Gasol JM, Montoya JM, Pumpanen J, Staehr PA, Trimmer M, Woodward G, Allen AP (2012) Reconciling the temperature dependence of respiration across timescales and ecosystem types. Nature 487:472–476. https://doi.org/10.1038/nature11205
Zarraonaindia I, Owens SM, Weisenhorn P, West K, Hampton-Marcell J, Lax S, Bokulich NA, Mills DA, Martin G, Taghavi S, van der Lelie D, Gilbert JA (2015) The soil microbiome influences grapevine-associated microbiota. mBio 6:e02527–e02514. https://doi.org/10.1128/mBio.02527-14
Zhang B, Bai Z, Hoefel D, Tang L, Yang Z, Zhuang G, Yang J, Zhang H (2008) Assessing the impact of the biological control agent Bacillus thuringiensis on the indigenous microbial community within the pepper plant phyllosphere. FEMS Microbiol Lett 284:102–108. https://doi.org/10.1111/j.1574-6968.2008.01178.x
Zhao Q, Niu H, Wang Y, Cui X, Li Y, Yu Z (2019a) Response of soil bacterial communities to moisture and grazing in the Tibetan alpine steppes on a small spatial scale. Geomicrobiol J 36:559–569. https://doi.org/10.1080/01490451.2019.1583697
Zhao ZB, He JZ, Geisen S, Han LL, Wang JT, Shen JP, Wei WX, Fang YT, Li PP, Zhang LM (2019b) Protist communities are more sensitive to nitrogen fertilization than other microorganisms in diverse agricultural soils. Microbiome 7:33. https://doi.org/10.1186/s40168-019-0647-0
Zhu YG, Yoshinaga M, Zhao FJ, Rosen BP (2014) Earth Abides Arsenic Biotransformations. Annu Rev Earth Planet Sci 42:443–467. https://doi.org/10.1146/annurev-earth-060313-054942
Zimmerman NB, Vitousek PM (2012) Fungal endophyte communities reflect environmental structuring across a Hawaiian landscape. Proc Natl Acad Sci USA 109:13022–13027. https://doi.org/10.1073/pnas.1209872109
Acknowledgments
This work was supported by The National Key Research and Development Program of China (2016YFC0501802).
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Additional information
Responsible editor: Jizheng He
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
ESM 1
(PDF 285 kb)
Rights and permissions
About this article
Cite this article
Zhao, Q., Wang, Y., Ayele, G. et al. Only mass migration of fungi runs through the biotopes of soil, phyllosphere, and feces. J Soils Sediments 21, 1151–1164 (2021). https://doi.org/10.1007/s11368-020-02873-z
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11368-020-02873-z