Abstract
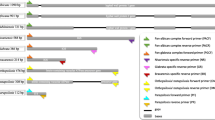
Non-albicans Candida species and other rare yeasts have emerged as major opportunistic pathogens in fungal infections. Identification of opportunistic yeasts in developing countries is mainly performed by phenotypic assay, which are time-consuming and prone to errors. The aim of the present study was to evaluate PCR-RFLP as a routinely used identification technique for the most clinically important Candida species in Iran and make a comparison with a novel multiplex PCR, called 21-plex PCR. One hundred and seventy-three yeast isolates from clinical sources were selected and identified with sequence analysis of the D1/D2 domains of rDNA (LSU rDNA) sequencing as the gold standard method. The results were compared with those obtained by PCR-RFLP using MspI restriction enzyme and the 21-plex PCR. PCR-RFLP correctly identified 93.4% of common pathogenic Candida species (C. albicans, C. glabrata, C. parapsilosis, C. tropicalis, and P. kudriavsevii (= C. krusei)) and was able to identify 45.5% of isolates of the uncommon yeast species compared to the D1/D2 rDNA sequencing. Compared with PCR-RFLP, all common Candida species and 72.7% of uncommon yeast species were correctly identified by the 21-plex PCR. The application of the 21-plex PCR assay as a non-sequence-based molecular method for the identification of common and rare yeasts can reduce turnaround time and costs for the identification of clinically important yeasts and can be applied in resource-limited settings.
Similar content being viewed by others
References
Lamoth F, Lockhart SR, Berkow EL, Calandra T. Changes in the epidemiological landscape of invasive candidiasis. J Antimicrob Chemother. 2018;73:i4–13.
Salehi M, Ahmadikia K, Mahmoudi S, Kalantari S, Jamalimoghadamsiahkali S, Izadi A, et al. Oropharyngeal candidiasis in hospitalised COVID-19 patients from Iran: Species identification and antifungal susceptibility pattern. Mycoses. 2020;63(8):771–8.
Fesharaki SH, Haghani I, Mousavi B, Kargar ML, Boroumand M, Anvari MS, et al. Endocarditis due to a co-infection of Candida albicans and Candida tropicalis in a drug abuser. J Med Microbiol. 2013;62(11):1763–7.
Sadrossadati SZ, Ghahri M, Fooladi AAI, Sayyahfar S, Beyraghi S, Baseri Z. Phenotypic and genotypic characterization of Candida species isolated from candideamia in Iran. Curr Med Mycol. 2018;4(2):14.
Arastehfar A, Daneshnia F, Kord M, Roudbary M, Zarrinfar H, Fang W, et al. Comparison of 21-Plex PCR and API 20C AUX, MALDI-TOF MS, and rDNA sequencing for a wide range of clinically isolated yeast species: improved identification by combining 21-Plex PCR and API 20C AUX as an alternative strategy for developing countries. Front Cell Infect Microbiol. 2019;9:21.
Arastehfar A, Wickes BL, Ilkit M, Pincus DH, Daneshnia F, Pan W, et al. Identification of mycoses in developing countries. J Fungi. 2019;5(4):90.
Aslani N, Janbabaei G, Abastabar M, Meis JF, Babaeian M, Khodavaisy S, et al. Identification of uncommon oral yeasts from cancer patients by MALDI-TOF mass spectrometry. BMC Infect Dis. 2018;18(1):24.
Ahangarkani F, Shokohi T, Rezai MS, Ilkit M, Mahmoodi Nesheli H, Karami H, et al. Epidemiological features of nosocomial candidaemia in neonates, infants and children: a multicentre study in Iran. Mycoses. 2020;63(4):382–94.
Jafari Z, Motamedi M, Jalalizand N. A comparison between CHROMagar, PCR-RFLP and PCR-FSP for identification of Candida species. Curr Med Mycol. 2017;3(3):10–5.
Baghdadi E, Khodavaisy S, Rezaie S, Abolghasem S, Sharifynia S, Kiasat N, et al. Antifungal susceptibility patterns of Candida species recovered from endotracheal tube biofilms in an intensive care unit. Adv Med. 2016. https://doi.org/10.1155/2016/9242031.
Kord M, Salehi MR, Khodavaisy S, Hashemi SJ, Daei R, Rezaei S, et al. Epidemiology of yeast species causing bloodstream infection in Tehran, Iran (2015–2017); Superiority of 21-plex PCR over the Vitek 2 system for yeast identification. J Med Microbiol. 2020;69(5):712–20.
Mirhendi H, Makimura K, Khoramizadeh M, Yamaguchi H. A one-enzyme PCR-RFLP assay for identification of six medically important Candida species. Nippon Ishinkin Gakkai Zasshi. 2006;47(3):225–9.
Munoz JF, Gade L, Chow NA, Loparev VN, Juieng P, Berkow EL, et al. Genomic insights into multidrug-resistance, mating and virulence in Candida auris and related emerging species. Nat Commun. 2018;9(1):1–13.
Arastehfar A, Fang W, Badali H, Vaezi A, Jiang W, Liao W, et al. Low-cost tetraplex PCR for the global spreading multi-drug resistant fungus, Candida auris and its phylogenetic relatives. Front Microbiol. 2018;9:1119.
Leaw SN, Chang HC, Sun HF, Barton R, Bouchara J-P, Chang TC. Identification of medically important yeast species by sequence analysis of the internal transcribed spacer regions. J Clin Microbiol. 2006;44(3):693–9.
Arastehfar A, Fang W, Pan W, Lackner M, Liao W, Badiee P, et al. YEAST PANEL multiplex PCR for identification of clinically important yeast species: stepwise diagnostic strategy, useful for developing countries. Diagn Microbiol Infect Dis. 2019;93(2):112–9.
Ahmed A, Azim A, Baronia A, Marak RS, Gurjar M. Invasive candidiasis in non neutropenic critically ill-need for region-specific management guidelines. Indian J Crit Care Med. 2015;19(6):333.
de Almeida JN, Del Negro GM, Grenfell RC, Vidal MS, Thomaz DY, De Figueiredo DS, et al. Matrix-assisted laser desorption ionization–time of flight mass spectrometry for differentiation of the dimorphic fungal species Paracoccidioides brasiliensis and Paracoccidioides lutzii. J Clin Microbiol. 2015;53(4):1383–6.
Keçeli SA, Dündar D, Tamer GS. Comparison of Vitek matrix-assisted laser desorption/ionization time-of-flight mass spectrometry versus conventional methods in Candida identification. Mycopathologia. 2016;181(1–2):67–73.
Castanheira M, Woosley LN, Diekema DJ, Jones RN, Pfaller MA. Candida guilliermondii and other species of Candida misidentified as Candida famata: assessment by Vitek 2, DNA sequencing analysis, and matrix-assisted laser desorption ionization–time of flight mass spectrometry in two global antifungal surveillance programs. J Clin Microbiol. 2013;51(1):117–24.
Shokohi T, Soteh MH, Pouri ZS, Hedayati M, Mayahi S. Identification of Candida species using PCR-RFLP in cancer patients in Iran. Indian J Med Microbiol. 2010;28(2):147.
Hasanvand S, Azadegan Qomi H, Kord M, Didehdar M. Molecular epidemiology and in vitro antifungal susceptibility of candida isolates from women with vulvovaginal candidiasis in northern cities of Khuzestan province, Iran. Jundishapur J Microbiol. 2017;10(8):e12804.
Mohammadi R, Mirhendi H, Yadegari MH, Ghahri M, Shadzi S, Jalalizand N, et al. Evaluation of Prevalence of the New Candida Species (C. orthopsilosis and C. metapsilosis) among C. parapsilosis Group Isolated from Patients by PCR-RFLP of SADH Gene in Iran. J Isfahan Med School. 2011;29(134):1–10.
Mohammadi R, Mirhendi H, Rezaei-Matehkolaei A, Ghahri M, Shidfar MR, Jalalizand N, et al. Molecular identification and distribution profile of Candida species isolated from Iranian patients. Med Mycol. 2013;51(6):657–63.
Khan Z, Ahmad S, Al-Sweih N, Khan S, Joseph L. Candida lusitaniae in Kuwait: prevalence, antifungal susceptibility and role in neonatal fungemia. PLoS ONE. 2019;14(3):e0213532.
Asner SA, Giulieri S, Diezi M, Marchetti O, Sanglard D. Acquired multidrug antifungal resistance in Candida lusitaniae during therapy. Antimicrob Agents Chemother. 2015;59(12):7715–22.
Atkinson BJ, Lewis RE, Kontoyiannis DP. Candida lusitaniae fungemia in cancer patients: risk factors for amphotericin B failure and outcome. Med Mycol. 2008;46(6):541–6.
Mirhendi H, Makimura K, Zomorodian K, Maeda N, Ohshima T, Yamaguchi H. Differentiation of Candida albicans and Candida dubliniensis using a single-enzyme PCR-RFLP method. Jpn J Infect Dis. 2005;58(4):235.
Loreto ÉS, Scheid LA, Nogueira CW, Zeni G, Santurio JM, Alves SH. Candida dubliniensis: epidemiology and phenotypic methods for identification. Mycopathologia. 2010;169(6):431–43.
Nikmanesh B, Ahmadikia K, Getso MI, Aghaei S. Candida africana and Candida dubliniensis as causes of pediatric candiduria: A study using HWP1 gene size polymorphism. AIMS Microbiol. 2020;6(3):272.
Davies GE, Thornton CR. Differentiation of the emerging human pathogens Trichosporon asahii and Trichosporon asteroides from other pathogenic yeasts and moulds by using species-specific monoclonal antibodies. PLoS ONE. 2014;9(1):e84789.
Acknowledgements
The authors would like to thank Mohammad Reza Safari and Azar Berahmeh for their assistance during the study.
Funding
This study was part of a PhD. dissertation (grant No. 9511353003) and M.S. dissertation (grant No. 9711352002) supported by Tehran University of Medical Sciences. This study has been funded and supported by Tehran University of Medical Sciences (TUMS); grant no. 99-1-99-46502.
Author information
Authors and Affiliations
Contributions
SK, SJH, RDG, and MK designed the study. AE, SK, MK, and AA collected yeast isolates. MK, MRS, and AA collected clinical data. MK, AE, AA, and MG performed molecular identification. SK, SR, TB, RDG, and SJH performed data analysis. MK, SK, MG, and AE prepared the first draft. All authors contributed in revision.
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Additional information
Handling Editor: Nancy L. Wengenack
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
About this article
Cite this article
Kord, M., Elmimoghaddam, A., Hashemi, S.J. et al. Comparison of PCR-RFLP with 21-plex PCR and rDNA Sequencing for Identification of Clinical Yeast Isolates. Mycopathologia 186, 213–220 (2021). https://doi.org/10.1007/s11046-020-00522-0
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11046-020-00522-0