Abstract
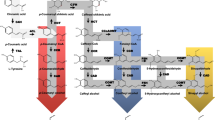
Lignin is the main component of secondary cell walls and is essential for plant development and defense. However, lignin is recognized as a major recalcitrant factor for efficiency of industrial biomass processing. Genes involved in general phenylpropanoid and monolignol-specific metabolism in sugarcane have been previously analyzed at the transcriptomic level. Nevertheless, the number of genes identified in this species is still very low. The recently released sugarcane genome sequence has allowed the genome-wide characterization of the 11 gene families involved in the monolignol biosynthesis branch of the phenylpropanoid pathway. After an exhaustive analysis of sugarcane genomes, 438 haplotypes derived from 175 candidate genes from Saccharum spontaneum and 144 from Saccharum hybrid R570 were identified as associated with this biosynthetic route. The phylogenetic analyses, combined with the search for protein conserved residues involved in the catalytic activity of the encoded enzymes, were employed to identify the family members potentially involved in developmental lignification. Accordingly, 15 candidates were identified as bona fide lignin biosynthesis genes: PTAL1, PAL2, C4H4, 4CL1, HCT1, HCT2, C3’H1, C3’H2, CCoAOMT1, COMT1, F5H1, CCR1, CCR2, CAD2, and CAD7. For this core set of lignin biosynthetic genes, we searched for the chromosomal location, the gene expression pattern, the promoter cis-acting elements, and microRNA targets. Altogether, our results present a comprehensive characterization of sugarcane general phenylpropanoid and monolignol-specific genes, providing the basis for further functional studies focusing on lignin biosynthesis manipulation and biotechnological strategies to improve sugarcane biomass utilization.






Similar content being viewed by others
Data availability
All data are available in the supplementary material.
References
Arruda P (2012) Genetically modified sugarcane for bioenergy generation. Curr Opin Biotechnol 23:315–322. https://doi.org/10.1016/j.copbio.2011.10.012
Aznar A, Chalvin C, Shih PM, Maimann M, Ebert B, Birdseye DS, Loqué D, Scheller HV (2018) Gene stacking of multiple traits for high yield of fermentable sugars in plant biomass. Biotechnol Biofuels 11:2. https://doi.org/10.1186/s13068-017-1007-6
Baltas M, Lapeyre C, Bedos-Belval F, Maturano M, Saint-Aguet P, Roussel L, Duran H, Grima-Pettenati J (2005) Kinetic and inhibition studies of cinnamoyl-CoA reductase 1 from Arabidopsis thaliana. Plant Physiol Biochem 43:746–753. https://doi.org/10.1016/j.plaphy.2005.06.003
Barrière Y, Argillier O (1993) Brown-midrib genes of maize: a review. Agronomie. 13:865–876
Barrière Y, Riboulet C, Mechin V, Maltese S, Pichon M, Cardinal A, Lapierre C, Lübberstedt T, Martinant JP (2007) Genetics and genomics of lignification in grass cell walls based on maize as model species. Genes Genomes Genom 1:133–156
Barros J, Serrani-Yarce JC, Chen F, Baxter D, Venables BJ, Dixon RA (2016) Role of bifunctional ammonia-lyase in grass cell wall biosynthesis. Nat Plants 2:16050
Barros J, Escamilla-Trevino L, Song L, Rao X, Serrani-Yarce JC, Palacios MD, Engle N, Choudhury FK, Tschaplinski TJ, Venables BJ, Mittler R, Dixon RA (2019) 4-Coumarate 3-hydroxylase in the lignin biosynthesis pathway is a cytosolic ascorbate peroxidase. Nat Commun 10:1994. https://doi.org/10.1038/s41467-019-10082-7
Bewg WP, Poovaiah C, Lan W, Ralph J, Coleman HD (2016) RNAi down-regulation of three key lignin genes in sugarcane improves glucose release without reduction in sugar production. Biotechnol Biofuels 9:270. https://doi.org/10.1186/s13068-016-0683-y
Bi C, Chen F, Jackson L, Gill BS, Li W (2010) Expression of lignin biosynthetic genes in wheat during development and upon infection by fungal pathogens. Plant Mol Biol Report 29:149–161. https://doi.org/10.1007/s11105-010-0219-8
Boerjan W, Ralph J, Baucher M (2003) Lignin biosynthesis. Annu Rev Plant Biol 54:519–546. https://doi.org/10.1146/annurev.arplant.54.031902.134938
Bonawitz ND, Kim JI, Tobimatsu Y, Ciesielski PN, Anderson NA, Ximenes E, Maeda J, Ralph J, Donohoe BS, Ladisch M, Chapple C (2014) Disruption of mediator rescues the stunted growth of a lignin-deficient Arabidopsis mutant. Nature 509:376–380. https://doi.org/10.1038/nature13084
Bottcher A, Cesarino I, Santos AB, Vicentini R, Mayer JL, Vanholme R, Morreel K, Goeminne G, Moura JCMS, Nobile PM, Carmello-Guerreiro SM, Anjos IA, Creste S, Boerjan W, Landell MGA, Mazzafera P (2013) Lignification in sugarcane: biochemical characterization, gene discovery, and expression analysis in two genotypes contrasting for lignin content. Plant Physiol 163:1539–1557. https://doi.org/10.1104/pp.113.225250
Bout S, Vermerris W (2013) A candidate-gene approach to clone the sorghum Brown midrib gene encoding caffeic acid O-methyltransferase. Mol Gen Genomics 269:205–214. https://doi.org/10.1007/s00438-003-0824-4
Brandes E (1956) Origin, dispersal and use in breeding of the Melanesian garden sugarcane and their derivatives, Saccharum officinarum L. Proc Int Soc Sugarcane Technol 9:709–750
Brill EM, Abrahams S, Hayes CM, Jenkins CLD, Watson JM (1999) Molecular characterization and expression of a wound-inducible cDNA encoding a novel cinnamyl-alcohol dehydrogenase enzyme in lucerne. Plant Mol Biol 41:279–291
Burhenne K, Kristensen BK, Rasmussen SK (2003) A new class of N-Hydroxycinnamoyltransferases. Purification, cloning and expression of a barley agmatine counaroyltransferase (EC 2.3.1.64). J Biol Chem 278:13919–13927. https://doi.org/10.1074/jbc.M213041200
Byeon Y, Lee HY, Lee K, Back K (2014) Caffeic acid O-methyltransferase is involved in the synthesis of melatonin by methylating N-acetylserotonin in Arabidopsis. J Pineal Res 57:219–227. https://doi.org/10.1111/jpi.12160
Carocha V, Soler M, Hefer C, Cassan-Wang H, Fevereiro P, Myburg AA, Paiva JAP, Grima-Pettenati J (2015) Genome-wide analysis of the lignin toolbox of Eucalyptus grandis. New Phytol 206:1297–1313. https://doi.org/10.1111/nph.13313
Cass CL, Peraldi A, Dowd PF, Mottiar Y, Santoro N, Karlen SD, Bukhman YV, Foster CE, Thrower N, Bruno LC, Moskvin OV, Johnson ET, Willhoit ME, Phutane M, Ralph J, Mansfield SD, Nicholson P, Sedbrook JC (2015) Effects of PHENYLALANINE AMMONIA LYASE (PAL) knockdown on cell wall composition, biomass digestibility, and biotic and abiotic stress responses in Brachypodium. J Exp Bot 66(14):4317–4335. https://doi.org/10.1093/jxb/erv269
Chapple CCS, Vogt T, Ellis BE, Somerville CR (1992) An Arabidopsis mutant defective in the general phenylpropanoid pathway. Plant Cell 4:1413–1424
Chiang VL, Funaoka M (1990) The difference between guaiacyl and guaiacyl-syringyl lignins in their response to kraft delignification. Holzforschung 44:309–313
Chiang VL, Puumala RJ, Takeuchi U, Eckert RE (1998) Comparison of softwood and hardwood kraft pulping. TAPPI J 71:173–176
Chow CN, Lee TY, Hung YC, Li GZ, Tseng KC, Liu YH, Kuo PL, Zheng HQ, Chang WC (2019) PlantPAN3.0: a new and updated resource for reconstructing transcriptional regulatory networks from ChIP-seq experiments in plants. Nucleic Acids Res 47(Database issue):D1155–D1163. https://doi.org/10.1093/nar/gky1081
Courtial A, Soler M, Chateigner-Boutin AL, Reymond M, Mechin V, Wang H, Grima-Pettenati J, Barrière Y (2013) Breeding grasses for capacity to biofuel production or silage feeding value: an updated list of genes involved in maize secondary cell wall biosynthesis and assembly. Maydica 58(1):67–102
D’Auria JC (2006) Acyltransferases in plants: a good time to be BAHD. Curr Opin Plant Biol 9:331–340. https://doi.org/10.1016/j.pbi.2006.03.016
D’Hont A, Ison D, Alix K, Roux C, Glaszmann JC (1998) Determination of basic chromosome numbers in the genus Saccharum by physical mapping of ribosomal RNA genes. Genome 41:221–225
Dai X, Zhuang Z, Zhao PX (2018) psRNATarget: a plant small RNA target analysis server (2017 release). Nucleic Acids Res 46:W49–W54. https://doi.org/10.1093/nar/gky316
Denness L, McKenna JF, Segonzac C, Wormit A, Madhou P, Bennett M, Mansfield J, Zipfel C, Hamann T (2011) Cell wall damage-induced lignin biosynthesis is regulated by a reactive oxygen species- and jasmonic acid-dependent process in Arabidopsis. Plant Physiol 156(3):1364–1374. https://doi.org/10.1104/pp.111.175737
Dias MOS, Ensinas AV, Nebra SA, Maciel Filho R, Rossell CEV, Maciel MRW (2009) Production of bioethanol and other bio-based materials from sugar cane bagasse: integration to convention al bioethanol production process. Chem Eng Res Des 87:1206–1216. https://doi.org/10.1016/j.cherd.2009.06.020
Do CT, Pollet B, Thévenin J, Sibout R, Denoue D, Barrière Y, Lapierre C, Jouanin L (2007) Both caffeoyl coenzyme A 3-O-methyltransferase 1 and caffeic acid O-methyltransferase 1 are involved in redundant functions for lignin, flavonoids and sinapoyl malate biosynthesis in Arabidopsis. Planta 226(5):1117–1129. https://doi.org/10.1007/s00425-007-0558-3
Edgar RC (2004) MUSCLE: multiple sequence alignment with high accuracy and high throughput. Nucleic Acids Res 32(5):1792–1797. https://doi.org/10.1093/nar/gkh340
Ehlting J, Hamberger B, Million-Rousseau R, Werck-Reichhart D (2006) Cytochromes P450 in phenolic metabolism. Phytochem Rev 5:239–270
Elkind Y, Edwards R, Mavandad M, Hedrick SA, Ribak O, Dixon RA, Lamb CJ (1990) Abnormal plant development and down-regulation of phenylpropanoid biosynthesis in transgenic tobacco containing a heterologous phenylalanine ammonia-lyase gene. Proc Natl Acad Sci U S A 87:9057–9061
Eloy NB, Voorend W, Lan W, Saleme MLS, Cesarino I, Vanholme R, Smith RA, Goeminne G, Pallidis A, Morreel K, Nicomedes J, Ralph J, Boerjan W (2017) Silencing CHALCONE SYNTHASE in maize impedes the incorporation of tricin into lignin and increases lignin content. Plant Physiol 173(2):998–1016. https://doi.org/10.1104/pp.16.01108
Escamilla-Treviño LL, Shen H, Uppalapati SR, Ra T, Tang Y, Hernandez T (2010) Switchgrass (Panicum virgatum) possesses a divergent family of cinnamoyl CoA reductases with distinct biochemical properties. New Phytol 185:143–155. https://doi.org/10.1111/j.1469-8137.2009.03018.x
Eudes A, Pollet B, Sibout R, Do CT, Seguin A, Lapierre C, Jouanin L (2006) Evidence for a role of AtCAD1 in lignification of elongating stems of Arabidopsis thaliana. Planta. 225:23–39. https://doi.org/10.1007/s00425-006-0326-9
Fan D, Li C, Fan C, Hu J, Li J, Yao S, Lu W, Yan Y, Luo K (2020) MicroRNA6443-mediated regulation of FERULATE 5-HYDROXYLASE gene alters lignin composition and enhances saccharification in Populus tomentosa. New Phytol 226(2):410–425. https://doi.org/10.1111/nph.16379
Ferreira SS, Simões MS, Carvalho GG, de Lima LGA, Svartman RMA, Cesarino I (2019) The lignin toolbox of the model grass Setaria viridis. Plant Mol Biol 101(3):235–255. https://doi.org/10.1007/s11103-019-00897-9
Ferrer JL, Zubieta C, Dixon RA, Noel JP (2005) Crystal structures of alfalfa caffeoyl coenzyme A 3-O-methyltransferase. Plant Physiol 137:1009–1017. https://doi.org/10.1104/pp.104.048751
Formalé S, Rencoret J, Garcia-Calvo L, Capellades M, Encina A, Santiago R, Rigau J, Gutiérrez A, del Río JC, Caparros-Ruiz D (2015) Cell wall modification triggere by down-regulationof coumarate 3-hydroxylase-1 in maize. Plant Sci 236:272–282. https://doi.org/10.1016/j.plantsci.2015.04.007
Franke R, McMichael M, Meyer K, Shirley AM, Cusumano JC, Chapple C (2000) Modified lignin in tobacco and poplar plants over-expressing the Arabidopsis gene encoding ferulate 5-hydroxylase. Plant J 22:223–234
Franke R, Humphreys JM, Hemm MR, Denault JW, Ruegger MO, Cusumano JC, Chapple C (2002) The Arabidopsis REF8 gene encodes the 3-hydroxylase of phenylpropanoid metabolism. Plant J 30:33–45. https://doi.org/10.1046/j.1365-313X.2002.01266.x
Freudenberg K, Nash AC (1968) Constitution and biosynthesis of lignin. Springer-Verlag, Berlin
Fu C, Mielenz JR, Xiao X, Ge Y, Hamilton CY, Rodriguez M, Fang Chen F, Foston M, Ragauskas A, Bouton J, Dixon RA, Wang ZY (2011) Genetic manipulation of lignin reduces recalcitrance and improves ethanol production from switchgrass. Proc Natl Acad Sci U S A 108:3803–3808. https://doi.org/10.1073/pnas.1100310108
Galliano H, Cabane M, Eckerskorn C, Lottspeich F, Sandermann H, Ernst D (1993) Molecular cloning, sequence analysis and elicitor-/ozone-induced accumulation of cinnamyl alcohol dehydrogenase from Norway spruce (Picea abies). Plant Mol Biol 23:145–156
García JR, Anderson N, Le-Feuvre R, Iturra C, Elissetche J, Chapple C, Valenzuela S (2014) Rescue of syringyl lignin and sinapate ester biosynthesis in Arabidopsis thaliana by a coniferaldehyde 5-hydroxylase from Eucalyptus globulus. Plant Cell Rep 33(8):1263–1274. https://doi.org/10.1007/s00299-014-1614-7
Garsmeur O, Droc G, Antonise R, Grimwood J, Potier B, Aitken K, Jenkins J, Martin G, Charron C, Hervouet C, Costet L, Yahiaoui N, Healey A, Sims D, Cherukuri Y, Sreedasyam A, Kilian A, Chan A, Sluys MAV, Swaminathan K, Town C, Bergès H, Simmons B, Glaszmann JC, van der Vossen E, Henry R, Schmutz J, D’Hont A (2018) A mosaic monoploid reference sequence for the highly complex genome of sugarcane. Nat Commun 9(1):2638. https://doi.org/10.1038/s41467-018-05051-5
Goujon T, Ferret V, Mila I, Pollet B, Ruel K, Burlat V, Joseleau JP, Barrière Y, Lapierre C, Jouanin L (2003) Down-regulation of the AtCCR1 gene in Arabidopsis thaliana: effects on phenotype, lignins and cell wall degradability. Planta. 217:218–228. https://doi.org/10.1007/s00425-003-0987-6
Grivet L, Glaszmann JC, D’Hont A (2006) Evolution and conservation of crops. In: Motley TJ (ed) Darwin’s harvest: new approaches to the origins. Columbia University Press, New York, pp 49–66
Gui J, Shen J, Li L (2011) Functional characterization of evolutionarily divergent 4-coumarate: coenzyme A ligases in rice. Plant Physiol 157:574–586. https://doi.org/10.1104/pp.111.178301
Guillaumie S, San-Clemente H, Deswarte C, Martinez Y, Lapierre C, Murigneux A, Barrière Y, Pichon M, Goffner D (2007) MAIZEWALL. Database and developmental gene expression profiling of cell wall biosynthesis and assembly in maize. Plant Physiol 143:339–363. https://doi.org/10.1104/pp.106.086405
Guindon S, Dufayard JF, Lefort V, Anisimova M, Hordijk W, Gascuel O (2010) New algorithms and methods to estimate maximum-likelihood phylogenies: assessing the performance of PhyML 3.0. Syst Biol 59:307–321. https://doi.org/10.1093/sysbio/syq010
Guo D, Chen F, Inoue K, Blount JW, Dixon RA (2001) Down-regulation of caffeic acid 3-O-methyltransferase and caffeoyl CoA 3-O-methyltransferase in transgenic alfalfa: impacts on lignin structure and implications for the biosynthesis of G and S lignin. Plant Cell 13:73–88
Ha CM, Escamilla-Trevino L, Yarce JC, Kim H, Ralph J, Chen F, Dixon RA (2016) An essential role of caffeoyl shikimate esterase in monolignol biosynthesis in Medicago truncatula. Plant J 86(5):363–375. https://doi.org/10.1111/tpj.13177
Halpin C, Holt K, Chojecki J, Oliver D, Chabbert B, Monties B, Edwards K, Barakate A, Foxon GA (1998) Brown-midrib maize (bm1) – a mutation affecting the cinnamyl alcohol dehydrogenase gene. Plant J 14(5):545–553. https://doi.org/10.1046/j.1365-313X.1998.00153.x
Hamann T, Bennett M, Mansfield J, Somerville C (2009) Identification of cell-wall stress as a hexose-dependent and osmosensitive regulator of plant responses. Plant J 57(6):1015–1026. https://doi.org/10.1111/j.1365-313X.2008.03744.x
Hickman R, Van Verk MC, Van Dijken AJH, Mendes MP, Vroegop-Vos IA, Caarls L, Steenbergen M, Van der Nagel I, Wesselink GJ, Jironkin A, Talbot A, Rhodes J, De Vries M, Schuurink RC, Denby K, Pieterse CMJ, Van Wees SCM (2017) Architecture and dynamics of the jasmonic acid gene regulatory network. Plant Cell 29(9):2086–2105. https://doi.org/10.1105/tpc.16.00958
Hirano K, Aya K, Kondo M, Okuno A, Morinaka Y, Matsuoka M (2012) OsCAD2 is the major CAD gene responsible for monolignol biosynthesis in rice culm. Plant Cell Rep 31:91–101. https://doi.org/10.1007/s00299-011-1142-7
Hsieh LS, Ma GJ, Yang CC, Lee PD (2010) Cloning, expression, sitedirected mutagenesis and immunolocalization of phenylalanine ammonia-lyase in Bambusa oldhamii. Phytochemistry. 71:1999–2009. https://doi.org/10.1016/j.phytochem.2010.09.019
Hu WJ, Harding SA, Lung J, Popko JL, Ralph J, Stokke DD et al (1999) Repression of lignin biosynthesis promotes cellu-lose accumulation and growth in transgenic trees. Nat Biotechnol 17:808–812
Huang J, Gu M, Lai Z, Fan B, Shi K, Zhou YH, Yu JQ, Chen Z (2010) Functional analysis of the Arabidopsis PAL gene family in plant growth, development, and response to environmental stress. Plant Physiol 153:1526–1538. https://doi.org/10.1104/pp.110.157370
Huntley SK, Ellis D, Gilbert M, Chapple C, Mansfield SD (2003) Significant increases in pulping efficiency in C4H-F5H-transformed poplars: improved chemical savings and reduced environmental toxins. J Agric Food Chem 51:6178
Jardim-Messeder D, Silva TF, Fonseca JP, Nicomedes J, Barzilai L, Felix-Cordeiro T, Pereira JC, Rodrigues-Ferreira C, Bastos I, Silva TC, Waldow VA, Cassol D, Pereira W, Flausino B, Carniel A, Faria J, Moraes T, Cruz FP, Loh R, Van Montagu M, Loureiro ME, Souza SR, Mangeon A, Sachetto-Martins G (2020) Identification of genes from the general phenylpropanoid and monolignol-specific metabolism in two sugarcane lignin-contrasting genotypes. Mol Gen Genomics 295:717–739
Jones L, Ennos AR, Turner SR (2001) Cloning and characterization of irregular xylem4 (irx4): a severely lignin-deficient mutant of Arabidopsis. Plant J 26:205–216. https://doi.org/10.1046/j.1365-313x.2001.01021.x
Jun SY, Sattler SA, Cortez GS, Vermerris W, Sattler SE, Kang C (2018a) Biochemical and structural analysis of substrate specificity of a phenylalanine ammonia-lyase. Plant Physiol 176(2):1452–1468. https://doi.org/10.1104/pp.17.01608
Jun SY, Sattler SA, Cortez GS, Vermerris W, Sattler SE, Kang C (2018b) Biochemical and structural analysis of substrate specificity of a phenylalanine ammonia-lyase. Plant Physiol 176(2):1452–1468. https://doi.org/10.1104/pp.17.01608
Jung HJG, Ni WT, Chapple CCS, Meyer K (1999) Impact of lignin composition on cell wall degradability in an Arabidopsis mutant. J Sci Food Agric 79:922–928
Jung JH, Fouad WM, Vermerris W, Gallo M, Altpeter F (2012) RNAi suppression of lignin biosynthesis in sugarcane reduces recalcitrance for biofuel production from lignocellulosic biomass. Plant Biotechnol J 10:1067–1076. https://doi.org/10.1111/j.1467-7652.2012.00734.x
Jung JH, Vermerris W, Gallo M, Fedenko JR, Erickson JE, Altpeter F (2013) RNA interference suppression of lignin biosynthesis increases fermentable sugar yields for biofuel production from field-grown sugarcane. Plant Biotechnol J 11:709–716. https://doi.org/10.1111/pbi.12061
Jung JH, Kannan B, Dermawan H, Moxley GW, Altpeter F (2016) Precision breeding for RNAi suppression of a major 4-coumarate:coenzyme A ligase gene improves cell wall saccharification from field grown sugarcane. Plant Mol Biol 92(4–5):505–517. https://doi.org/10.1007/s11103-016-0527-y
Kajita S, Hishiyama S, Tomimura Y, Katayama Y, Omori S (1997) Structural characterization of modified lignin in transgenic tobacco plants in which the activity of 4-coumarate:coenzyme A ligase is depressed. Plant Physiol 114(3):871–879
Kasirajan L, Hoang NV, Furtado A, Botha FC, Henry RJ (2018) Transcriptome analysis highlights key differentially expressed genes involved in cellulose and lignin biosynthesis of sugarcane genotypes varying in fiber content. Sci Rep 8(1):11612. https://doi.org/10.1038/s41598-018-30033-4
Khandal H, Singh AP, Chattopadhyay D (2020) MicroRNA397b-LACCASE2 module regulates root lignification under water- and phosphate deficiency. Plant Physiol 182(3):1387–1403. https://doi.org/10.1104/pp.19.00921
Kim SJ, Kim MR, Bedgar D, Moinuddin SGA, Cardenas CL, Davin LB, Kang CCK, Lewis NG (2004) Functional reclassification of the putative cinnamyl alcohol dehydrogenase multigene family in Arabidopsis. Proc Nat Acad Sci U S A 101:1455–1460. https://doi.org/10.1073/pnas.0307987100
Kim RJ, Kim HJ, Shim D, Suh MC (2016) Molecular and biochemical characterizations of the monoacylglycerol lipase gene family of Arabidopsis thaliana. Plant J 85:758–771. https://doi.org/10.1111/tpj.13146
Koshiba T, Hirose N, Mukai M, Yamamura M, Hattori T, Suzuki S, Sakamoto M, Umezawa T (2013) Characterization of 5-hydroxyconiferaldehyde Omethyltransferase in Oryza sativa. Plant Biotechnol 30:157–167. https://doi.org/10.5511/plantbiotechnology.13.0219a
Lallemand LA, Zubieta C, Lee SG, Wang Y, Acajjaoui S, Timmins J, McSweeney S, Jez JM, McCarthy JG, McCarth AA (2012) A structural basis for the biosynthesis of the major chlorogenic acids found in coffee. Plant Physiol 160:249–260. https://doi.org/10.1104/pp.112.202051
Lauvergeat V, Lacomme C, Lacombe E, Lasserre E, Roby D, Grima-Pettenati J (2001) Two cinnamoyl-CoA reductase (CCR) genes from Arabidopsis thaliana are differentially expressed during development and in response to infection with pathogenic bacteria. Phytochemistry. 57:1187–1195. https://doi.org/10.1016/S0031-9422(01)00053-X
Lee D, Meyer K, Chapple C, Douglas CJ (1997) Antisense suppression of 4-coumarate: coenzyme A ligase activity in Arabidopsis leads to altered lignin subunit composition. Plant Cell 9:1985–1998. https://doi.org/10.1105/tpc.9.11.1985
Levsh O, Chiang YC, Tung CF, Noel JP, Wang Y, Weng JK (2016) Dynamic conformational states dictate selectivity toward the native substrate in a substrate-permissive acyltransferase. Biochemistry. 55:6314–6326. https://doi.org/10.1021/acs.biochem.6b00887
Li X, Chapple C (2010) Understanding lignification: challenges beyond monolignol biosynthesis. Plant Physiol 154:449–452. https://doi.org/10.1104/pp.110.162842
Li X, Yang Y, Yao J, Chen G, Li X, Zhang Q, Wu C (2009) FLEXIBLE CULM 1 encoding a cinnamyl-alcohol dehydrogenase controls culm mechanical strength in rice. Plant Mol Biol 69:685–697. https://doi.org/10.1007/s11103-008-9448-8
Li X, Chen W, Zhao Y, Xiang Y, Jiang H, Zhu S, Cheng B (2013) Downregulation of caffeoyl-CoA O-methyltransferase (CCoAOMT) by RNA interference leads to reduced lignin production in maize straw. Genet Mol Biol 36(4):540–546. https://doi.org/10.1590/S1415-47572013005000039
Li J, Fan F, Wang L, Zhan Q, Wu P, Du J, Yang X, Liu Y (2016) Cloning and expression analysis of cinnamoyl-CoA reductase (CCR) genes in sorghum. PeerJ. 4:e2005. https://doi.org/10.7717/peerj.2005
Li M, Liang Z, He S, Zeng Y, Jing Y, Fang W, Wu K, Wang G, Ning X, Wang L, Li S, Tan H, Tan F (2017a) Genome-wide identification of leaf abscission associated microRNAs in sugarcane (Saccharum officinarum L.). BMC Genomics 18(1):754. https://doi.org/10.1186/s12864-017-4053-3
Li W, Zhang F, Wu R, Jia L, Li G, Guo Y, Liu C, Wang G (2017b) A novel N-methyltransferase in Arabidopsis appears to feed a conserved pathway for nicotinate detoxification among land plants and is associated with lignin biosynthesis. Plant Physiol 174(3):1492–1504. https://doi.org/10.1104/pp.17.00259
Llerena JPP, Figueiredo R, Brito MDS, Kiyota E, Mayer JLS, Araujo P, Schimpl FC, Dama M, Pauly M, Mazzafera P (2019) Deposition of lignin in four species of Saccharum. Sci Rep 9(1):5877. https://doi.org/10.1038/s41598-019-42350-3
Lu S, Li Q, Wei H, Chang MJ, Tunlaya-Anukit S, Kim H, Liu J, Song J, Sun YH, Yuan L, Yeh TF, Peszlen I, Ralph J, Sederoff RR, Chiang VL (2013) Ptr-miR397a is a negative regulator of laccase genes affecting lignin content in Populus trichocarpa. Proc Natl Acad Sci U S A 110(26):10848–10853. https://doi.org/10.1073/pnas.1308936110
Ma QH (2010) Functional analysis of a cinnamyl alcohol dehydrogenase involved in lignin biosynthesis in wheat. J Exp Bot 61(10):2735–2744. https://doi.org/10.1093/jxb/erq107
Ma D, Constabel CP (2019) MYB repressors as regulators of phenylpropanoid metabolism in plants. Trends Plant Sci 24(3):275–289. https://doi.org/10.1016/j.tplants.2018.12.003
Ma QH, Luo HR (2015) Biochemical characterization of caffeoyl coenzyme A 3-O-methyltransferase from wheat. 2015. Planta 242:113–122
Maeda HA (2016) Lignin biosynthesis: tyrosine shortcut in grasses. Nat Plants 2:16080
Martin AP, Palmer WM, Brown C, Abel C, Lunn JE, Furbank RT, Grof CPL (2016) A developing Setaria viridis internode: an experimental system for the study of biomass generation in a C4 model species. Biotechnol Biofuels 9:45. https://doi.org/10.1186/s13068-016-0457-6
Memelink J, Verpoorte R, Kijne JW (2001) ORCAnization of jasmonate-responsive gene expression in alkaloid metabolism. Trends Plant Sci 6(5):212–219. https://doi.org/10.1016/s1360-1385(01)01924-0
Meyer K, Shirley AM, Cusumano JC, Bell-Lelong DA, Chapple C (1998) Lignin monomer composition is determined by the expression of a cytochrome P450-dependent monooxygenase. Proc Natl Acad Sci U S A 95:6619–6623
Muthamilarasan M, Khan Y, Jaishankar J, Shweta S, Lata C, Prasad M (2015) Integrative analysis and expression profiling of secondary cell wall genes in C4 biofuel model Setaria italica reveals targets for lignocellulose bioengineering. Front Plant Sci 6:965. https://doi.org/10.3389/fpls.2015.00965
O’Malley DM, Porter S, Sederoff RR (1992) Purification, characterization, and cloning of cinnamyl alcohol dehydrogenase in loblolly pine (Pinus taeda L). Plant Physiol 98:1364–1371
OECD/FAO (2018) OECD-FAO Agricultural Outlook 2018-2027. OECD Publishing, Paris/FAO, Rome. https://doi.org/10.1787/agr_outlook-2018-en
Pallas JA, Paiva NL, Lamb C, Dixon RA (1996) Tobacco plants epigenetically suppressed in phenylalanine ammonia-lyase expression do not develop systemic acquired resistance in response to infection by tobacco mosaic virus. Plant J 10:281–293
Panje R, Babu C (1960) Studies in Saccharum spontaneum. Distribution and geographical association of chromosome numbers. Cytologia 25:152–172
Park HL, Bhoo SH, Kwon M, Lee SW, Cho MH (2017) Biochemical and expression analyses of the rice cinnamoyl-CoA reductase gene family. Front Plant Sci 8:2099. https://doi.org/10.3389/fpls.2017.02099
Pauwels L, Morreel K, De Witte E, Lammertyn F, Montagu MV, Boerjan W, Inzé D, Goossens A (2008) Mapping methyl jasmonate-mediated transcriptional reprogramming of metabolism and cell cycle progression in cultured Arabidopsis cells. Proc Natl Acad Sci U S A 105(4):1380–1385. https://doi.org/10.1073/pnas.0711203105
Pichon M, Courbou I, Beckert M, Boudet AM, Grima-Pettenati J (1998) Cloning and characterization of two maize cDNAs encoding cinnamoyl-CoA reductase (CCR) and differential expression of the corresponding genes. Plant Mol Biol 38:671–676. https://doi.org/10.1023/A:1006060101866
Pincon G, Maury S, Hoffmann L, Geoffroy P, Lapierre C, Pollet B, Legrand M (2002) Repression of O-methyltransferase genes in transgenic tobacco affects lignin synthesis and plant growth. Phytochemistry 57:1167–1176. https://doi.org/10.1016/S0031-9422(01)00098-X
Piquemal J, Chamayou S, Nadaud I, Beckert M, Barrière Y, Mila I, Lapierre C, Rigau J, Puigdomenech P, Jauneau A, Digonnet C, Boudet AM, Goffner D, Pichon M (2002) Down-regulation of caffeic acid o-methyltransferase in maize revisited using a transgenic approach. Plant Physiol 130(4):1675–1685. https://doi.org/10.1104/pp.012237
Poelking VGC, Giordano A, Ricci-Silva ME, Williams TCR, Peçanha DA, Ventrella MC, Rencoret J, Ralph J, Barbosa MHP, Loureiro M (2015) Analysis of a modern hybrid and an ancient sugarcane implicates a complex interplay of factors in affecting recalcitrance to cellulosic ethanol production. PLoS One 10(8):e0134964. https://doi.org/10.1371/journal.pone.0134964
Ponniah SK, Shang Z, Akbudak MA et al (2017) Down-regulation of hydroxycinnamoyl CoA: shikimate hydroxycinnamoyl transferase, cinnamoyl CoA reductase, and cinnamyl alcohol dehydrogenase leads to lignin reduction in rice (Oryza sativa L. ssp. japonica cv. Nipponbare). Plant Biotechnol Rep 11:17–27. https://doi.org/10.1007/s11816-017-0426-y
Poovaiah CR, Nageswara-Rao M, Soneji JR, Baxte HL, Stewart CN (2014) Altered lignin biosynthesis using biotechnology to improve lignocellulosic biofuel feedstocks. Plant Biotechnol J 12:1163–1173. https://doi.org/10.1111/pbi.12225
Prouse MB, Campbell MM (2012) The interaction between MYB proteins and their target DNA binding sites. Biochim Biophys Acta 1819(1):67–77. https://doi.org/10.1016/j.bbagrm.2011.10.010
Raes J, Rohde A, Christensen JH, Van de Peer Y, Boerjan W (2003) Genome-wide characterization of the lignification toolbox in Arabidopsis. Plant Physiol 133:1051–1071. https://doi.org/10.1104/pp.103.026484
Rakoczy M, Femiak I, Alejska M, Figlerowicz M, Podkowinski J (2018) Sorghum CCoAOMT and CCoAOMT-like gene evolution, structure, expression and the role of conserved amino acids in protein activity. Mol Gen Genomics 293:1077–1089. https://doi.org/10.1007/s00438-018-1441-6
Ralph J, Akiyama T, Kim H, Lu F, Schatz PF, Marita JM, Ralph SA, Reddy MSS, Chen F, Dixon RA (2006) Effects of coumarate 3-hydroxylase down-regulation on lignin structure. J Biol Chem 281:8843–8853. https://doi.org/10.1074/jbc.M511598200
Ralph J, Brunow G, Boerjan W (2007) Lignins. In: Rose F, Osborne K (eds) Encyclopedia of life sciences. Wiley, Chichester, pp 1–10. https://doi.org/10.1002/9780470015902.a0020104
Ramos RLB, Tovar FJ, Junqueira RM, Lino FB, Sachetto-Martins G (2001) Sugarcane expressed sequences tags (ESTs) encoding enzymes involved in lignin biosynthesis pathways. Genet Mol Biol 24:235–241. https://doi.org/10.1590/S1415-47572001000100031
Rohde A, Morreel K, Ralph J, Goeminne G, Hostyn V, De Rycke R, Kushnir S, Doorsselaere JV, Joseleau JP, Vuylsteke M, Driessche GV, Beeumen JV, Messens E, Boerjan W (2004) Molecular phenotyping of the pal1 and pal2 mutants of Arabidopsis thaliana reveals far-reaching consequences on phenylpropanoid, amino acid, and carbohydrate metabolism. Plant Cell 16(10):2749–2771. https://doi.org/10.1105/tpc.104.023705
Romero I, Fuertes A, Benito MJ, Malpica JM, Leyva A, Paz-Ares J (1998) More than 80R2R3-MYB regulatory genes in the genome of Arabidopsis thaliana. Plant J 14(3):273–284
Rösler J, Krekel F, Amrhein N, Schmid J (1997) Maize phenylalanine ammonia-lyase has tyrosine ammonia-lyase activity. Plant Physiol 113:175–179
Röther D, Poppe L, Morlock G, Viergutz S, Rétey J (2002) An active site homology model of phenylalanine ammonia-lyase from Petroselinum crispum. Eur J Biochem 269:3065–3075. https://doi.org/10.1046/j.1432-1033.2002.02984.x
Rubinelli PM, Chuck G, Li X, Meilan R (2013) Constitutive expression of the Corngrass1 microRNA in poplar affects plant architecture and stem lignin content and composition. Biomass Bioenergy 54:312–321
Ruegger M, Chapple C (2001) Mutations that reduce sinapoylmalate accumulation in Arabidopsis thaliana define loci with diverse roles in phenylpropanoid metabolism. Genetics 159:1741–1749
Saballos A, Vermerris W, Rivera L, Ejeta G (2008) Allelic association, chemical characterization and saccharification properties of brown midribmutants of sorghum (Sorghum bicolor (L.) Moench). Bioenergy Res 2:198–204. https://doi.org/10.1007/s12155-008-9025-7
Saballos A, Ejeta G, Sanche E, Kang C, Vermerris W (2009) A genome wide analysis of the cinnamyl alcohol dehydrogenase family in sorghum (Sorghum bicolor (L.) Moench) identifies SbCAD2 as the brown midrib6 gene. Genetics. 181:783–795. https://doi.org/10.1534/genetics.108.098996
Saballos A, Sattler SE, Sanchez E, Foster TP, Xin Z, Kang C, Pedersen JF, Vermerris W (2012) Brown midrib2 (Bmr2) encodes the major 4-coumarate:coenzyme A ligase involved in lignin biosynthesis in sorghum (Sorghum bicolor (L.) Moench). Plant J 70:818–830. https://doi.org/10.1111/j.1365-313X.2012.04933.x
Saleme MLS, Cesarino I, Vargas L, Kim H, Vanholme R, Goeminne G, Van Acker R, Fonseca FCA, Pallidis A, Voorend W, Junior JN, Padmakshan D, Van Doorsselaere J, Ralph J, Boerjan W (2017) Silencing CAFFEOYL SHIKIMATE ESTERASE affects lignification and improves saccharification in poplar. Plant Physiol 175(3):1040–1057. https://doi.org/10.1104/pp.17.00920
Salzman RA, Brady JA, Finlayson SA, Buchanan CD, Summer EJ, Sun F, Klein PE, Klein RR, Pratt LH, Cordonnier-Pratt MM, Mullet JE (2005) Transcriptional profiling of sorghum induced by methyl jasmonate, salicylic acid, and aminocyclopropane carboxylic acid reveals cooperative regulation and novel gene responses. Plant Physiol 138(1):352–368. https://doi.org/10.1104/pp.104.058206
Santos AB, Bottcher A, Kiyota E, Mayer JL, Vicentini R, Brito MS, Creste S, Landell MGA, Mazzafera P (2015) Water stress alters lignin content and related gene expression in two sugarcane genotypes. J Agric Food Chem 63(19):4708–4720. https://doi.org/10.1021/jf5061858
Sato Y, Takehisa H, Kamatsuki K, Minami H, Namiki N, Ikawa H, Ohyanagi H, Sugimoto K, Antonio BA, Nagamura Y (2013) RiceXPro version 3.0: expanding the informatics resource for rice transcriptome. Nucleic Acids Res 41(Database issue):D1206–D1213. https://doi.org/10.1093/nar/gks1125
Sattler SE, Saathoff AJ, Haas EJ, Palmer NA, Funnell-Harris DL, Sarath G, Pedersen JF (2009) A nonsense mutation in a cinnamyl alcohol dehydrogenase gene is responsible for the sorghum brown midrib6 phenotype. Plant Physiol 150:584–595. https://doi.org/10.1104/pp.109.136408
Sattler SE, Saballos A, Xin Z, Funnell-Harris DL, Vermerris W, Pedersen JF (2014) Characterization of novel sorghum brown midrib mutants from an EMS-mutagenized population. G3 (Bethesda) 4(11):2115–2124. https://doi.org/10.1534/g3.114.014001
Sattler SA, Walker AM, Vermerris W, Sattler SE, Kang C (2017) Structural and biochemical characterization of Cinnamoyl-CoA reductases. Plant Physiol 173(2):1031–1044. https://doi.org/10.1104/pp.16.01671
Schilmiller AL, Stout J, Weng JK, Humphreys J, Ruegger MO, Chapple C (2009) Mutations in the cinnamate 4-hydroxylase gene impact metabolism, growth and development in Arabidopsis. Plant J 60(5):771–782. https://doi.org/10.1111/j.1365-313X.2009.03996.x
Schneider K, Hövel K, Witzel K, Hamberger B, Schomburg D, Kombrink E, Stuible HP (2003) The substrate specificity-determining amino acid code of 4-coumarate:CoA ligase. Proc Natl Acad Sci U S A 100(14):8601–8606. https://doi.org/10.1073/pnas.1430550100
Screenivasan TV, Ahloowalia BS, Heinz DJ (1987) Sugarcane improvement through breeding. Cytogenetics. 5:211–253
Shadle G, Chen F, Srinivasa Reddy MS, Jackson L, Nakashima J, Dixon RA (2007) Down-regulation of hydroxycinnamoyl CoA: shikimate hydroxycinnamoyl transferase in transgenic alfalfa affects lignification, development and forage quality. Phytochemistry 68(11):1521–1529. https://doi.org/10.1016/j.phytochem.2007.03.022
Sharma D, Tiwari M, Pandey A, Bhatia C, Sharma A, Trivedi PK (2016) MicroRNA858 is a potential regulator of phenylpropanoid pathway and plant development. Plant Physiol 71(2):944–959. https://doi.org/10.1104/pp.15.01831
Shi R, Su YH, Li Q, Heber S, Sederoff R, Chiang VL (2010) Towards a systems approach for lignin biosynthesis in populus trichocarpa: transcript abundance and specificity of the monolignol biosynthetic genes. Plant Cell Physiol 51:144–163. https://doi.org/10.1093/pcp/pcp175
Sibout R, Eudes A, Mouille G, Pollet B, Lapierre C, Jouanin L, Seguin A (2005) CINNAMYL ALCOHOL DEHYDROGENASE C and D are the primary genes involved in lignin biosynthesis in the floral stem of Arabidopsis. Plant Cell 17:2059–2076. https://doi.org/10.1105/tpc.105.030767
St-Pierre B, De Luca V (2000) Evolution of acyltransferase genes: origin and diversification of the BAHD superfamily of acyltransferases involved in secondary metabolism. In: Romeo JT et al (eds) Recent advances in phytochemistry. Evolution of metabolic pathway 34:285–315
Stuible HP, Kombrink E (2001) Identification of the substrate specificity-conferring amino acid residues of 4-coumarate:coenzyme a ligase allows the rational design of mutant enzymes with new catalytic properties. J Biol Chem 276:26893–26897. https://doi.org/10.1074/jbc.M100355200
Takeda Y, Koshiba T, Tobimatsu Y, Suzuki S, Murakami S, Yamamura M, Rahman MM, Takano T, Hattori T, Sakamoto M, Umezawa T (2017) Regulation of CONIFERALDEHYDE 5-HYDROXYLASE expression to modulate cell wall lignin structure in rice. Planta 246(2):337–349. https://doi.org/10.1007/s00425-017-2692-x
Takeda Y, Suzuki S, Tobimatsu Y, Osakabe K, Osakabe Y, Ragamustari SK, Sakamoto M, Umezawa T (2018a) Lignin characterization of rice CONIFERALDEHYDE 5-HYDROXYLASE loss-of-function mutants generated with the CRISPR/Cas9 system. Plant J 97:543–554. https://doi.org/10.1111/tpj.14141
Takeda Y, Tobimatsu Y, Karlen SD, Koshiba T, Suzuki S, Yamamura M, Murakami S, Mukai M, Hattori T, Osakabe K, Ralph J, Sakamoto M, Umezawa T (2018b) Downregulation of p-COUMAROYL ESTER 3-HYDROXYLASE in rice leads to altered cell wall structures and improves biomass saccharification. Plant J 95:796–811. https://doi.org/10.1111/tpj.13988
Tamasloukht B, Wong Quai Lam MSJ, Martinez Y, Tozo K, Barbier O, Jourda C, Jauneau A, Borderies G, Balzergue S, Renou JP, Huguet S, Martinant JP, Tatout C, Lapierre C, Barrière Y, Goffner D, Pichon M (2011) Characterization of a cinnamoyl-CoA reductase 1 (CCR1) mutant in maize: effects on lignification, fibre development, and global gene expression. J Exp Bot 62:3837–3848. https://doi.org/10.1093/jxb/err077
Tamura K, Stecher G, Peterson D, Filipski A, Kumar S (2013) MEGA6: molecular evolutionary genetics analysis version 6.0. Mol Biol Evol 30:2725–2729. https://doi.org/10.1093/molbev/mst197
Trumbo JL, Zhang B, Stewart CN Jr (2015) Manipulating microRNAs for improved biomass and biofuels from plant feedstocks. Plant Biotechnol J 13(3):337–354. https://doi.org/10.1111/pbi.12319
Tu Y, Rochfort S, Liu Z, Ran Y, Griffith M, Badenhorst P, Louie GV, Bowman ME, Smith KF, Noel JP, Mouradov A, Spangenberg G (2010) Functional analyses of caffeic acid O-methyltransferase and Cinnamoyl-CoA-reductase genes from perennial ryegrass (Lolium perenne). Plant Cell 22:3357–3373. https://doi.org/10.1105/tpc.109.072827
Vanholme R, Storme V, Vanholme B, Sundin L, Christensen JH, Goeminne G, Halpin C, Rohde A, Morreel K, Boerjan W (2012) A systems biology view of responses to lignin biosynthesis perturbations in Arabidopsis. Plant Cell 24:3506–3529. https://doi.org/10.1105/tpc.112.102574
Vanholme R, Cesarino I, Rataj K, Xiao Y, Sundin L, Goeminn G, Kim H, Cross J, Morreel K, Araujo P, Welsh L, Haustraete J, McClellan C, Vanholme B, Ralph J, Simpson GG, Halpin C, Boerjan W (2013) Caffeoyl shikimate esterase (CSE) is an enzyme in the lignin biosynthetic pathway in Arabidopsis. Science 341(6150):1103–1106. https://doi.org/10.1126/science.1241602
Vargas L, Cesarino I, Vanholme R et al (2016) Improving total saccharification yield of Arabidopsis plants by vessel-specific complementation of caffeoyl shikimate esterase (cse) mutants. Biotechnol Biofuels 9:139. https://doi.org/10.1186/s13068-016-0551-9
Vélez-Bermúdez IC, Salazar-Henao JE, Fornalé S, López-Vidriero I, Franco-Zorrilla JM, Grotewold E, Gray J, Solano R, Schmidt W, Pagés M, Riera M, Caparros-Ruiz D (2015) A MYB/ZML complex regulates wound-induced lignin genes in maize. Plant Cell 27(11):3245–3259. https://doi.org/10.1105/tpc.15.00545
Verma SR, Dwivedi UN (2014) Lignin genetic engineering for improvement of wood quality: applications in paper and textile industries, fodder and bioenergy production. S Afr J Bot 91:107–125. https://doi.org/10.1016/j.sajb.2014.01.002
Vignols F, Rigau J, Torres MA, Puigdomenech P (1995) The brown midrib3 (bm3) mutation in maize occurs in the gene encoding caffeic acid O- methyltransferase. Plant Cell 7:407–416
Voelker SL, Lachenbruch B, Meinzer FC, Jourdes M, Ki C, Patten AM et al (2010) Antisense down-regulation of 4CL expression alters lignification, tree growth, and sac-charification potential of field-grown poplar. Plant Physiol 154:874–886. https://doi.org/10.1104/pp.110.159269
Wagner A, Tobimatsu Y, Phillips L, Flint H, Geddes B, Lu F, Ralph J (2015) Syringyl lignin production in conifers: proof of concept in a pine tracheary element system. Proc Natl Acad Sci U S A 112:6218–6223. https://doi.org/10.1073/pnas.1411926112
Walker AM, Hayes RP, Youn B, Vermerris W, Sattler SE, Kang C (2013) Elucidation of the structure and reaction mechanism of sorghum hydroxycinnamoyltransferase and its structural relationship to other coenzyme a-dependent transferases and synthases. Plant Physiol 162(2):640–651. https://doi.org/10.1104/pp.113.217836
Wang H, Li K, Hu X, Liu Z, Wu Y, Huang C (2016) Genome-wide association analysis of forage quality in maize mature stalk. BMC Plant Biol 16:227. https://doi.org/10.1186/s12870-016-0919-9
Wasternack C, Strnad M (2019) Jasmonates are signals in the biosynthesis of secondary metabolites - Pathways, transcription factors and applied aspects - A brief review. N Biotechnol 48:1–11. https://doi.org/10.1016/j.nbt.2017.09.007
Watts KT, Mijts BN, Lee PC, Manning AJ, Schmidt-Dannert C (2006) Discovery of a substrate selectivity switch in tyrosine ammonia-lyase, a member of the aromatic amino acid lyase family. Chem Biol 13:1317–1326. https://doi.org/10.1016/j.chembiol.2006.10.008
Weng JK, Akiyama T, Bonawitz ND, Li X, Ralph J, Chapple C (2010) Convergent evolution of syringyl lignin biosynthesis via distinct pathways in the lycophytes Selaginella and flowering plants. Plant Cell 22:1033–1045. https://doi.org/10.1105/tpc.109.073528
Xia J, Liu Y, Yao S, Li M, Zhu M, Huang K, Gao L, Xia T (2017) Characterization and expression profiling of Camellia sinensis cinnamate 4-hydroxylase genes in phenylpropanoid pathways. Genes (Basel) 8(8):193. https://doi.org/10.3390/genes8080193
Xiong W, Wu Z, Liu Y, Li Y, Su K, Bai Z et al (2019) Mutation of 4-coumarate: coenzyme A ligase 1 gene affects lignin biosynthesis and increases the cell wall digestibility in maize brown midrib5 mutants. Biotechnol Biofuels 12:82. https://doi.org/10.1186/s13068-019-1421-z.158
Xu Z, Zhang D, Hu J, Zhou X, Ye X, Reichel KL, Stewart NR, Syrenne RD, Yang X, Gao P, Shi W, Doeppke C, Sykes RW, Burris JN, Bozell JJ, Cheng MZM, Hayes DG, Labbe N, Davis M, Stewart CN, Yuan JS (2009) Comparative genome analysis of lignin biosynthesis gene families across the plant kingdom. BMC Bioinf 10(Suppl 1):S3. https://doi.org/10.1186/1471-2105-10-S11-S3
Xu B, Escamilla-Treviño LL, Sathitsuksanoh N, Shen Z, Shen H, Zhang YH, Dixon RA, Zhao B (2011) Silencing of 4-coumarate:coenzyme A ligase in switchgrass leads to reduced lignin content and improved fermentable sugar yields for biofuel production. New Phytol 192(3):611–625. https://doi.org/10.1111/j.1469-8137.2011.03830.x
Yang Q, He Y, Kabahuma M, Chaya T, Kelly A, Borrego E, Bian Y, El Kasmi F, Yang L, Teixeira P, Kolkman J, Nelson R, Kolomiets M, Dangl JL, Wisser R, Caplan J, Li X, Lauter N, Balint-Kurti P (2017) A gene encoding maize caffeoyl-CoA O-methyltransferase confers quantitative resistance to multiple pathogens. Nat Genet 49(9):1364–1372. https://doi.org/10.1038/ng.3919
Ye ZH, Kneusel RE, Matern U, Varner JE (1994) An alternative methylation pathway in lignin biosynthesis in Zinnia. Plant Cell 6(10):1427–1439
Youn B, Camacho R, Moinuddin SGA, Lee C, Davin LB, Lewis NG, Kang C (2006) Crystal structures and catalytic mechanism of the Arabidopsis cinnamyl alcohol dehydrogenases AtCAD5 and AtCAD4. Org Biomol Chem 4:1687–1697. https://doi.org/10.1039/b601672c
Zhang K, Qian Q, Huang Z, Wang Y, Li M, Hong L, Zeng D, Gu M, Chu C, Cheng Z (2006) GOLD HULL AND INTERNODE2 encodes a primarily multifunctional cinnamyl-alcohol dehydrogenase in rice. Plant Physiol 140(3):972–983. https://doi.org/10.1104/pp.105.073007
Zhang G, Zhang Y, Xu J, Niu X, Qi J, Tao A, Zhang L, Fang P, Lin L, Su J (2014) The CCoAOMT1 gene from jute (Corchorus capsularis L.) is involved in lignin biosynthesis in Arabidopsis thaliana. Gene. 546:398–402. https://doi.org/10.1016/j.gene.2014.05.011
Zhang J, Zhang X, Tang H, Zhang Q, Hua X, Ma X et al (2018) Allele-defined genome of the autopolyploid sugarcane Saccharum spontaneum L. Nat Genet 50:1565–1573. https://doi.org/10.1038/s41588-018-0237-2
Zhong R, Ye Z-H (2012) MYB46 and MYB83 bind to the SMRE sites and directly activate a suite of transcription factors and secondary wall biosynthetic genes. Plant Cell Physiol 53:368–380. https://doi.org/10.1093/pcp/pcr185
Zhong R, Morrison WH, Himmelsbach DS, Poole FL, Ye Z-H (2000) Essential role of caffeoyl coenzyme A o-methyltransferase in lignin biosynthesis in woody poplar plants. Plant Physiol 124:563–577
Zhong R, Lee C, Ye Z-H (2010) Global analysis of direct targets of secondary wall NAC master switches in Arabidopsis. Mol Plant 3:1087–1103. https://doi.org/10.1093/mp/ssq062
Zhou R, Jackson L, Shadle G, Nakashima J, Temple S, Chen F, Dixon RA (2010) Distinct cinnamoyl CoA reductases involved in parallel routes to lignin in Medicago truncatula. Proc Natl Acad Sci 107:17803–17808. https://doi.org/10.1073/pnas.1012900107
Ziebell A, Gjersing EL, Hinchee MA, Katahira R, Johnson DK, Davis MF (2016) Down-regulation of ρ-coumaroyl quinate/shikimate 3′-hydroxylase (C3ˊH) or cinnamate 4-hydroxylase (C4H) in Eucalyptus urophylla 9 Eucalyptus grandis leads to increased extractability. Bioenergy Res 9:691–699. https://doi.org/10.1186/s13068-015-0316-x
Acknowledgments
This work is part of the PhD thesis of Thais Felix-Cordeiro. DJM was supported by Post-doctoral fellowship CAPES-PNPD. TFC was supported by CAPES PhD fellowship. YSV, VG, GVA, and YRAA were recipient of CNPq PIBIC fellowship.
Funding
This work was supported by grants from the Conselho Nacional de Desenvolvimento Científico e Tecnológico – CNPq (grant number 474911/2012-8); Coordenação de Aperfeiçoamento de Pessoal de Nível Superior - CAPES (PNPD N ° 02691/09-4 - linha MEC / CAPES); Fundação Carlos Chagas Filho de Amparo à Pesquisa do Estado do Rio de Janeiro – FAPERJ (E26/111.234/2014); and Petrobras (grant number ANP 01070 0050.0076123.12.9) to GSM.
Author information
Authors and Affiliations
Contributions
DJM and GSM conceived and designed the experiments.
DJM, TFC, LB, YSV, VG, GAB, GVA, YRAA, AFR, RLC, and GSM performed the experiments.
DJM, LB, RLC, and GSM analyzed the data.
DJM and GSM wrote the manuscript.
All authors read and approved the manuscript.
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Ethics approval
This article does not contain any studies with human participants or animals performed by any of the authors.
Additional information
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
About this article
Cite this article
Jardim-Messeder, D., Felix-Cordeiro, T., Barzilai, L. et al. Genome-wide analysis of general phenylpropanoid and monolignol-specific metabolism genes in sugarcane. Funct Integr Genomics 21, 73–99 (2021). https://doi.org/10.1007/s10142-020-00762-9
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10142-020-00762-9