Abstract
Brain stroke is an emergency medical condition which occurs mainly due to insufficient blood flow to the brain. It results in permanent cellular-level damage. There are two main types of brain stroke, ischemic and hemorrhagic. Ischemic brain stroke is caused by a lack of blood flow, and the haemorrhagic form is due to internal bleeding. The affected part of brain will not function properly after this attack. Hence, early detection is important for more efficacious treatment. Computer-aided diagnosis is a type of non-invasive diagnostic tool which can help in detecting life-threatening disease in its early stage by utilizing image processing and soft computing techniques. In this paper, we have developed one such model to assess intracerebral haemorrhage by employing non-linear features combined with a probabilistic neural network classifier and computed tomography (CT) images. Our model achieved a maximum accuracy of 97.37% in discerning normal versus haemorrhagic subjects. An intracerebral haemorrhage index is also developed using only three significant features. The clinical and statistical validation of the model confirms its suitability in providing for improved treatment planning and in making strategic decisions.
Similar content being viewed by others
Introduction
Stroke is one of the life-threatening cerebrovascular diseases that occurs due to the abrupt termination of blood supply to the brain. As the brain tissues cannot receive oxygen and nutrients, they begin to die, which can result in substantial lifetime disability and even death [1, 2]. Stroke is the second leading cause of lifetime disability as well as death worldwide [3]. Intracerebral haemorrhage (ICH) is one of the devastating stroke subtypes with poor outcome and high mortality rate within the first year. Hence, early diagnosis and management of ICH are essential for improved functional outcomes and to save patient lives [4].
Computed tomography (CT) is the preferred modality of choice for assessment of haemorrhage due to its wide availability, lesser cost, rapidity, and high sensitivity for haemorrhage [5]. In CT imagery, haemorrhagic lesions can be characterised as brighter areas as compared to the surroundings. Manual segmentation of haemorrhage from CT imagery remains challenging due to its uneven boundaries, overlapping pixel intensities, noise, and artefacts in CT scans. Even though manual demarcation and estimation appears accurate, it is a time-consuming task, heavily dependent on the expertise of clinicians, and subject to intraobserver and interobserver variability [6,7,8,9]. Furthermore, the irregularities and complexities associated with varied shapes and sizes of haemorrhagic lesions with time will also make the process more difficult and strenuous. Moreover, the process can become a laborious and daunting task, particularly in large clinical settings, which can introduce inadvertent error and delay [10,11,12,13,14]. This can cause additional morbidity and even mortality to the patient. Thus, development of automated methods which will support clinicians for making rapid, reliable, and efficient diagnosis of haemorrhage from CT images would be highly beneficial.
Computer aided diagnosis (CAD) for haemorrhage detection
CAD systems can play an important role in early diagnosis of haemorrhage. CAD offers quick and consistent results with improved accuracy, thus enabling clinicians to make better treatment planning and strategic decisions [15, 16]. The triage workflow can also be streamlined and optimized by automating the routine scanning of CT images with little or no human intervention. The following subsection discusses the existing works related to detection of haemorrhage using CAD. Several semi-automated and automated methods have been developed to detect ICH from CT images. Prakash et al. [17] applied the modified distance regularized level set (MDLRSE) for the detection of ICH and intraventricular haemorrhage (IVH) in CT scans. This two-step method involves shrinking and expansion and is highly sensitive to several DRLSE parameters; it requires a long computation time. Shahangian and Pourghassem [18] presented another modified version of the DRLSE method to detect small and uncertain haemorrhages. A hierarchical classifier was further used to distinguish the segmented regions into three different types of haemorrhage. However, control parameters require proper adjustment for accurate segmentation. Gillebert et al. [19] developed a voxel-based outlier detection procedure using a set of control images to classify infarct and haemorrhage in stroke CT slices. They reported a dice similarity index (DSI) of 0.52–0.89 for 500 CT scans. Muschelli et al. [20] proposed a fully automated method based on voxel-level probabilities to segment the haemorrhage. They obtained a higher median DSI of 0.899 using the random forest algorithm on 112 3D CT scan images. Al-Ayoob et al. [21] presented an automated segmentation method using Otsu’s thresholding, morphological operations, and a region-growing technique. The method was tested on 76 CT images, and achieved a 92% accuracy for classification. Zhang et al. [22] applied adaptive thresholding and case-based reasoning to detect ICH and IVH in CT head scans. They achieved a segmentation accuracy of 95% for a set of ten CT images with different haemorrhage sizes ranging from small to large. Kumar et al. [23] combined fuzzy c-mean (FCM) clustering and DRLSE for haemorrhage segmentation using CT images. An entropy-based thresholding is applied to automatically selected FCM clustered imagery, and DRLSE is applied to the resultant images for ICH segmentation. They reported an average sensitivity of 87.06% for 35 brain CT images. Phan et al. [24] developed another method using Hounsfield values and k-nearest neighbour to classify four types of haemorrhage. The method obtained an accuracy of 93.33% on 150 images. Gautam and Raman [25] have used white matter fuzzy c-means (WMFCM) and wavelet based thresholding to detect and localise haemorrhage in 20 CT images. Nag et al. [26] has delineated haemorrhagic regions in 48 CT images by combining an autoencoder and the Chan Vese model. Foo et al. [27] developed a CAD system using multiple thresholding and symmetry detection to identify acute ICH in 108 CT volumes.
From the above studies, it is evident that a limited dataset of CT images has been utilized for ICH detection and classification. Furthermore, a few of the existing techniques involves intricate engineering techniques such as skull stripping, rigid body transformation and feature extraction based on intensity of voxels and information on local moments. It can also be noted that some of the segmentation methods are time consuming and complex due to the intricacies associated with the initialisation of mathematical models and also require manual intervention. In addition, some of these methods are based on hardcoded logic and presumptive rules, which will limit their application to certain specific scenarios. Hence, there is a need for a quick, robust, efficient, and cost-effective automated method which can accurately detect and classify ICH for a large set of CT images. The main goal of this paper is to identify the ICH very quickly in the CT images using efficient feature extraction method. The contributions of the paper are two-fold:
-
Entropy based non-linear features are efficiently extracted on a significantly large dataset to characterise the haemorrhagic brain stroke using CT images.
-
ICH risk index is formulated using only three significant features. This helps to perform the categorisation using a unique number.
The remainder of the paper is prepared as follows: “Materials” designates the used materials and the acquisition method. The framework of the proposed assessment tool is described in “Framework of our approach”. The experimental results and its analysis is discussed in “Experimental results and discussions”. Finally, we conclude the paper in “Conclusion”.
Materials
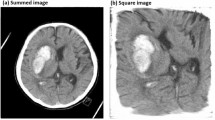
The CT images were collected from ICH patients who are admitted for the CT scan in the Department of Radio-diagnosis of Kalinga Institute of Medical Science (KIMS), Bhubaneswar, India. The images were retrospectively acquired on a Digital p64-Slice CT scan machine in consultation with the radiologist. Based on visual inspection by the experts, the presence or absence of lesions was considered for the image collection. A total of 1603 non-contrast head CT axial images were collected from 48 patients, of which 784 were normal and 819 were ICH. The CT dataset was properly anonymized by an individual who was not part of the study. The CT slice which includes the largest area of haemorrhage was selected and saved in JPEG format. Figure 1 displays utilized samples of CT images in the current study.
Framework of our approach
We propose a new automated technique that can classify normal and haemorrhagic CT imagery using a minimal set of highly discriminating non-linear features along with normal classifiers. The proposed brain pathology identification system utilizes the following stages: (1) Feature generation, (2) feature organization, and (3) classification. In the first stage, brain imagery is pre-processed and nonlinear features are extracted with seven different types of entropy measures. Thereafter, features are rearranged using Student's t test. Finally, the significant features are categorized using the probabilistic neural network classifier. The details of the proposed technique are explained in the following sections. The overview of the proposed technique is illustrated in Fig. 2.
Feature generation
This stage generates the features to characterise brain CT imagery. It involves pre-processing and nonlinear feature extraction stages, which are briefly explained below.
Pre-processing
The main aim of the initial pre-processing phase is to improve image quality for further processing. This phase involves removal of noise and unwanted regions from the input images, which will contribute to an overall improvement in the performance of the automated system. The following steps are applied to each input image as part of pre-processing. (a) A mask is initially created to remove the skull from the CT image [28], (b) contrast limited adaptive histogram equalisation (CLAHE) is applied to enhance the contrast and intensity of the image [29], and (c) the mask is then applied to the thresholded image to extract the brain region [28]. Figure 3 depicts the output of the pre-processing stage.
Non-linear feature extraction
Morphological alteration in brain tissue can be captured by automated extraction of key non-linear texture-based features. In this work, textural features are extracted using statistical moments of the image gray level histogram. Entropy is one such robust textural descriptor which can be used to accurately classify normal versus abnormal images. Entropy indicates the degree of randomness in the intensity distribution of a grayscale image. Consider an image f(x, y) with a gray level histogram hi where i = 0,1,…L − 1 denotes the various gray levels present. Then the normalised histogram Hi of image f with dimensions M and N in the X and Y directions is given as:
Various entropy measures for textural feature extraction are as follows:
Shannon entropy (Shan.entropy) [30]: It is defined as:
Rényi entropy (Ren. entropy) [30, 31]: It is defined as:
where \(\propto \) is the diversity index, and it is considered that \(\propto =3\).
Kapur entropy (Kap. entropy) [30, 31]: It is defined as:
where \(\alpha and \beta \) are diversity indices with \(\alpha \ne 1\), \(\alpha + \beta -1 >0\), and \(\alpha \) = 0.5 and \(\beta \) = 0.7.
Yager entropy (Yag. entropy) [31]: It is defined as:
Vajda entropy (Vaj. entropy) [32]: It is defined as:
Maximum entropy (Max. entropy) [33]:
Assume (i, j) where i is the gray level of a pixel and j is the average gray level of its neighborhood in the image f(x, y) and p(i, j) is the gray level probability. Supposing that the threshold of maximum entropy is set at [s, t] gray level pair, then the target segmented areas will consist of pairs (i1, j1) where i1 = 0,1,..,s and j1 = 0,1,..,t and background areas will have pairs (i2, j2) where i2 = s + 1,..,L − 1 and j2 = t + 1,..,L − 1. Then the total entropy H(s, t) can be computed as,
The pair (s*, t*) with highest entropy will be selected as the optimal threshold [33], such that:
Log energy entropy: It is defined as:
Feature organization
Student's t test is used for optimal feature selection. It measures the statistical difference between two sets. This method helps to compute a ratio between the difference of the two class means and the variability of the two classes [34]. The t value indicates the significance of the difference between the two groups, and a significant p value denotes that the sample data does not occur by chance. The t value and p value will be calculated for the features of both classes, and better ranking of features can be done by selecting a higher t value with corresponding lesser p value [35].
Classification
To classify normal versus ICH images, we have used three conventional classifiers, namely, k-nearest neighbour (k-NN), probabilistic neural network (PNN) and support vector machine (SVM) to evaluate feature performance.
k-Nearest neighbour
k-NN is a supervised learning method which assigns a class label to the unknown sample based on its resemblance with the known sample [36]. The k-nearest points in proximity to the unknown data are evaluated, and the most common class among k-nearest neighbours will be the class label for the unknown data [37, 38]. The distance criteria used to measure the resemblance of new test data with the known data is the Euclidean distance. The set of samples with accurate classification should be selected as neighbors to predict the class for the new test data [38].
Probabilistic neural network
PNN is a multiclass classifier which uses a kernel discriminant analysis algorithm to classify the new test data into one of the various classes [39]. This multi-layered feedforward neural network architecture consists of four layers, namely the input layer, pattern layer, summation layer, and output layer [40].
Support vector machine
This supervised classifier is aimed at determining the hyperplane which maximises the margin among the different input classes, which are viewed in n-dimensional space [41]. The hyperplane that forms the maximum separation between the two input classes is selected for classification. Initially, SVM is designed for two-class problems and was extended to address multi-class problems. The Radial Basis Function (RBF) and polynomial kernel are the most frequently used kernel functions utilized by the classifier to solve non-linear boundary problems [42].
ICH indexing
Indexing is employed by several research groups to demonstrate the implication of generated features. As a result, a single number is composed to discriminate healthy versus pathology images [43,44,45,46,47,48,49,50,51]. Hence, we have developed a novel ICH index, which is formulated using only three clinically significant features. The mathematical equation for the ICH index is given as
where \({{\varvec{F}}}_{1}\) is Max. entropy, \({{\varvec{F}}}_{2}\) is Yag. entropy, and \({{\varvec{F}}}_{3}\) is Kap. entropy. The above equation is developed with the assistance of mathematical simulation. All the constant values and the equation itself are formulated empirically to obtain a maximum separation between the normal and ICH classes using a single index value.
Experimental results and discussion
The proposed model is tested on a set of 1603 axial CT images (normal: 784, ICH: 819). Initially, the brain tissue was separated from non-brain tissues (such as skull, fat, etc.) from all the images by thresholding in the pre-processing stage to obtain better classification results. Then, the entropies of each contrast enhanced image are calculated, i.e., in total seven entropy features. Hence, a complete feature set for these brain CT images is generated, and the significance is tested using t values, as shown in Fig. 4. These features are arranged in order of decreasing t value for classification. A further set of seven features are input to the classifier, such as {Max. entropy and Yag. entropy}, {Max. entropy, Yag. entropy, and Kap. entropy},…….., and {Max. entropy, Yag. entropy, Kap. entropy, Ren. entropy, Shan. entropy, Log entropy, and Vaj. entropy}. The classifiers employed in this study are k-NN, PNN, and SVM. The parameters for system evaluation are computed using true positive and negative (TP and TN), and false positive and negative (FP and FN) statistical metrics. For generality, tenfold cross-validation is used. The average value of the tenfold is taken to assess the system performance. k-NN achieved an accuracy of 97.06% for k = 5, PNN achieved an accuracy of 97.37% for \(\sigma =0.03\), and SVM with RBF (i.e., radial basis function) kernel achieved an accuracy of 97% using only six features. Table 1 shows the obtained maximum results for various classifiers using only six features. The complete algorithm is executed under the MATLAB environment using a standalone personal computer.
Discussion
The main goal of this study was to develop a quick-assessment tool for ICH estimation with CT brain imagery. In the current study, the identification of ICH is explored, with an accuracy of 97.37%. It was observed that the entropies are efficiently used to analyse medical data, i.e., 1D signals and 2D signals [52,53,54,55]. Hence, seven entropies were used in our study. These entropies were able to capture the sudden changes in the pixular distribution. They quantify these variations so as to suitably discern the presence of intracerebral haemorrhage in brain CT imagery. It is observed that the system achieved a maximum performance using only six features. Table 2 shows the means and standard deviations (SD) of these features.
It is noted that the first six features are statistically significant with a p value < 0.05, and these features are arranged in descending order using t value. These entropies powerfully distinguish brain CT imagery, and can be analysed using a box plot as shown in Fig. 5. It is noted from Fig. 5 that the range of distributions of the entropy features to characterise brain CT imagery differ. Thus, the features are able to provide clear structural information about imagery and play a key role in performing accurate classification. Therefore, the combination of all six features produces a remarkable performance, shown in Fig. 6. As observed in Table 1, PNN clearly distinguishes normal versus ICH imagery efficiently, with an accuracy of 97.37%, an achieved sensitivity of 96.94%, and a specificity of 97.83%. Both k-NN and SVM classifies the normal versus abnormal group with a sensitivity of 97.43 and 96.21% as compared to PNN. It is also observed that the system reached a PPV of 97.90%. Figure 6 reveals that the performance of the proposed algorithm increases monotonically with the number of features incorporated. To the best of our knowledge, this is the first method which deploys a minimal set of highly discriminative entropy-based non-linear features to classify a large dataset of 1603 CT images. The sensitivity of the system showed that it is robust and can support doctor’s decisions at a high level. Hence, we expect such tools will be incorporated in hospital settings for improved interpretation.
To perform improved discrimination between normal and ICH classes, we have introduced an ICH index using Max. entropy, Yag. entropy, and Vaj. entropy. Figure 7 shows the plot of ICH index for the two classes. It can be observed from Fig. 7 that the range of ICH indexing values provides a clear distinction between normal versus ICH classes. This helps to provide quick and accurate results using a single quantity. Hence, we can conclude as follows. From the result, it is evident that the false positive and negative values are 17 and 25 samples, respectively. It is noted that the specificity approaches 97.83%, which reduces doctor workload by approximately 50%. Furthermore, only three features are employed to develop ICH risk indexing. It is noted that for ICH, mean values of Max. entropy and Kap. entropy are high when compared to normal brain images. This helps to formulate a unique equation to enhance the level of discernment between the two classes. Furthermore, the proposed methodology can be used to analyse medical images with various modalities. In the future, we would like to develop a generalised risk index using a greater number of images.
Several studies have designed a CAD tool for assessment of ICH using brain CT imagery [56,57,58]. A summary of these methods is shown in Table 3. Most of the approaches are segment-based; however the proposed approach is feature-based. Hence, it avoids localization of the region of interest. Our proposed method requires minimal features to model a large set of data. After validating the algorithm with a greater number of subjects, the proposed method can be used as an assistive tool for clinicians in hospital and polyclinic settings. In the future, we intend to extend this work by incorporating deep learning models with an even larger data set, as deep learning models outperform all conventional machine-learning models [59,60,61,62]. Figure 8 shows the proposed architecture for future digital healthcare using the Internet of Things (IoT) for detecting ICH, wherein patients will receive an instant diagnostic result to their mobile phone numbers through cloud-based services. This will automatically reduce the overall turnaround time for diagnosis and decision-making.
To summarize the advantages of the proposed model:
-
It is a feature-based method requiring a minimal number of features, and can be implemented in systems with limited resources. It avoids the localization of clinical features for brain CT images.
-
Speedup of the estimation is done using ICH indexing, which helps to categorise brain CT imagery with a unique value.
Conclusion
We have developed a fully automated system for early diagnosis and management of intracerebral haemorrhage in CT imagery. The technique, which consists of extraction of a minimal set of entropy features and supervised learning methods (k-NN, PNN and SVM), can clearly discern normal versus ICH imagery. Four evaluation metrics were used to assess system performance, and the PNN was found to successfully classify CT imagery with an accuracy of 97.37%. This shows that our approach can perform better as compared to existing state-of-the-art methods. Furthermore, the developed ICH indexing offers a swift, cost effective, and highly accurate solution for identification of ICH lesions. Moreover, this method simplifies the entire diagnostic procedure, and enables doctors to analyse a large number of CT scans in quick succession with highly accurate and consistent results. We would like to extend this work by incorporating more subjects, as well as by including additional features and deeper convolutional neural network techniques.
References
Al-Mufti F, Thabet AM, Singh T, El-Ghanem M, Amuluru K, Gandhi CD (2018) Clinical and radiographic predictors of intracerebral hemorrhage outcome. Interv Neurol 7:118–136
Wijman CAC, Venkatasubramanian C, Bruins S, Fischbein N, Schwartz N (2010) Utility of early MRI in the diagnosis and management of acute spontaneous intracerebral hemorrhage. Cerebrovasc Dis 30:456–463
Gorelick PB (2019) The global burden of stroke: persistent and disabling. Lancet Neurol 18(5):417–418
Krishnan K, Mukhtar SF, Lingard J, Houlton A, Walker E, Jones T, Sprigg N, Cala LA, Becker JL, Dineen RA, Koumellis P (2015) Performance characteristics of methods for quantifying spontaneous intracerebral haemorrhage: data from the Efficacy of Nitric Oxide in Stroke (ENOS) trial. J Neurol Neurosurg Psychiatry 86(11):258–1266
Patel A, Schreuder FH, Klijn CJ, Prokop M, van Ginneken B, Marquering HA, Roos YB, Baharoglu MI, Meijer FJ, Manniesing R (2019) Intracerebral haemorrhage segmentation in non-contrast CT. Sci Rep 9(1):1–11
Sharif M, Tanvir U, Munir EU, Khan MA, Yasmin M (2018) Brain tumor segmentation and classification by improved binomial thresholding and multi-features selection. J Ambient Intell Humaniz Comput. https://doi.org/10.1007/s12652-018-1075-x
Khan SA, Khan MA, Song OY, Nazir M (2020) Medical imaging fusion techniques: a survey benchmark analysis, open challenges and recommendations. J Med Imaging Health Inform 10(11):2523–2531
Naheed N, Shaheen M, Khan SA, Alawairdhi M, Khan MA (2020) Importance of features selection, attributes selection, challenges and future directions for medical imaging data: a review. Comput Model Eng Sci 125(1):314–344
Hussain UN, Khan MA, Lali IU, Javed K, Ashraf I, Tariq J, Ali H, Din A (2020) A Unified design of ACO and skewness based brain tumor segmentation and classification from MRI scans. J Control Eng Appl Inform 22(2):43–55
Zahoor S, Lali IU, Khan MA, Javed K, Mehmood W (2020) Breast cancer detection and classification using traditional computer vision techniques: a comprehensive review. Curr Med Imaging. https://doi.org/10.2174/1573405616666200406110547
Nasir M, Khan MA, Sharif M, Javed MY, Saba T, Ali H, Tariq J (2020) Melanoma detection and classification using computerized analysis of dermoscopic systems: a review. Curr Med Imaging 16(7):794–822
Nazar U, Khan MA, Lali IU, Lin H, Ali H, Ashraf I, Tariq J (2020) Review of automated computerized methods for brain tumor segmentation and classification. Curr Med Imaging 16(7):823–834
Sharif MI, Li JP, Khan MA, Saleem MA (2020) Active deep neural network features selection for segmentation and recognition of brain tumors using MRI images. Pattern Recognit Lett 129:181–189
Khan MA, Rubab S, Kashif A, Sharif MI, Muhammad N (2020) An integrated design of contrast based classical features fusion and selection. Pattern Recognit Lett 129:77–85
Kakhandaki N, Kulkarni SB (2020) Identification of normal and abnormal brain hemorrhage on magnetic resonance images. Cogn Inform Comput Model Cogn Sci Academic Press 1:71–91
Khan MA, Sarfraz MS, Alhaisoni M, Albesher AA, Wang S, Ashraf I (2020) StomachNet: optimal deep learning features fusion for stomach abnormalities classification. IEEE Access 8:197969–197981
Prakash KNB, Zhou S, Morgan TC, Hanley DF, Nowinski WL (2012) Segmentation and quantification of intra-ventricular/cerebral hemorrhage in CT scans by modified distance regularized level set evolution technique. Int J Comput Assist Radiol Surg 7(5):785–798
Shahangian B, Pourghassem H (2016) Automatic brain hemorrhage segmentation and classification algorithm based on weighted grayscale histogram feature in a hierarchical classification structure. Biocybern Biomed Eng 36(1):217–232
Gillebert CR, Humphreys GW, Mantini D (2014) Automated delineation of stroke lesions using brain CT images. Neuroimage Clin 4:540–548
Muschelli J, Sweeney EM, Ullman NL, Vespa P, Hanley DF, Crainiceanu CM (2017) PItcHPERFeCT: primary intracranial hemorrhage probability estimation using random forests on CT. NeuroImage Clin 14:379–439
Al-Ayyoub M, Alawad D, Al-Darabsah K, Aljarrah I (2013) Automatic detection and classification of brain hemorrhages. WSEAS Trans Comput 12(10):395–405
Zhang Y, Chen M, Qingmao Hu, Huang W (2013) Detection and quantification of intracerebral and intraventricular hemorrhage from computed tomography images with adaptive thresholding and case-based reasoning. Int J Comput Assist Radiol Surg 8(6):917–927
Kumar I, Bhatt C, Singh KU (2020) Entropy based automatic unsupervised brain intracranial hemorrhage segmentation using CT images. J King Saud Univ Comput Inf Sci
Phan A-C, Vo V-Q, Phan T-C (2019) A Hounsfield value-based approach for automatic recognition of brain haemorrhage. J Inf Telecommun 3(2):196–209
Gautam A, Raman B (2019) Automatic segmentation of intracerebral hemorrhage from brain CT images. Machine intelligence and signal analysis. Springer, Berlin, pp 753–764
Nag MK, Chatterjee S, Sadhu AK, Chatterjee J, Ghosh N (2019) Computer-assisted delineation of hematoma from CT volume using autoencoder and Chan Vese model. Int J Comput Assist Radiol Surg 14(2):259–269
Foo YH, Wong JHD, Azman RR, Leong YL, Tan LK (2020) Identification of acute intracranial bleed on computed tomography using computer aided detection. J Phys Conf Ser 1497:012019
Sharma B, Venugopalan K (2014) Classification of hematomas in brain CT images using neural network. In: 2014 international conference on issues and challenges in intelligent computing techniques (ICICT), IEEE, pp 41–46
Pizer SM, Amburn EP, Austin JD et al (1987) Adaptive histogram equalization and its variations. Comput Vis Graph Image Process 39:355–368
Pharwaha APS, Singh B (2009) Shannon and non-shannon measures of entropy for statistical texture feature extraction in digitized mammograms. In: Proceedings of the world congress on engineering and computer science, vol 2, pp 20–22
Acharya UR, Mookiah MRK, Sree SV, Yanti R, Martis RJ, Saba L, Molinari F, Guerriero S, Suri JS (2014) Evolutionary algorithm-based classifier parameter tuning for automatic ovarian cancer tissue characterization and classification. Ultraschall in der Medizin Eur J Ultrasound 35(3):237–245
Sen H, Agarwal A (2017) A comparative analysis of entropy based segmentation with Otsu method for grey and color images. In International conference of electronics, communication and aerospace technology (ICECA), vol 1, pp 113–118
Zhou M, Hong X, Tian Z, Dong H, Wang M, Xu K (2014) Maximum entropy threshold segmentation for target matching using speeded-up robust features. J Electr Comput Eng 2014:1–12
Zhou N, Wang L (2007) A modified t-test feature selection method and its application on the hapmap genotype data. Geno Prot Bioinform 5(3–4):242–249
Available in: https://www.statisticshowto.datasciencecentral.com/probability-and-statistics/t-test. Last Accessed 29 Apr 2020
Larose DT (2004) Discovering knowledge in data: an introduction to data mining. Wiley-Interscience, New York
Han J, Pei J, Kamber M (2011) Data mining: concepts and techniques. Elsevier, Amsterdam
Acharya UR, Fujita H, Sudarshan VK, Sree VS, Eugene LWJ, Ghista DN, Tan RS (2015) An integrated index for detection of sudden cardiac death using discrete wavelet transform and nonlinear features. Knowl Based Syst 83:149–158
Specht DF (1990) Probabilistic neural networks. Neural Netw 3(1):109–118
Kecman V (2001) Learning and soft computing. MIT Press, Cambridge
Hastie T, Tibshirani R, Friedman J (2008) The elements of statistical learning, 2nd edn. Springer, New York
Christianini N, Shawe-Taylor J (2000) An introduction to support vector machines and other kernel-based learning methods. Cambridge University Press, Cambridge
Ghista DN (2009) Nondimensional physiological indices for medical assessment. J Mech Med Biol 9:643–669
Acharya UR, Raghavendra U, Fujita H, Hagiwara Y, Koh JEW, Hong TJ, Sudarshan VK, Vijayananthan A, Yeong CH, Gudigar A, Ng KH (2016) Automated characterization of fatty liver disease and cirrhosis using curvelet transform and entropy features extracted from ultrasound images. Comput Biol Med 79:250–258
Fujita H, Acharya UR, Sudarshan VK, Ghista DN, Sree SV, Eugene LWJ, Koh JEW (2016) Sudden cardiac death (SCD) prediction based on nonlinear heart rate variability features and SCD index. Appl Soft Comput 43:510–519
Acharya UR, Fujita H, Sudarshan VK, Mookiah MRK, Koh JE, Tan JH, Hagiwara Y, C. KC, J. S. Padmakumar, A. Vijayananthan, K. H. Ng, (2015) An integrated index for identification of fatty liver disease using radon transform and discrete cosine transform features in ultrasound images. Inf Fusion 31:43–53
Raghavendra U, Rajendra Acharya U, Ng EYK, Tan J-H, Gudigar A (2016) An integrated index for breast cancer identification using histogram of oriented gradient and kernel locality preserving projection features extracted from thermograms. Quant Infrared Thermogr 13(2):195–209
Raghavendra U, Fujita H, Gudigar A, Shetty R, Nayak K, Pai U, Jyothi Samanth U, Acharya R (2017) Automated technique for coronary artery disease characterization and classification using DD-DTDWT in ultrasound images. Biomed Signal Process Control 40:324–334
Raghavendra U, Rajendra Acharya U, Gudigar A, Tan JH, Fujita H, Hagiwara Y, Molinari F, Kongmebhol P, Ng KH (2017) Fusion of spatial grey level dependency and fractal texture features for the characterization of thyroid lesions. Ultrasonics 77:202–210
Acharya UR, Mookiah MR, Koh JE, Tan JH, Noronha K, Bhandary SV, Rao AK, Hagiwara Y, Chua CK, Laude A (2016) Novel risk index for the identification of age-related macular degeneration using radon transform and DWT features. Comput Biol Med 73:131–140
Acharya UR, Fujita H, Sudarshan VK, Mookiah MR, Koh JE, Tan JH, Hagiwara Y, Chua CK, Junnarkar SP, Vijayananthan A, Ng KH (2016) An integrated index for identification of fatty liver disease using radon transform and discrete cosine transform features in ultrasound images. Inf Fusion 31:43–53
Vranković A, Lerga J, Saulig N (2020) A novel approach to extracting useful information from noisy TFDs using 2D local entropy measures. EURASIP J Adv Signal Process 1:1–19
Borowska M (2015) Entropy-based algorithms in the analysis of biomedical signals. Stud Logic Gramm Rhetor 43(1):21–32
Cornforth DJ, Tarvainen MP, Jelinek HF (2013) Using Renyi entropy to detect early cardiac autonomic neuropathy. In: Engineering in medicine and biology society (EMBC), 35th annual international conference of the IEEE, pp 5562–5565
Liang Z, Wang Y, Sun X, Li D, Voss LJ, Sleigh JW, Li X et al (2015) EEG entropy measures in anaesthesia. Front Comput Neurosci. https://doi.org/10.3389/fncom.2015.00016
Subudhi A, Jena SS, Sabut SK (2018) Delineation of the ischemic stroke lesion based on watershed and relative fuzzy connectedness in Brain MRI. Med Biol Eng Comput 56(5):795–807
Subudhi A, Acharya UR, Dash M, Jena S, Sabut SK (2018) Automated approach for detection of ischemic stroke using Delaunay triangulation in brain MRI images. Comput Biol Med 103:116–129
Subudhi A, Dash M, Sabut SK (2020) Automated segmentation and classification of brain stroke using expectation maximization and random forest classifier. Biocybern Biomed Eng 40(1):277–289
Oh SL, Hagiwara Y, Raghavendra U, Yuvaraj R, Arunkumar N, Murugappan M, Acharya UR (2018) A deep learning approach for Parkinson’s disease diagnosis from EEG signals. Neural Comput Appl 1–7 (in press)
Raghavendra U, Fujita H, Bhandary SV, Gudigar A, Tan JH, Acharya UR (2018) Deep convolution neural network for accurate diagnosis of glaucoma using digital fundus images. Inf Sci 441:41–49
Acharya UR, Fujita H, Oh SL, Raghavendra U, Tan JH, Adam M, Gertych A, Hagiwara Y (2018) Automated identification of shockable and non-shockable life-threatening ventricular arrhythmias using convolutional neural network. Future Gener Comput Syst 79:952–959
Tan JH, Bhandary SV, Sivaprasad S, Hagiwara Y, Bagchi A, Raghavendra U, Rao AK, Raju B, Shetty NS, Gertych A, Chua KC, Acharya UR (2018) Age-related macular degeneration detection using deep convolutional neural network. Future Gener Comput Syst 87:127–135
Funding
The authors received no financial support for the research.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
None of the authors have any conflict of interest.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Raghavendra, U., Pham, TH., Gudigar, A. et al. Novel and accurate non-linear index for the automated detection of haemorrhagic brain stroke using CT images. Complex Intell. Syst. 7, 929–940 (2021). https://doi.org/10.1007/s40747-020-00257-x
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s40747-020-00257-x