Abstract
Echeveria ‘Perle von Nürnberg’ is a popular ornamental potted plant. In 2017–2018, stem rot lesions caused by Fusarium spp. were repeatedly observed on Echeveria ‘Perle von Nürnberg’ in Zhangzhou, Fujian Province, China. In this study, samples were collected to isolate the pathogen. After morphological observations, molecular identification, and pathogenicity measurements, the pathogen was confirmed as Fusarium oxysporum. Green fluorescent protein was used to label F. oxysporum to observe the pathogenic process in the host plant. Two to three days after inoculation, the pathogen conidia adhered to the surface of the host stem and germinated into hyphae. The hyphae invaded the host cortex through wounds on the plant or the stem-leaf junction. The host stem produced greyish-brown circular or irregular lesions. Five days after inoculation, the expanded hyphae invaded the vascular cylinder and proliferated. The lesions on the host stem became larger and the neighboring leaves gradually fell off. Seven days after inoculation, the thalli continued to proliferate in the host, the hyphae continued to expand, and the host pith was damaged. The lesions on the stem continued to expand; the stems and branches became withered and lodged, and numerous leaves fell off. Given the rapid infection of F. oxysporum in Echeveria ‘Perle von Nürnberg’ plants, necessary prevention and control measures should be employed prior to infection. In summary, this study demonstrated that the pathogen responsible for stem rot disease in Echeveria ‘Perle von Nürnberg’ is F. oxysporum and further revealed the invasion modes and pathways of F. oxysporum in this plant, providing an empirical basis for the diagnosis and treatment of this stem rot pathogen.
Similar content being viewed by others
Introduction
Succulents are plants featuring thick and succulent organs (i.e., roots, stems, and leaves) that can store large amounts of water (Griffiths and Males 2017). Petite, interesting and cute, succulents have become popular ornamental plants in offices and public spaces (Wang et al. 2009). In 2016, the cultivation area of Echeveria spp. in China was 53.33 hm2 and the annual yield was 300 million pots (Ke and Tang 2016). In 2017 and 2018, the cultivation area of Echeveria spp. increased drastically in China, with annual yields increasing to 400 and 500 million pots, respectively (Wang et al. 2018; Yan 2018). Echeveria ‘Perle von Nürnberg’ has great economic value. Also known as purple pearl, and its beautiful pink and purple lotus constellation-shaped flowers make it a consumer favorite. It is typically propagated by removing fresh branch and leaf cuttings and allowing them to callous over for a few days, after which they are placed directly into substrate. However, this cultivation process makes Echeveria ‘Perle von Nürnberg’ susceptible to pathogen infection. Disease is one of the biggest threats to succulents such as Echeveria ‘Perle von Nürnberg’ and often occurs as a result of shed cultivation conditions, such as high temperature and humidity and poor ventilation (Wang et al. 2009). In greenhouse plantings, some Echeveria ‘Perle von Nürnberg’ plants exhibit greyish-brown lesions, shedding of neighboring leaves, or stem rot and withering. In some severe cases, the entire stem withers completely. It is important it identify the pathogen causing stem rot disease in Echeveria ‘Perle von Nürnberg’.
The identification of Fusarium species involves systematic morphological identification. However, relying on morphological characteristics alone to identify Fusarium tends to result in errors, producing uncertain results and affecting subsequent research. Currently, molecular techniques are being used in the identification of Fusarium species. Commonly used DNA sequences include internal transcribed spacer (ITS) regions, intragenic spacer (IGS) regions, actin genes, and translation elongation factor-1α (EF-1α) genes. These sequences can be used to effectively differentiate between species, variants, and geographical isolates (Cabral et al. 2018; Liu et al. 2012; Silva et al. 2014; Wang et al. 2018; Yao et al. 2018). A systems approach combining morphology and molecular biology, enables the rapid and reliable identification of Fusarium species.
The genus Fusarium contains some species that are parasitic and infect the vascular cylinder system of plants. Species of Fusarium can cause diseases that are difficult to control and prevent (Darnetty and Salleh 2017; Sarrocco et al. 2007). To effectively control and prevent stem rot disease, it is necessary to understand the host pathogenesis of Fusarium in order to apply suitable agricultural measures. However, observing Fusarium attachment, germination, invasion, and expansion by conventional observation methods is tedious. It is therefore necessary to utilize an effective real-time dynamic tracking technique to analyze infection and pathogenesis (Yao et al. 2018, 2019). Green fluorescent protein (GFP) is widely used for studying the pathogenesis of plant pathogens (Yang et al. 2019; Yao et al. 2019) and has been successfully used to label F. oxysporum growing within host plants such as banana (Visser et al. 2004; Zhang 2007), musk melon (Nonomura et al. 2003), and carnation (Sarrocco et al. 2007). However, these aforementioned hosts are herbs with structural features that are characterized by an external stem layer with a high fiber content and tough mechanical tissue. Typically, the vascular tissue of plants contains a large number of tightly packed parenchyma cells (Neuhaus 2013). However, the structural characteristics of Echeveria spp. differ greatly from the aforementioned herbs and are unique in that there is a tender epidermis and thick and succulent roots, stems, and leaves that can store large volumes of water. Furthermore, Echeveria spp. develop vascular tissues lacking tough mechanical tissue and with a loose parenchyma cell arrangement. It is thus unclear whether the application of GFP can sufficiently elucidate the patterns or the characteristics of Fusarium infection in host plants with unique structures, such as Echeveria spp.
In this study, a combination of morphological and molecular identification techniques was employed to determine the type of stem rot disease pathogen in Echeveria ‘Perle von Nürnberg’. GFP labeled Fusarium was used to reveal the pathogenicity characteristics of the Echeveria ‘Perle von Nürnberg’ stem rot disease.
Materials and methods
Sample collection
In June and August of 2017, and in April, June, and November of 2018, 75 diseased samples were obtained from plants with stem rot at three cultivation greenhouses in Zhangzhou, Fujian Province, China, and the incidence of stem rot disease was investigated. The sites were located in Jiuhu Town (124°29′3.73″N, 117°38′40.20″E), Chengxi Town (24°23′33.89″N, 117°35′9.34″E), and Guanxun Town (24°19′19.26″N, 117°44′52.29″E) (Fig. 1a). Small pieces of stem tissues (0.5 cm × 0.5 cm) were surface-disinfested with 1% sodium hypochlorite for 5 min and rinsed three times on sterile water. The tissues were cultured in potato dextrose agar (PDA) medium for 5 d at 28 °C (Yao et al. 2018). One hundred and five isolates that showed similar morphological characteristics were obtained (Table 1).
Field symptoms, pathogen morphological characteristics, and pathogenicity of “Echeveria ‘Perle von Nürnberg” plants infected with stem rot pathogen. a Plants infected with stem rot pathogen in the field exhibit symptoms such as stem rot and falling leaves; b morphological characteristics of the colonies after 7 days of growth on potato dextrose agar (PDA) medium; c macroconidia (scale bar 20 μm); d microconidia (scale bar 20 μm); e chlamydospores (scale bar 20 μm); f plant inoculation [left: non-wounding inoculation (CK), right: wounding inoculation, j shows the inoculation site]; g comparison of CK and treated plants on 5 days post inoculation (DPI) (left: CK, right: inoculated plant); h At 5 DPI, lesions on the stem become larger and the neighboring leaves were infected and gradually fell off; i comparison of CK and treated plants on 7 DPI (left: CK, right: inoculated plant, k shows the stem lodging point)
Plant materials
Healthy one-year-old plants in pots with flat leaves and trauma-free stems were selected from the Jiuhu Town cultivation site and brought back to the laboratory. Sterile water was used to gently wash the stems and leaves of Echeveria ‘Perle von Nürnberg’. After 15 d, the plants were used for pathogenicity measurements and observation of host invasion.
Materials for GFP insertion
The transformation vector pCAMBIA1300 − ptrpC−hph − gfp and transformation carrier Agrobacterium AGL-1 strain were obtained from the Agricultural Bioresources Research Institute, Fujian Academy of Agricultural Sciences (Fuzhou, Fujian, China). Various antibiotics (hygromycin B, kanamycin, and timentin) were purchased from Sigma-Aldrich Inc. (St, Louis, MO, USA). The storage solution and minimal medium (MM), induction medium (IM), and complete medium (CM) used in the experiments were prepared according to the method of Yao et al. (2019).
Pathogen identification
Pathogen purification
The colony color, hyphal growth status and colony characteristics of the 105 pathogenic isolates were not different. Nine representative isolates were randomly selected according to the year and cultivation site. Monospores of these isolates were purified and renumbered from FJVP-1 to FJVP-9 according to a previously described method (Unnithan and Thammaiah 2017).
Morphological characterization
The purified isolates (FJVP-1 to FJVP-9) were inoculated on PDA plates and cultured in a light incubator (Xinmiao Instrument Manufacturing Co., Ltd., Shanghai, China) at 28 ± 0.5 °C with a light cycle of 16 L: 8 D for 7 d. The colony color and hyphal growth status were then observed. After 15 d, the conidia morphological characteristics of the isolates (FJVP-1 to FJVP-9) were observed under a CX51 optical microscope (Olympus, Japan). Twenty spores were observed in each microscopic field, and five fields were observed in total for each isolate (Bechem and Afanga 2018; Lin et al. 2018).
Molecular identification
The methods of Yao et al. (2018) and Tian et al. (2018) were used as a reference to extract the genomic DNA from monospores of the nine monospore isolates (FJVP-1 to FJVP-9). The rDNA-ITS universal primers (ITS1/ITS4), actin gene primers (ACT-512F/ACT-783R), and EF-1α gene primers (EF1/EF2) were used for PCR amplification of the genomic DNA of the pathogen (White et al. 1990; Carbone and Kohn 1999; O'Donnell et al. 1998). The rDNA-ITS PCR reaction conditions were as follows: pre-denaturation at 94 °C for 5 min; followed by 30 cycles of denaturation at 94 °C for 1 min, annealing at 55 °C for 1 min, and extension at 72 °C for 1.5 min, before a final extension at 72 °C for 8 min. The annealing temperatures for actin and EF-1α were 58 °C and 56 °C, respectively, and the remaining reaction conditions were identical to rDNA-ITS. The single band of PCR amplifications was cut out from the agarose gel and purified using the agarose gel DNA fragment recovery kit [Tiangen Biotech (Beijing, China) Co., Ltd] following the manufacturer’s protocol and ligated into the plasmid pMD 18-T vector [TaKaRa Biotech (Dalian, China) Co., Ltd], the reactions (5.5 μL) contained 2.5 μL of PCR product (80 ng), 2.5 μL Ligation Mix [SolutionI, TaKaRa Biotech (Dalian, China) Co., Ltd], 0.5 μL pMD18-T Vector (25 ng), at 16 °C over 4 h and followed at 4 °C overnight.. The ligated DNA was transformed into competent cells of Escherichia coli DH5α. The positive clones were selected on ampicillin agar and screened by colony PCR for the presence of insert. Inserts from positive colonies were sequenced in both forward and reverse directions using ITS1/ITS4, ACT-512F/ACT-783R and EF1/EF2 primers by Sangon (Shanghai Sangon Biotech Co., Ltd). NCBI BLAST (https://blast.ncbi.nlm.nih.gov/) and MEGA 6.0 software (https://www.megasoftware.net) were used for homology and phylogenetic analysis of the sequencing results. To further reveal the taxonomic status of the fungal isolates, their rDNA-ITS, actin, and EF-1α gene sequences were aligned separately using the MAFFT program (Katoh and Standley 2013) and concatenated subsequently using PhyloSuite 1.2.1 (Zhang et al. 2020). Phylogenetic analysis was carried out using MrBayes 3.2.6 (Ronquist et al. 2012) implemented in PhyloSuite under the best data partitioning schemes, which were determined using PartitionFinder 2 (Lanfear et al. 2017).
Pathogenicity
The FJVP-6 and FJVP-9 isolates were used to determine pathogenicity measurements according to the method of Yao et al. (2018). Both wounding and non-wounding inoculations on the stems of host plants (Echeveria ‘Perle von Nürnberg’ plants) were employed. For wounding inoculation: three healthy leaves were randomly selected from the lower part of the plant, and the upper epidermis of each leaf was punctured with a sterile needle. Mycelial discs (5 mm diameter) removed from a 5-day-old colony of isolate FJVP-6 or FJVP-9 were placed on the wounds on the leaf. and a sterile PDA agar block was used as a control. The inoculation site was then wrapped with sterile water-soaked cotton to retain moisture. The process was the same for the non-wounding inoculation, except that the epidermis of the leaves was not punctured. Ten host plants were inoculated for each treatment, with three replicates tested. The plants were all maintained at 90% relative humidity with a 12-h photoperiod at 25 °C in climate chambers. Within 7 days post inoculation (DPI), signs of disease were observable in the stems of the host plants, and the pathogens were isolated and purified from hosts that demonstrated classical symptoms. There was an attempt to isolate and purify from control plants. Morphological observations and molecular identification were then carried out. The experiments were performed three times.
Observation of the host infection process
GFP transformation and culture of the pathogen
The purified FJVP-6 isolates (monospores) were randomly selected, and the method of Yao et al. (2019) was used for GFP transformation and culture to obtain 25 pathogenic transformants.
Transformant genetic stability and pathogenicity measurements. Six purified transformants were randomly selected (monospores, named PTM-1 to PTM-6) and subjected to genetic stability assays as described by Yao et al. (2019). Three tenth-generation transformants (PTM-1, PTM-4, and PTM-5) were randomly selected, and wounding and non-wounding inoculations were performed for each transformant. The wild-type FJVP-6 strain was used as a control. Host plants were inoculated with mycelial discs (5 mm diameter) removed from a 5-day-old colony of isolate PTM-1, PTM-4, and PTM-5 and the sterile PDA agar block was used as a control. Wounding and non-wounding inoculations were each used to inoculate three host plants for each transformant, with three replicates (i.e., nine plants for one treatment). Control plants were inoculated using similar methods. Within 10 DPI, the stems of the host plants were observed for disease symptoms, and pathogens were isolated and purified from hosts demonstrating typical stem rot disease symptoms. Morphological observations and molecular identification were carried out, and incidence was determined. The experiments were performed three times.
Observation of the host infection process of the pathogen. Tenth-generation transformants (PTM-1, PTM-4, and PTM-5) were selected. Host plants were inoculated with mycelial discs (5 mm diameter) removed from a 5-day-old colony of isolate PTM-1, PTM-4, and PTM-5 and the sterile PDA agar block was used as a control. Wounding and non-wounding inoculations were each used to inoculate 30 host plants for each transformant. Control plants were inoculated using similar methods. At 1, 2, 3, 5, and 7 DPI, three host plants from each treatment were randomly selected for sectioning. Stem sections of 0.5 cm were sliced, starting from the base of the stem and continuing upwards. Slides exhibiting the best observation results under the IX73 fluorescence microscope (Olympus, Japan) were selected for observation and photography using a TCS SP5 laser scanning confocal microscope (Leica, Germany) to examine the host infection process by the pathogen. The experiments were performed three times.
Results
Field investigation
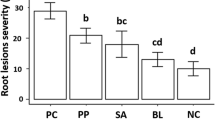
A survey of the Echeveria ‘Perle von Nürnberg’ cultivation areas in three towns of Fujian Province, China, indicated that some plants exhibited greyish-brown lesions, shedding of neighboring leaves, or stem rot and withering. In some severe cases, the entire stem had withered completely. According to the survey, in June 2017, August 2017, April 2018, June 2018, and November 2018, the average incidences of disease in the three towns were 29.8%, 35.3%, 35.6%, 37.6%, and 37.1%, respectively, suggesting that the disease may be increasing. Seventy-five diseased samples were obtained from plants with stem rot.
Pathogen identification
Morphological characterization
The result showed that the growth rate of the isolates (FJVP-1 to FJVP-9) was relatively fast on the PDA culture medium, reaching 83.6–85.4 mm in diameter after 7 d of culture, with dense aerial mycelia. The colonies appeared flocculent and were pale purplish-white in color (Fig. 1b). After 15 d of growth, the isolates produced macroconidia, microconidia, and chlamydospores. The macroconidia generally were sickle-shaped, with some appearing curved. The macroconidia had 2–6 septae and were 12.6–39.4 μm in length × 3.5–5.0 μm in diameter (Fig. 1c). The microconidia were produced on solitary phialides and typically formed spherical clumps at the tip of phialides. The microconidia were single-celled and oval-shaped, and were 3–12 μm in length and 2–3.5 μm in diameter (Fig. 1d). The chlamydospores were intercalary or terminal, spherical, with a smooth surface and thick walls, and were 4.2–14.3 μm in length × 4.4–11.7 μm in diameter (Fig. 1e). According to the taxonomic system of Booth (1971), these morphological characteristics were similar to F. oxysporum and different from other Fusarium species.
Molecular analyses
Molecular identification was used to further classify the isolates. Fragments of about 500 bp, 250 bp, and 700 bp of the rDNA-ITS, actin gene, and EF-1α gene, respectively, of the nine isolates (FJVP-1 to FJVP-9) were amplified. After sequencing and alignment, the related sequences were identical, revealing that the nine representative isolates were the same species. The rDNA-ITS, actin gene, and EF-1α gene sequences of FJVP-6 and FJVP-9 were submitted to NCBI/GenBank (www.ncbi.nlm.nih.gov/genbank/) and accession numbers were obtained: FJVP-6 (Accession No. MG825179, MH511658 and MK810784) and FJVP-9 (Accession No. MG825180, MH511657 and MK810785).
NCBI/BLAST (blast.ncbi.nlm.nih.gov/Blast.cgi) was used to align the rDNA-ITS, actin gene, and EF-1α gene sequences of the two isolates, and the results indicated that they shared 100% similarity with the ITS sequences (Accession No. MK416124, MK250067) and actin gene sequences (Accession No. MK001023, LR131915) of F. oxysporum, while the EF-1α gene sequences (Accession No. KF574851, LT970767) had similarities of 99.85% and 99.42%, respectively. There were four bases that differed from the LT970767 strain (183 T → C, 252C → T, 322 base deletion→T, and 676C → G), and one base that differed from KF574851 (676 base deletion→T). The rDNA-ITS, actin and EF-1α sequences were subjected to Phylogenetic analysis (Table 2; Fig. 2). Using previously-published sequences in NCBI/GenBank, it was found that the FJVP-6 and FJVP-9 isolates clustered along the same branch with F. oxysporum (Accession No. MH221085, KY798316, KY798315, MK949084, MK949085, MK880499). These results indicated that the FJVP-6 and FJVP-9 isolates were F. oxysporum.
Pathogenicity
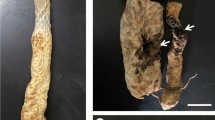
Monospore isolates of FJVP-6 and FJVP-9 were inoculated onto Echeveria ‘Perle von Nürnberg’ plants using wounding and non-wounding methods (Fig. 1f). The results showed that after 2 d after wounding inoculation, the epidermis of the inoculated stem started to rupture and produced greyish-brown circular or irregular lesions. Disease onset was delayed by 1 d in plants that received non-wounding inoculation, and lesions were visible at 3 DPI. At 5 DPI, the lesions on the host stem became larger, and the neighboring leaves gradually became infected and were shed (Fig. 1g and h). At 7 DPI, the host pith was damaged, and the greyish-brown lesions on the stem continued to expand, the stems and branches became withered and lodged, and a large number of leaves fell off, which was very similar to the natural disease symptoms on plants grown in the field (Fig. 1i). The host disease incidence was 100% after inoculation. No disease occurred in the host plants that were inoculated with the PDA agar block control. Hosts with typical disease symptoms were again used for isolation and purification of pathogens, and morphological observations and molecular identifications were carried out. The results showed that the re-isolated strain was similar to the inoculated strain. We did attempt to isolate and purify from control plants, but no pathogens were isolated from the control plants. The experiment was performed three times, and similar results were obtained. From these results, the pathogen causing Echeveria ‘Perle von Nürnberg’ stem rot disease was confirmed to be F. oxysporum.
Observation of the host infection process
Pathogen GFP transformation and culture
Through Agrobacterium-mediated transformation, the hyphae and conidia of 25 pathogenic transformants were able to emit stable green fluorescence (Fig. 3) under excitation by a 480 nm blue laser. Therefore, the GFP gene was successfully integrated into the genome of FJVP-6 and was expressed.
Transformant genetic stability and pathogenicity
Six transformants underwent monospore purification (PTM-1 to PTM-6) and were cultured for 10 generations. On PDA medium, the tenth generation transformants had hyphal growth and colony morphology similar to the wild-type. Hyphal growth was dense and purplish-white or pale purplish-white. Hygromycin resistance was stable in the transformants. Under fluorescence microscopy, the hyphae and conidia of the tenth generation of transformants could stably emit green fluorescence. Therefore, the GFP gene was not only successfully integrated into the genome of the F. oxysporum FJVP-6 strain, but also exhibited genetic stability. The tenth generation of transformants (PTM-1, PTM-4, and PTM-5) and the wild-type FJVP-6 strain were inoculated into Echeveria ‘Perle von Nürnberg’ plants, and similar stem rot disease and symptoms were produced: the stems showed greyish-brown lesions, the neighboring leaves fell off, and the stems gradually withered and lodged (Fig. 4). The host disease incidence was 100% after isolate inoculation. No disease occurred in the host plants that were inoculated with the PDA agar block control. The pathogen was re-isolated from inoculated plants and showed very similar morphological characteristics to the wild-type FJVP-6 strain. The experiment was performed three times and similar results were obtained. This indicated that the GFP-labeled Echeveria ‘Perle von Nürnberg’ stem rot pathogen F. oxysporum retained similar pathogenicity to the wild-type strain. This confirmed that the GFP gene had been successfully transferred into the FJVP-6 strain, its genetic characteristics were stable, and its ability to cause infection was not affected. Therefore, the labeled strain could be used to further study the mechanism and process of pathogen infection.
Symptoms from 5 to 10 days post inoculation (DPI) when the wild-type and green fluorescent protein (GFP)-labeled FJVP-6 F. oxysporum were inoculated on Echeveria ‘Perle von Nürnberg’ plants. A: 5 DPI; B: 6 DPI; C: 7 DPI; D: 8 DPI; E: 9 DPI; F: 10 DPI. Notes: Capital letters indicate plants inoculated with the wild-type strain, while small letters indicate plants inoculated with the GFP-labeled strain
Host plant infection process of the pathogen
To directly observe the infection process of the pathogen, the stems of Echeveria ‘Perle von Nürnberg’ plants that were infected with the tenth generation of transformants were cut horizontally (Fig. 5Α and B), and laser confocal microscopy was used to observe the host plant infection process by GFP-labeled F. oxysporum. The results showed that at 1 DPI and 2 DPI by the wounding method, large numbers of mycelia germinate around the wound of the infected plant, and they expanded in an organized manner to form a hyphal network. The hyphae invaded the host cortex through wounds on the plant and the cortical tissues were significantly damaged (Fig. 5C). The infection process in the non-wounding inoculated plants was slightly slower, and epidermis-germinated hyphae had just started to invade the cellular spaces in the plants at 2 DPI (Fig. 5D). At 3 DPI of non-wounding inoculation, epidermis-germinated hyphae preferentially expanded at the stem-leaf junction to form a hyphal network. The hyphae invaded the host cortex from the stem-leaf junction (Fig. 5E). The epidermal and cortical layers of the stem were separated to form cavities. Some of the epidermis was ruptured (Fig. 5F). At 5 DPI, the expanded hyphae invaded the vascular bundle and proliferated in the gaps in the vascular cylinder and parenchyma cells. During repeated hyphal invasion of the vascular cylinder, the vascular system was severely damaged (Fig. 5G and H). At 7 DPI, the thalli continued to proliferate in the host, and the hyphae continued to expand horizontally and vertically (Fig. 5I and J). The thalli preferentially expanded along the growth direction in the vascular tissues and parenchyma cells, and the host pith was largely damaged (Fig. 5K). The experiment was repeated three times and similar results were obtained.
Stem microscopic characteristics when green fluorescent protein (GFP)-labeled F. oxysporum was used to infect Echeveria ‘Perle von Nürnberg’ plants. A Echeveria ‘Perle von Nürnberg’ pathogen that was GFP-labeled. B horizontally cut stem of infected Echeveria ‘Perle von Nürnberg’ plants. C on day 2 of wounding inoculation, large numbers of germinating hyphae were concentrated around the wound in infected plants, and they expanded to form a hyphal network. The hyphae invaded the host cortex through wounds on the plant, and the cortical tissues were significantly damaged. D on day 2 of non-wounding inoculation, the epidermis-germinated hyphae started to invade the cellular spaces in the plants. E on day 3 of non-wounding inoculation, the epidermis-germinated hyphae preferentially expanded at the stem-leaf junction to form a hyphal network. The hyphae invaded the host cortex from the stem-leaf junction. F the epidermal and cortical layers of the stem were separated to form cavities. Some of the epidermis was ruptured, and the thallus exhibited vigorous growth inside the cortex. G and H At 5 days post inoculation, the expanded hyphae invaded the vascular bundle and proliferated in the gaps in the vascular cylinder and parenchyma cells. I and J At 7 days post inoculation, the thallus continued to proliferate in the host, and the hyphae continued to grow horizontally and vertically. K the thallus preferentially expanded along the growth direction in the vascular cylinder, and the parenchyma cells and host pith were damaged. Note: capital letters E–K indicate the visual field under blue excitation light, while the small letters e–k represent the bright field
Discussion
In recent years, stem rot disease has increasingly hindered the large-scale cultivation of Echeveria ‘Perle von Nürnberg’. Stem rot disease has a rapid onset and strong transmissibility and has become one of the major diseases affecting this plant. There is thus an urgent need for the prevention and control of this disease. Plant-pathogenic Fusarium species are widely distributed globally. Fusarium is also a complex genus with obvious individual species polymorphisms, different infective and pathogenic abilities, and high susceptibility to variation with environment. Its accurate identification remains challenging (Booth 1971; Ibrahim et al. 2017). In the current study, an Echeveria ‘Perle von Nürnberg’ stem rot pathogen was isolated, and purified. The morphological oobservations, molecular identification, and pathogenicity measurements, mutually validated each other. The results confirmed that this disease was caused by F. oxysporum infection. This is the first report that stem rot disease in Echeveria ‘Perle von Nürnberg’ is caused by F. oxysporum. It is important to determine the pathogenicity and characteristics so as to formulate effective prevention and control protocols.
Fusarium oxysporum is a parasite of the vascular cylinder system in plants and can proliferate and spread in infected plants and disrupt host tissue structure (Darnetty and Salleh 2017; Sarrocco et al. 2007). A detailed examination of the initiation and expansion process during host infection and a thorough understanding of the crucial time-points for pathogen infection are extremely important for preventing and controlling Echeveria ‘Perle von Nürnberg’ stem rot disease. However, determining the crucial time-points of F. oxysporum infection in Echeveria ‘Perle von Nürnberg’ during growth by physical observation is challenging. This is mainly because Echeveria ‘Perle von Nürnberg’ plants are thick and succulent and do not clearly demonstrate symptoms of dehydration and wilting, which masks the early-stage symptoms of the disease. When the leaves have fallen and lodging has occurred, the disease is in an advanced stage and cannot be controlled or treated. Therefore, there is a need to develop a simple, intuitive, and effective real-time dynamic tracking technique to adequately evaluate the infection and pathogenesis of F. oxysporum. The GFP marker has been widely used in labeling F. oxysporum to observe its infection and pathogenesis in bananas, musk melons, flax, and carnations (Blum et al. 2018; Nonomura et al. 2003; Sarrocco et al. 2007; Visser et al. 2004; Zhang 2007). In the current study, the GFP labelling results showed that F. oxysporum infects Echeveria ‘Perle von Nürnberg’, exhibits strong genetic stability, and has similar growth characteristics and pathogenicity to the wild-type strain. This is the first report on the use of GFP gene labeling to observe the infection process of F. oxysporum on Echeveria ‘Perle von Nürnberg.’ This method revealed the infection modes and pathways of F. oxysporum in Echeveria ‘Perle von Nürnberg’. Previous studies showed that the infection rate of F. oxysporum was relatively slow. For example, F. oxysporum can infect Lilium (Zhang et al. 2018), Cymbidium ensifolium (Yao et al. 2018), Cyclamen (Kim et al. 2003), Euphorbia pulcherrima (Orlikowski and Ptaszek 2013), Chrysanthemum (Singh and Kumar 2011), Eustoma grandiflorum (Li et al. 2010), and Dianthus caryophyllus (Ardila et al. 2011). These hosts are herbaceous plants, and it takes at least 30 d or even 2–3 months from infection to plant death. The present study found that the speed of Echeveria ‘Perle von Nürnberg’ infection by F. oxysporum was significantly faster than that in herbaceous plants, as the fungus was able to invade the host cortex through wounds or stem-leaf junctions within 2–3 days and then proliferate in the plant. After 5–7 DPI, the pathogen had invaded the vascular cylinder and host pith, causing leaves to fall and lodging to occur. This may be primarily due to the stem structure of the host plant. The outer layer of the stems of herbaceous plants constitutes tough mechanical tissue, the parenchyma cells in the vascular cylinder are arranged tightly, and the water content is relatively low. These factors greatly limit the growth, proliferation, and spread of F. oxysporum in plants (Neuhaus 2013; Visser et al. 2004). When GFP-labeled F. oxysporum was used to infect these herbs, the hyphae were confined within the vascular cylinder by the endodermal cells beginning from the zone of differentiation of the vascular tissues and were able to grow inside the vessels (Visser et al. 2004). In contrast, Echeveria ‘Perle von Nürnberg’ is a succulent plant with a unique stem tissue structure that, unlike herbs, has developed vascular tissues lacking tough mechanical structures, a tender epidermis, thick and succulent tissues, and an ability to store large amounts of water. Hence, F. oxysporum can proliferate vigorously in the plant cortex and rapidly proliferate and spread in the spaces between the vascular cylinders and parenchyma cells. This study showed that the infection characteristics of F. oxysporum in the succulent tissues of Echeveria ‘Perle von Nürnberg’ were different from those of herbs, demonstrating the infection characteristics of F. oxysporum in these tissues for the first time. In view of the rapid infection of F. oxysporum in Echeveria ‘Perle von Nürnberg’ plants, the necessary prevention and control measures should be employed 2–3 d before infection. This is a crucial timepoint for preventing and controlling Echeveria ‘Perle von Nürnberg’ stem rot disease, as missing this timepoint will result in significant yield losses. Additionally, this study found that the key site of infection of Echeveria ‘Perle von Nürnberg’ plants by F. oxysporum were the wounds on the stem and at the stem-leaf junction. Considering that leaf cutting propagation is usually used for the production of Echeveria ‘Perle von Nürnberg’ plants, which causes wounding, it is recommended that wounds be disinfected after leaf cutting, followed by culture in sterile culture medium for 5–7 d. The cuttings should be used for cultivation after the wounds have healed and new roots have grown in order to reduce F. oxysporum infection in progeny plants.
Conclusion
This is the first study to demonstrate that F. oxysporum is the pathogen responsible for Echeveria ‘Perle von Nürnberg’ stem rot disease. It is also the first to report that GFP gene labeling can be used to reveal the infection and pathogenesis of F. oxysporum in Echeveria ‘Perle von Nürnberg.’ This study showed that the infection speed of this pathogen in Echeveria ‘Perle von Nürnberg’ proceeds faster than in herbaceous species, as the fungus can invade the host cortex through wounds or at the stem-leaf junction within 2–3 d and proliferate in the plant. After 5–7 DPI, the pathogen invaded the vascular cylinder and host pith, causing many leaves to fall and lodging to occur. The key infection sites of F. oxysporum were found to be stem wounds and stem-leaf junctions. This work provides a basis for the diagnosis and prevention of stem rot disease in succulent plants with unique structures.
References
Ardila, H. D., Martínez, S. T., & Higuera, B. L. (2011). Regulación espacio-temporal de fenilalanina amonio liasa en clavel (Dianthus caryophyllus L.) durante su interacción con el patógeno Fusarium oxysporum f. sp. dianthi. Revista Colombiana de Quimica, 40(1), 7–24.
Bechem, E. T., & Afanga, Y. A. (2018). Morphological and molecular identification of fungi associated with corm rot and blight symptoms on plantain (Musa paradisiaca) in macro-propagators. International Journal of Biological Sciences, 11(6), 2793–2808.
Blum, A., Bressan, M., Zahid, A., Trinsoutrot-Gattin, I., Driouich, A., & Laval, K. (2018). Verticillium wilt on fiber flax: Symptoms and pathogen development in planta. Plant Disease, 102(12), 2421–2429.
Booth, C. (1971). The genus Fusarium. Kew, Surrey, England: CMI Press.
Cabral, C. S., de N. Fonseca, M. E., Brunelli, K. R., Rossato, M., Costa, H., Boiteux, L. S., and Reis, A. (2018). Relationships among Brazilian and worldwide isolates of Fusarium oxysporum f. sp. lactucae race 1 inferred from ribosomal intergenic spacer (IGS-rDNA) region and EF-1α gene sequences. European Journal of Plant Pathology, 152(1), 81–94.
Carbone, I., & Kohn, L. M. (1999). A method for designing primer sets for speciation studies in filamentous ascomycetes. Mycologia, 91(3), 553–556.
Griffiths, H., & Males, J. (2017). Succulent plants. Current Biology, 27(17), R890–R896.
Ibrahim, N. F., Mohd, M. H., Mohamed Nor, N. M. I., & Zakaria, L. (2017). Characterization of Fusarium spp. associated with pineapple fruit rot and leaf spot in peninsular Malaysia. Journal of Phytopathology, 165(11–12), 718–726.
Katoh, K., & Standley, D. M. (2013). MAFFT multiple sequence alignment software version 7: Improvements in performance and usability. Molecular Biology and Evolution, 30, 772–780.
Ke, Q. H., & Tang, S. B. (2016). The development status and analysis of succulent plant in China. Xiandai Horticulture, 39–42 (in Chinese).
Kim, J. Y., Kim, H. G., Hong, S. S., Kim, J. W., & Park, K. Y. (2003). Occurrence of Fusarium wilt on Cyclamen casued by Fusarium oxysporum f. sp. cyclaminis and selection of resistant cultivars. Research in Plant Disease, 9(2), 79–84.
Lanfear, R., Frandsen, P. B., Wright, A. M., Senfeld, T., & Calcott, B. (2017). PartitionFinder 2: New methods for selecting partitioned models of evolution for molecular and morphological phylogenetic analyses. Molecular Biology and Evolution, 34, 772–773.
Li, Y., Garibaldi, A., & Gullino, M. L. (2010). Genetic variability analysis and molecular detection of Fusarium oxysporum f. sp. eustomae isolated from Eustoma grandiflorum in northern Italy. Journal of Phytopathology, 158(7–8), 546–553.
Lin, S., Taylor, N. J., & Peduto Hand, F. (2018). Identification and characterization of fungal pathogens causing fruit rot of deciduous holly. Plant Disease, 102(12), 2430–2445.
Liu, J., Zhang, Q., Chang, Q., Zhuang, H., Huang, L. L., & Kang, Z. S. (2012). Cloning and characterization of the actin gene from Puccinia striiformis f. sp. tritici. World Journal of Microbiology and Biotechnology, 28(6), 2331–2339.
Neuhaus, G. (2013). Strasburger's plant sciences: Including prokaryotes and Fungi (pp. 129–159). Berlin, Heidelberg: Springer, Berlin Heidelberg.
Nonomura, T., Tajima, H., Kitagawa, Y., Sekiya, N., Shitomi, K., Tanaka, M., Maeda, K., Matsuda, Y., & Toyoda, H. (2003). Distinguishable staining with neutral red for GFP-marked and GFP-nonmarked Fusarium oxysporum isolates simultaneously colonizing root surfaces. Journal of General Plant Pathology, 69(1), 45–48.
O'Donnell, K., Kistler, H. C., Cigelnik, E., & Ploetzm, R. C. (1998). Multiple evolutionary origins of the fungus causing Panama disease of banana: Concordant evidence from nuclear and mitochondrial gene genealogies. Proceedings of the National Academy of Sciences of the United States of America, 95(5), 2044–2049.
Orlikowski, L. B., & Ptaszek, M. (2013). First notice of Phytophthora crown and root rot of Euphorbia pulcherrima in polish greenhouses. Journal of Plant Protection Research, 53(4), 307–311.
Ronquist, F., Teslenko, M., van der Mark, P., Ayres, D. L., Darling, A., Hohna, S., Larget, B., Liu, L., Suchard, M. A., & Huelsenbeck, J. P. (2012). MrBayes 3.2: Efficient Bayesian phylogenetic inference and model choice across a large model space. Systematic Biology, 61(3), 539–542.
Darnetty, & Salleh, B. (2017). Morphological characteristics and mating populations of Fusarium species in Gibberella fujikuroi species complex (Gfsc) associated with stalk rot disease of maize in Indonesia, Malaysia and Thailand. Plant Pathology Journal, (Faisalabad), 16(1), 33–40.
Sarrocco, S., Falaschi, N., Vergara, M., Nicoletti, F., & Vannacci, G. (2007). Use of Fusarium oxysporum f. sp. dianthi transformed with marker genes to follow colonization of carnation roots. Journal of Plant Pathology, 89(1), 47–54.
Silva, F. P. d., Vechiato, M. H., & Harakava, R. (2014). EF-1α gene and IGS rDNA sequencing of Fusarium oxysporum f. sp. vasinfectum and F. oxysporum f. sp. phaseoli reveals polyphyletic origin of isolates. Tropical Plant Pathology, 39(1), 64–73.
Singh, P. K., & Kumar, V. (2011). Variability among isolates of Fusarium oxysporum f. sp. chrysanthemi pathogenic to Chrysanthemum. International Journal of Plant Pathology, 2(3), 136–143.
Tian, Y., Zhao, Y., Sun, T., Wang, L., Liu, J., Ma, X., & Hu, B. (2018). Identification and characterization of Phomopsis amygdali and Botryosphaeria dothidea associated with peach shoot blight in Yangshan, China. Plant Disease, 102(12), 2511–2518.
Unnithan, R., & Thammaiah, N. (2017). Isolation, identification and proving the pathogenicity of banana anthracnose pathogen Colletotrichum musae. International Journal of Plant Protection, 10(2), 399–403.
Visser, M., Gordon, T. R., Wingfield, B. D., Wingfield, M. J., & Viljoen, A. (2004). Transformation of Fusarium oxysporum f. sp. cubense, causal agent of Fusarium wilt of banana, with the green fluorescent protein (GFP) gene. Australasian Plant Pathology, 33(1), 69–75.
Wang, C. C., Chen, H. B., Li, W. J., & Zhou, Q. (2009). Outdoor cultivation succulents screening and utilization in Xiamen. Subtropical Plant Science, 38(4), 69–73 (in Chinese).
Wang, S. W., Wang, Q. H., Li, X. P., Zhang, Q. F., Ma, Z. K., & Tang, H. P. (2018). Progress in molecular identification in the genus Fusarium and its important applications. Microbiology, 45(4), 907–919 (in Chinese).
White, T. J., Bruns, T., Lee, S., Taylor, J.(1990). Amplification and direct sequencing of fungal ribosomal RNA genes for phylogenetics. In:Innis MA, Gelfand DH, Sninsky JJ, White TJ, eds. Pcr protocols: A guide to methods and applications. San Diego, CA:Academic Press, 315-322.
Yan, C. X. (2018). Analysis on the production and marketing situation of potted plants in China in 2017. China Flowers & Horticulture, 9, 30–33 (in Chinese).
Yang, S., Hu, Y., Cheng, Z., Rice, J. H., Miao, L., Ma, J., Hewezi, T., Li, Y., & Gai, J. (2019). An efficient Agrobacterium-mediated soybean transformation method using green fluorescent protein as a selectable marker. Plant Signaling Behavior, 14(7). https://doi.org/10.1080/15592324.2019.1612682.
Yao, J. A., Huang, P., Lan, C. Z., & Yu, D. Y. (2018). Stem rot on Cymbidium ensifolium (Orchidaceae) caused by Fusarium oxysporum in China. Canadian Journal of Plant Pathology, 40(1), 105–108.
Yao, J. A., Zhang, H., Huang, P., Chen, F., & Yu, D. Y. (2019). Green fluorescent protein genetic marker of Fusarium oxysporum F-02 of stem rot disease on Cymbidium ensifolium. Fujian Journal of Agricultural Sciences, 34(1), 70–75 (in Chinese).
Yao, J. A., Huang, P., Chen, H. X., Hou, X. Y., & Yu, D. Y. (2020). Anthracnose pathogen of the succulent plant Echeveria ‘Perle von Nürnberg’. Australasian Plant Pathology, 49, 209–212.
Zhang, Y. P., Qu, S. P., Yang, X. M., Xu, F., Zhang, L. F., Wang, J. H., & He, Y. Q. (2018). Compare to ultrastructure of root of resistant and susceptible oriental lily clones by Fusarium oxysporum f. sp. lilii. Physiological and Molecular Plant Pathology, 104, 103–107.
Zhang, D., Gao, F., Jakovlic, I., Zou, H., Zhang, J., Li, W. X., & Wang, G. T. (2020). PhyloSuite: An integrated and scalable desktop platform for streamlined molecular sequence data management and evolutionary phylogenetics studies. Molecular Ecology Resources, 20(1), 348–355.
Zhang, X. (2007). Studies on genetic diversity and transformation of the green fluorescent protein gene in Fusarium oxysporum f. sp. cubense, causal agent of banana Fusarium wilt. Doctoral dissertation. South China University of Tropical Agriculture. (in Chinese)
Acknowledgments
The authors would like to thank the Program of Fujian Provincial Science and Technology Department (grant number 2019R1024-5 and 2020 J011357) and Projects of Fujian Academy of Agricultural Sciences (grant number STIT2017-2-2, DEC201907), which funded the research.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no conflicts of interest.
Human and animal rights
This article does not contain any studies with human or animal subjects.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Yao, J., Huang, P., Chen, H. et al. Fusarium oxysporum is the pathogen responsible for stem rot of the succulent plant Echeveria ‘Perle von Nürnberg’ and observation of the infection process. Eur J Plant Pathol 159, 555–568 (2021). https://doi.org/10.1007/s10658-020-02186-4
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10658-020-02186-4