Abstract
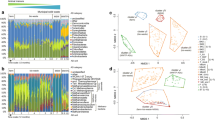
The diversity and assembly of activated sludge microbiomes play a key role in the performances of municipal wastewater treatment plants (WWTPs), which are the most widely applied biotechnological process systems. In this study, we investigated the microbiomes of municipal WWTPs in Bangkok, Wuhan, and Beijing that respectively represent tropical, subtropical, and temperate climate regions, and also explored how microbiomes assembled in these municipal WWTPs. Our results showed that the microbiomes from these municipal WWTPs were significantly different. The assembly of microbiomes in municipal WWTPs followed deterministic and stochastic processes governed by geographical location, temperature, and nutrients. We found that both taxonomic and phylogenetic α-diversities of tropical Bangkok municipal WWTPs were the highest and were rich in yet-to-be-identified microbial taxa. Nitrospirae and β-Proteobacteria were more abundant in tropical municipal WWTPs, but did not result in better removal efficiencies of ammonium and total nitrogen. Overall, these results suggest that tropical and temperate municipal WWTPs harbored diverse and unique microbial resources, and the municipal WWTP microbiomes were assembled with different processes. Implications of these findings for designing and running tropical municipal WWTPs were discussed.
Key points
• Six WWTPs of tropical Thailand and subtropical and temperate China were investigated.
• Tropical Bangkok WWTPs had more diverse and yet-to-be-identified microbial taxa.
• Microbiome assembly processes were associated with geographical location.






Similar content being viewed by others

References
Alawi M, Off S, Kaya M, Spieck E (2009) Temperature influences the population structure of nitrite-oxidizing bacteria in activated sludge. Environ Microbiol Rep 1(3):184–190. https://doi.org/10.1111/j.1758-2229.2009.00029.x
Altschul SF, Gish W, Miller W, Myers EW, Lipman DJ (1990) Basic local alignment search tool. J Mol Biol 215(3):403–410. https://doi.org/10.1016/S0022-2836(05)80360-2
Bolyen E, Rideout JR, Dillon MR, Bokulich NA, Abnet CC, Al-Ghalith GA, Alexander H, Alm EJ, Arumugam M, Asnicar F (2019) Reproducible, interactive, scalable and extensible microbiome data science using QIIME 2. Nat Biotechnol 37(8):852–857. https://doi.org/10.1038/s41587-019-0209-9
Brown JH, Gillooly JF, Allen AP, Savage VM, West GB (2004) Toward a metabolic theory of ecology. Ecology 85(7):1771–1789. https://doi.org/10.1890/03-9000
Burger W, Krysiak-Baltyn K, Scales PJ, Martin GJ, Stickland AD, Gras SL (2017) The influence of protruding filamentous bacteria on floc stability and solid-liquid separation in the activated sludge process. Water Res 123:578–585. https://doi.org/10.1016/j.watres.2017.06.063
Cai L, Ju F, Zhang T (2013) Tracking human sewage microbiome in a municipal wastewater treatment plant. Appl Microbiol Biotechnol 98:3317–3326. https://doi.org/10.1007/s00253-013-5402-z
Cai X, Mao Y, Xu J, Tian L, Wang Y, Iqbal W, Yang B, Liu C, Zhao X, Wang Y (2020) Characterizing community dynamics and exploring bacterial assemblages in two activated sludge systems. Appl Microbiol Biotechnol 104:1795–1808. https://doi.org/10.1007/s00253-019-10279-2
Chen Y, Lan S, Wang L, Dong S, Zhou H, Tan Z, Li X (2017) A review: driving factors and regulation strategies of microbial community structure and dynamics in wastewater treatment systems. Chemosphere 174:173–182. https://doi.org/10.1016/j.chemosphere.2017.01.129
Chen H, Wang M, Chang S (2020) Disentangling community structure of ecological system in activated sludge: core communities, functionality, and functional redundancy. Microb Ecol 2020:1–13. https://doi.org/10.1007/s00248-020-01492-y
Daims H, Taylor MW, Wagner M (2006) Wastewater treatment: a model system for microbial ecology. Trends Biotechnol 24:483–489. https://doi.org/10.1016/j.tibtech.2006.09.002
Edgar RC (2010) Search and clustering orders of magnitude faster than BLAST. Bioinformatics 26:2460–2461. https://doi.org/10.1093/bioinformatics/btq461
Edgar RC (2013) UPARSE: highly accurate OTU sequences from microbial amplicon reads. Nat Methods 10:996–998. https://doi.org/10.1038/nmeth.2604
Edgar RC, Haas BJ, Clemente JC, Quince C, Knight R (2011) UCHIME improves sensitivity and speed of chimera detection. Bioinformatics 27:2194–2200. https://doi.org/10.1093/bioinformatics/btr381
Fierer N, Jackson RB (2006) The diversity and biogeography of soil bacterial communities. Proc Natl Acad Sci U S A 103:626–631. https://doi.org/10.1073/pnas.0507535103
Fuhrman JA, Steele JA, Hewson I, Schwalbach MS, Brown MV, Green JL, Brown JH (2008) A latitudinal diversity gradient in planktonic marine bacteria. Proc Natl Acad Sci U S A 105:7774–7778. https://doi.org/10.1073/pnas.0803070105
Gao P, Xu W, Sontag P, Li X, Xue G, Liu T, Sun W (2016) Correlating microbial community compositions with environmental factors in activated sludge from four full-scale municipal wastewater treatment plants in Shanghai, China. Appl Microbiol Biotechnol 100:4663–4673. https://doi.org/10.1007/s00253-016-7307-0
Griffin JS, Wells GF (2017) Regional synchrony in full-scale activated sludge bioreactors due to deterministic microbial community assembly. ISME J 11:500–511. https://doi.org/10.1038/ismej.2016.121
Guo F, Zhang T (2012) Profiling bulking and foaming bacteria in activated sludge by high throughput sequencing. Water Res 46:2772–2782. https://doi.org/10.1016/j.watres.2012.02.039
Hollander M, Wolfe DA, Chicken E (2015) Nonparametric Statistical Methods, Third Edition. John Wiley & Sons, Inc., Hoboken. https://doi.org/10.1002/9781119196037
Hu M, Wang X, Wen X, Xia Y (2012) Microbial community structures in different wastewater treatment plants as revealed by 454-pyrosequencing analysis. Bioresour Technol 117:72–79. https://doi.org/10.1016/j.biortech.2012.04.061
Huang Z, Gedalanga PB, Asvapathanagul P, Olson BH (2010) Influence of physicochemical and operational parameters on Nitrobacter and Nitrospira communities in an aerobic activated sludge bioreactor. Water Res 44:4351–4358. https://doi.org/10.1016/j.watres.2010.05.037
Ibarbalz FM, Orellana E, Figuerola EL, Erijman L (2016) Shotgun metagenomic profiles have a high capacity to discriminate samples of activated sludge according to wastewater type. Appl Environ Microbiol 82:5186–5196. https://doi.org/10.1128/AEM.00916-16
Jenkins D, Wanner J (2014) Activated sludge – 100 years and counting. Volume 13. IWA Publishing. https://doi.org/10.2166/9781780404943
Jiang XT, Ye L, Ju F, Wang YL, Zhang T (2018) Toward an intensive longitudinal understanding of activated sludge bacterial assembly and dynamics. Environ Sci Technol 52:8224–8232. https://doi.org/10.1021/acs.est.7b05579
Johnston J, Behrens S (2020) Seasonal dynamics of the activated sludge microbiome in sequencing batch reactors using 16S rRNA transcript amplicon sequencing. Appl Environ Microbiol 86. https://doi.org/10.1128/AEM.00597-20
Ju F, Zhang T (2015) Bacterial assembly and temporal dynamics in activated sludge of a full-scale municipal wastewater treatment plant. ISME J 9:683–695. https://doi.org/10.1038/ismej.2014.162
Ju F, Guo F, Ye L, Xia Y, Zhang T (2014) Metagenomic analysis on seasonal microbial variations of activated sludge from a full-scale wastewater treatment plant over 4 years. Environ Microbiol Rep 6:80–89. https://doi.org/10.1111/1758-2229.12110
Kottek M, Grieser J, Beck C, Rudolf B, Rubel F (2006) World map of the Köppen-Geiger climate classification updated. Meteorol Z 15:259–263. https://doi.org/10.1127/0941-2948/2006/0130
Kozich JJ, Westcott SL, Baxter NT, Highlander SK, Schloss PD (2013) Development of a dual-index sequencing strategy and curation pipeline for analyzing amplicon sequence data on the MiSeq Illumina sequencing platform. Appl Environ Microbiol 79:5112–5120. https://doi.org/10.1128/AEM.01043-13
Krause S, Le Roux X, Niklaus PA, Van Bodegom PM, Lennon JT, Bertilsson S, Grossart HP, Philippot L, Bodelier PL (2014) Trait-based approaches for understanding microbial biodiversity and ecosystem functioning. Front Microbiol 5:251. https://doi.org/10.3389/fmicb.2014.00251
Liu F, Hu X, Zhao X, Guo H, Zhao Y (2019) Microbial community structures’ response to seasonal variation in a full-scale municipal wastewater treatment plant. Environ Eng Sci 36:172–179. https://doi.org/10.1089/ees.2018.0280
Lopez-Vazquez CM, Hooijmans CM, Brdjanovic D, Gijzen HJ, van Loosdrecht MC (2008) Factors affecting the microbial populations at full-scale enhanced biological phosphorus removal (EBPR) wastewater treatment plants in The Netherlands. Water Res 42:2349–2360. https://doi.org/10.1016/j.watres.2008.01.001
Magoc T, Salzberg SL (2011) FLASH: fast length adjustment of short reads to improve genome assemblies. Bioinformatics 27:2957–2963. https://doi.org/10.1093/bioinformatics/btr507
Martin M (2011) Cutadapt removes adapter sequences from high-throughput sequencing reads. EMBnet J 17:10–12. https://doi.org/10.14806/ej.17.1.200
Martiny JB, Eisen JA, Penn K, Allison SD, Horner-Devine MC (2011) Drivers of bacterial β-diversity depend on spatial scale. Proc Natl Acad Sci U S A 108:7850–7854. https://doi.org/10.1073/pnas.1016308108
McGill BJ, Etienne RS, Gray JS, Alonso D, Anderson MJ, Benecha HK, Dornelas M, Enquist BJ, Green JL, He F (2007) Species abundance distributions: moving beyond single prediction theories to integration within an ecological framework. Ecol Lett 10:995–1015. https://doi.org/10.1111/j.1461-0248.2007.01094.x
McIlroy SJ, Kirkegaard RH, McIlroy B, Nierychlo M, Kristensen JM, Karst SM, Albertsen M, Nielsen PH (2017) MiDAS 2.0: an ecosystem-specific taxonomy and online database for the organisms of wastewater treatment systems expanded for anaerobic digester groups. Database 2017. https://doi.org/10.1093/database/bax016
Newton RJ, McLellan SL, Dila DK, Vineis JH, Morrison HG, Eren AM, Sogin ML (2015) Sewage reflects the microbiomes of human populations. mBio 6:e02574–e02514. https://doi.org/10.1128/mBio.02574-14
Nielsen PH, Saunders AM, Hansen AA, Larsen P, Nielsen JL (2012) Microbial communities involved in enhanced biological phosphorus removal from wastewater-a model system in environmental biotechnology. Curr Opin Biotechnol 23:452–459. https://doi.org/10.1016/j.copbio.2011.11.027
Nierychlo M, Andersen KS, Xu Y, Green N, Albertsen M, Dueholm MS, Nielsen PH (2019) Species-level microbiome composition of activated sludge-introducing the MiDAS 3 ecosystem-specific reference database and taxonomy. bioRxiv:842393. https://doi.org/10.1101/842393
Ofiţeru ID, Lunn M, Curtis TP, Wells GF, Criddle CS, Francis CA, Sloan WT (2010) Combined niche and neutral effects in a microbial wastewater treatment community. Proc Natl Acad Sci U S A 107:15345–15350. https://doi.org/10.1073/pnas.1000604107
Parks DH, Tyson GW, Hugenholtz P, Beiko RG (2014) STAMP: statistical analysis of taxonomic and functional profiles. Bioinformatics 30:3123–3124. https://doi.org/10.1093/bioinformatics/btu494
Pedregosa F, Varoquaux G, Gramfort A, Michel V, Thirion B, Grisel O, Blondel M, Prettenhofer P, Weiss R, Dubourg V (2011) Scikit-learn: machine learning in Python. J Mach Learn Res 12:2825–2830
Pinel-Alloul B, André A, Legendre P, Cardille JA, Patalas K, Salki A (2013) Large-scale geographic patterns of diversity and community structure of pelagic crustacean zooplankton in Canadian lakes. Glob Ecol Biogeogr 22:784–795. https://doi.org/10.1111/geb.12041
Quast C, Pruesse E, Yilmaz P, Gerken J, Schweer T, Yarza P, Peplies J, Glockner FO (2013) The SILVA ribosomal RNA gene database project: improved data processing and web-based tools. Nucleic Acids Res 41:590–596. https://doi.org/10.1093/nar/gks1219
Saunders AM, Albertsen M, Vollertsen J, Nielsen PH (2016) The activated sludge ecosystem contains a core community of abundant organisms. ISME J 10:11–20. https://doi.org/10.1038/ismej.2015.117
Segata N, Izard J, Waldron L, Gevers D, Miropolsky L, Garrett WS, Huttenhower C (2011) Metagenomic biomarker discovery and explanation. Genome Biol 12:R60. https://doi.org/10.1186/gb-2011-12-6-r60
Segura A, Calliari D, Kruk C, Fort H, Izaguirre I, Saad JF, Arim M (2015) Metabolic dependence of phytoplankton species richness. Glob Ecol Biogeogr 24:472–482. https://doi.org/10.1111/geb.12258
Soddell JA, Seviour RJ (1995) Relationship between temperature and growth of organisms causing Nocardia foams in activated sludge plants. Water Res 29:1555–1558. https://doi.org/10.1016/0043-1354(94)00222-S
Song Y, Jiang CY, Liang ZL, Wang BJ, Jiang Y, Yin Y, Zhu HZ, Qin YL, Cheng RX, Liu ZP, Liu Y, Jin T, Corvini PF, Rabaey K, Wang AJ, Liu SJ (2020) Casimicrobium huifangae gen. nov., sp. nov., a ubiquitous “most-wanted” core bacterial taxon from municipal wastewater treatment plants. Appl Environ Microbiol 86(4). https://doi.org/10.1128/AEM.02209-19
Speirs LBM, Rice DTF, Petrovski S, Seviour RJ (2019) The phylogeny, biodiversity, and ecology of the Chloroflexi in activated sludge. Front Microbiol 10:2015. https://doi.org/10.3389/fmicb.2019.02015
Stegen JC, Lin X, Fredrickson JK, Chen X, Kennedy DW, Murray CJ, Rockhold ML, Konopka A (2013) Quantifying community assembly processes and identifying features that impose them. ISME J 7:2069–2079. https://doi.org/10.1038/ismej.2013.93
Stegen JC, Lin X, Fredrickson JK, Konopka AE (2015) Estimating and mapping ecological processes influencing microbial community assembly. Front Microbiol 6:370. https://doi.org/10.3389/fmicb.2015.00370
Steudel B, Hector A, Friedl T, Löfke C, Lorenz M, Wesche M, Kessler M (2012) Biodiversity effects on ecosystem functioning change along environmental stress gradients. Ecol Lett 15:1397–1405. https://doi.org/10.1111/j.1461-0248.2012.01863.x
Tilman D (1982) Resource Competition and Community Structure. (MPB-17), Volume 17. Princeton University Press. https://doi.org/10.1515/9780691209654
Tribelli PM, Lopez NI (2018) Reporting key features in cold-adapted bacteria. Life (Basel) 8:8. https://doi.org/10.3390/life8010008
Valentín-Vargas A, Toro-Labrador G, Massol-Deya AA (2012) Bacterial community dynamics in full-scale activated sludge bioreactors: operational and ecological factors driving community assembly and performance. PLoS One 7:e42524. https://doi.org/10.1371/journal.pone.0042524
Vuono DC, Benecke J, Henkel J, Navidi WC, Cath TY, Munakatamarr J, Spear JR, Drewes JE (2015) Disturbance and temporal partitioning of the activated sludge metacommunity. ISME J 9:425–435. https://doi.org/10.1038/ismej.2014.139
Vuono DC, Munakata-Marr J, Spear JR, Drewes JE (2016) Disturbance opens recruitment sites for bacterial colonization in activated sludge. Environ Microbiol 18:87–99. https://doi.org/10.1111/1462-2920.12824
Wang P, Yu Z, Qi R, Zhang H (2016) Detailed comparison of bacterial communities during seasonal sludge bulking in a municipal wastewater treatment plant. Water Res 105:157–166. https://doi.org/10.1016/j.watres.2016.08.050
Wang YH, Huang Z, Liu SJ (2019) Chemotaxis towards aromatic compounds: insights from Comamonas testosteroni. Int J Mol Sci 20:2701. https://doi.org/10.3390/ijms20112701
Wei Z, Liu Y, Feng K, Li S, Wang S, Jin D, Zhang Y, Chen H, Yin H, Xu M, Deng Y (2018) The divergence between fungal and bacterial communities in seasonal and spatial variations of wastewater treatment plants. Sci Total Environ 628-629:969–978. https://doi.org/10.1016/j.scitotenv.2018.02.003
Wu L, Ning D, Zhang B, Li Y, Zhang P, Shan X, Zhang Q, Brown M, Li Z, Van Nostrand JD, Ling F, Xiao N, Zhang Y, Vierheilig J, Wells GF, Yang Y, Deng Y, Tu Q, Wang A, Zhang T, He Z, Keller J, Nielsen PH, Alvarez PJJ, Criddle CS, Wagner M, Tiedje JM, He Q, Curtis TP, Stahl DA, Alvarez-Cohen L, Rittmann BE, Wen X, Zhou J (2019) Global diversity and biogeography of bacterial communities in wastewater treatment plants. Nat Microbiol 4:1183–1195. https://doi.org/10.1038/s41564-019-0426-5
Xia Y, Kong Y, Thomsen TR, Halkjaer Nielsen P (2008) Identification and ecophysiological characterization of epiphytic protein-hydrolyzing Saprospiraceae (“Candidatus Epiflobacter” spp.) in activated sludge. Appl Environ Microbiol 74:2229–2238. https://doi.org/10.1128/AEM.02502-07
Xia Y, Wen X, Zhang B, Yang Y (2018) Diversity and assembly patterns of activated sludge microbial communities: a review. Biotechnol Adv 36:1038–1047. https://doi.org/10.1016/j.biotechadv.2018.03.005
Zhang X, Qu Y, Ma Q, Zhang Z, Li D, Wang J, Shen W, Shen E, Zhou J (2015) Illumina MiSeq sequencing reveals diverse microbial communities of activated sludge systems stimulated by different aromatics for indigo biosynthesis from indole. PLoS One 10:e0125732. https://doi.org/10.1371/journal.pone.0125732
Zhang Z, Deng Y, Feng K, Cai W, Li S, Yin H, Xu M, Ning D, Qu Y (2019) Deterministic assembly and diversity gradient altered the biofilm community performances of bioreactors. Environ Sci Technol 53:1315–1324. https://doi.org/10.1021/acs.est.8b06044
Zhang B, Ning D, Van Nostrand JD, Sun C, Yang Y, Zhou J, Wen X (2020a) Biogeography and assembly of microbial communities in wastewater treatment plants in China. Environ Sci Technol 54:5884–5892. https://doi.org/10.1021/acs.est.9b07950
Zhang B, Ning D, Yang Y, Van Nostrand JD, Zhou J, Wen X (2020b) Biodegradability of wastewater determines microbial assembly mechanisms in full-scale wastewater treatment plants. Water Res 169:115276. https://doi.org/10.1016/j.watres.2019.115276
Zhang K, Delgado-Baquerizo M, Zhu YG, Chu H (2020c) Space is more important than season when shaping soil microbial communities at a large spatial scale. mSystems 5. https://doi.org/10.1128/mSystems.00783-19
Zhao Y, Park HD, Park JH, Zhang F, Chen C, Li X, Zhao D, Zhao F (2016) Effect of different salinity adaptation on the performance and microbial community in a sequencing batch reactor. Bioresour Technol 216:808–816. https://doi.org/10.1016/j.biortech.2016.06.032
Zhou J, Deng Y, Shen L, Wen C, Yan Q, Ning D, Qin Y, Xue K, Wu L, He Z, Voordeckers JW, Nostrand JD, Buzzard V, Michaletz ST, Enquist BJ, Weiser MD, Kaspari M, Waide R, Yang Y, Brown JH (2016) Temperature mediates continental-scale diversity of microbes in forest soils. Nat Commun 7:12083. https://doi.org/10.1038/ncomms12083
Funding
This work was supported by research grants from the Ministry of Science and Technology of China (KY201701011; 2019YFA0905500; 2019YFC1905001), International Partnership Program of Chinese Academy of Sciences (153211KYSB20160029), and National Center for Genetic Engineering and Biotechnology, National Science and Technology Development Agency of Thailand (P-18-52413).
Author information
Authors and Affiliations
Contributions
YS, WM, and SYL performed research and analyzed data. NP collected samples and analyzed physio-chemical parameters. SW conducted the molecular works. YS wrote the paper. CYJ, PK, and DRQ conceived and designed research. SJL, AJW, and VC supervised the project, revised the manuscript, and acquired funding. All authors read and approved the manuscript.
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no competing interests.
Ethical approval
This article does not contain any studies with human participants or animals performed by any of the authors.
Additional information
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
About this article
Cite this article
Song, Y., Mhuantong, W., Liu, SY. et al. Tropical and temperate wastewater treatment plants assemble different and diverse microbiomes. Appl Microbiol Biotechnol 105, 853–867 (2021). https://doi.org/10.1007/s00253-020-11082-0
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00253-020-11082-0