Abstract
Seaweeds have received considerable attention as sources of dietary fiber and biomass for manufacturing valuable products. The major polysaccharides of red seaweeds include agar and porphyran. In a marine environment, marine bacteria utilize agar and porphyran through the agarase and porphyranase genes encoded in their genomes. Most of these enzymes identified and characterized so far originate from marine bacteria. Recently, Bacteroides plebeius, a human gut bacterium isolated from seaweed-eating Japanese individuals, was revealed to contain a polysaccharide utilization locus (PUL) targeting the porphyran and agarose of red seaweeds. For example, B. plebeius contains an endo-type β-agarase, BpGH16A, belonging to glycoside hydrolase family 16. BpGH16A cleaves the β-1,4-glycosidic linkages of agarose and produces neoagarooligosccharides from agarose. Since it is crucial to study the characteristics of BpGH16A to understand the depolymerization pathway of red seaweed polysaccharides by B. plebeius in the human gut and to industrially apply the enzyme for the depolymerization of agar, we characterized BpGH16A for the first time. According to our results, BpGH16A is an extracellular endo-type β-agarase with an optimal temperature of 40 °C and an optimal pH of 7.0, which correspond to the temperature and pH of the human colon. BpGH16A depolymerizes agarose into neoagarotetraose (as the main product) and neoagarobiose (as the minor product). Thus, BpGH16A is suggested to be an important enzyme that initiates the depolymerization of red seaweed agarose or agar in the human gut by B. plebeius.
Key points
• Bacteroides plebeius is a human gut bacterium isolated from seaweed-eating humans.
• BpGH16A is an extracellular endo-type β-agarase with optimal conditions of 40 °C and pH 7.0.
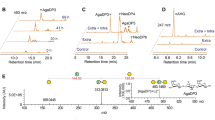
• BpGH16A depolymerizes agarose into neoagarotetraose and neoagarobiose.






Similar content being viewed by others
Data availability
All data generated or analyzed during this study are included in this published article (and its supplementary information files).
References
Allouch J, Jam M, Helbert W, Barbeyron T, Kloareg B, Henrissat B, Czjzek M (2003) The three-dimensional structures of two β-agarases. J Biol Chem 278(47):47171–47180. https://doi.org/10.1074/jbc.m308313200
Araki C (1956) Structure of the agarose constituent of agar-agar. Bull Chem Soc Jpn 29:543–544. https://doi.org/10.1246/bcsj.29.543
Chi W-J, Chang Y-K, Hong S-K (2012) Agar degradation by microorganisms and agar-degrading enzymes. Appl Microbiol Biotechnol 94(4):917–930. https://doi.org/10.1007/s00253-012-4023-2
Cui FY, Dong SJ, Shi XC, Zhao X, Zhang X-H (2014) Overexpression and characterization of a novel thermostable beta-agarase YM01-3, from marine bacterium Catenovulum agarivorans YM01T. Mar Drugs 12(5):2731–2747. https://doi.org/10.3390/md12052731
Fu XT, Kim SM (2010) Agarase: review of major sources, categories, purification method, enzyme characteristics and applications. Mar Drugs 8(1):200–218. https://doi.org/10.3390/md8010200
Fu XT, Pan C-H, Lin H, Kim SM (2009) Gene cloning, expression, and characterization of a β-agarase, AgaB34, from Agarivorans albus YKW-34. J Microbiol Biotechnol 19(3):257–264. https://doi.org/10.4014/jmb.0801.026
Guarner F, Malagelada JR (2003) Gut flora in health and disease. Lancet 361(9356):512–519. https://doi.org/10.1016/s0140-6736(03)12489-0
Hehemann J-H, Correc G, Barbeyron T, Helbert W, Czjzek M, Michel G (2010) Transfer of carbohydrate-active enzymes from marine bacteria to Japanese gut microbiota. Nature 464(7290):908–U123. https://doi.org/10.1038/nature08937
Hehemann J-H, Kelly AG, Pudlo NA, Martens EC, Boraston AB (2012) Bacteria of the human gut microbiome catabolize red seaweed glycans with carbohydrate-active enzyme updates from extrinsic microbes. Proc Natl Acad Sci U S A 109(48):19786–19791. https://doi.org/10.1073/pnas.1211002109
Kim HT, Lee S, Lee D, Kim H-S, Bang W-G, Kim KH, Choi I-G (2010) Overexpression and molecular characterization of Aga50D from Saccharophagus degradans 2-40: an exo-type β-agarase producing neoagarobiose. Appl Microbiol Biotechnol 86(1):227–234. https://doi.org/10.1007/s00253-009-2256-5
Kim JH, Yun EJ, Seo N, Yu S, Kim DH, Cho KM, An HJ, Kim J-H, Choi I-G, Kim KH (2017) Enzymatic liquefaction of agarose above the sol-gel transition temperature using a thermostable endo-type β-agarase, Aga16B. Appl Microbiol Biotechnol 101(3):1111–1120. https://doi.org/10.1007/s00253-016-7831-y
Knutsen S, Myslabodski D, Larsen B, Usov AI (1994) A modified system of nomenclature for red algal galactans. Bot Mar 37:163–170. https://doi.org/10.1515/botm.1994.37.2.163
Lahaye M, Yaphe W, Viet MTP, Rochas C (1989) 13C-n.m.r. spectroscopic investigation of methylated and charged agarose oligosaccharides and polysaccharides. Carbohydr Res 190(2):249–265. https://doi.org/10.1016/0008-6215(89)84129-1
Li MM, Li GS, Zhu LY, Yin YS, Zhao XL, Xiang C, Yu GL, Wang X (2014) Isolation and characterization of an agaro-oligosaccharide (AO)-hydrolyzing bacterium from the gut microflora of Chinese individuals. PLoS One 9(3):9. https://doi.org/10.1371/journal.pone.0091106
Li MM, Shang QS, Li GS, Wang X, Yu GL (2017) Degradation of marine algae-derived carbohydrates by Bacteroidetes isolated from human gut microbiota. Mar Drugs 15(4). https://doi.org/10.3390/md15040092
Miller GL (1959) Use of dinitrosalicylic acid reagent for determination of reducing sugar. Anal Chem 31(3):426–428. https://doi.org/10.1021/ac60147a030
Ohta Y, Hatada Y, Nogi Y, Miyazaki M, Li Z, Akita M, Hidaka Y, Goda S, Ito S, Horikoshi K (2004) Enzymatic properties and nucleotide and amino acid sequences of a thermostable β-agarase from a novel species of deep-sea Microbulbifer. Appl Microbiol Biotechnol 64(4):505–514. https://doi.org/10.1007/s00253-004-1573-y
Rios-Covian D, Ruas-Madiedo P, Margolles A, Gueimonde M, de los Reyes-Gavilan CG, Salazar N (2016) Intestinal short chain fatty acids and their link with diet and human health. Front Microbiol 7:9. https://doi.org/10.3389/fmicb.2016.00185
Sievers F, Wilm A, Dineen D, Gibson TJ, Karplus K, Li WZ, Lopez R, McWilliam H, Remmert M, Soding J, Thompson JD, Higgins DG (2011) Fast, scalable generation of high-quality protein multiple sequence alignments using Clustal Omega. Mol Syst Biol 7:539. https://doi.org/10.1038/msb.2011.75
Temuujin U, Chi W-J, Lee S-Y, Chang Y-K, Hong S-K (2011) Overexpression and biochemical characterization of DagA from Streptomyces coelicolor A3(2): an endo-type β-agarase producing neoagarotetraose and neoagarohexaose. Appl Microbiol Biotechnol 92(4):749–759. https://doi.org/10.1007/s00253-011-3347-7
Wei N, Quarterman J, Jin Y-S (2013) Marine macroalgae: an untapped resource for producing fuels and chemicals. Trends Biotechnol 31(2):70–77. https://doi.org/10.1016/j.tibtech.2012.10.009
Yun EJ, Lee S, Kim JH, Kim BB, Kim HT, Lee SH, Pelton JG, Kang NJ, Choi I-G, Kim KH (2013) Enzymatic production of 3,6-anhydro-l-galactose from agarose and its purification and in vitro skin whitening and anti-inflammatory activities. Appl Microbiol Biotechnol 97(7):2961–2970. https://doi.org/10.1007/s00253-012-4184-z
Yun EJ, Choi I-G, Kim KH (2015) Red macroalgae as a sustainable resource for bio-based products. Trends Biotechnol 33(5):247–249. https://doi.org/10.1016/j.tibtech.2015.02.006
Yun EJ, Yu S, Kim KH (2017) Current knowledge on agarolytic enzymes and the industrial potential of agar-derived sugars. Appl Microbiol Biotechnol 101(14):5581–5589. https://doi.org/10.1007/s00253-017-8383-5
Acknowledgments
The experiments were performed at the facilities of the Institute of Biomedical Science and Food Safety at the Food Safety Hall, Korea University.
Funding
This research was supported by the Mid-career Researcher Program funded by the National Research Foundation of Korea (NRF) (2020R1A2B5B02002631) as well as the Ministry of Oceans and Fisheries, Korea (20200367). EJY acknowledges a grant support from the NRF (2020R1C1C1008048). SY was financially supported by the NRF (2020R1A6A3A01100292) and Korea University.
Author information
Authors and Affiliations
Contributions
K.H.K. and E.J.Y. conceived and designed the research. N.J.P., S.Y., and D.H.K. performed the experiments. N.J.P., S.Y., and E.J.Y. analyzed the data. K.H.K., E.J.Y., N.J.P., and S.Y. wrote the manuscript. All authors read and approved the manuscript.
Corresponding authors
Ethics declarations
Conflict of interest
K.H.K., E.J.Y., N.J.P., S.Y., and D.H.K. have filed a patent on a part of this work.
Ethical approval
This article does not contain any studies with human participants or animals performed by any of the authors.
Additional information
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
ESM 1
(PDF 281 kb).
Rights and permissions
About this article
Cite this article
Park, N.J., Yu, S., Kim, D.H. et al. Characterization of BpGH16A of Bacteroides plebeius, a key enzyme initiating the depolymerization of agarose in the human gut. Appl Microbiol Biotechnol 105, 617–625 (2021). https://doi.org/10.1007/s00253-020-11039-3
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00253-020-11039-3