Abstract
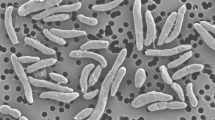
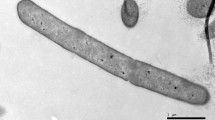
An endospore producing, strict aerobic, Gram-stain-positive, orange-colored colony forming bacterium designated as strain JC1013T was isolated from an orange pond near a solar saltern of Tamil Nadu, India. Phylogenetic analysis of the 16S rRNA gene sequences indicated that strain was affiliated to the family Bacillaceae of the phylum Firmicutes. Strain showed highest 16S rRNA gene sequence identity of 98.7% with Mesobacillus selenatarsenatis SF-1 T and below 98.3% with other members of the genus Mesobacillus. Strain JC1013T produced carotenoid pigments and indole compounds. Major cellular fatty acids of strain JC1013T were iso-C15:0, anteiso-C15:0, C16:0 3-OH, iso-C17:0ω10c and summed feature 4 (iso-C17:1 I/ anteisoB). Polar lipids were diphosphatidylglycerol, phosphatidylethanolamine, phosphatidylglycerol, two unidentified aminolipids and four unidentified phospholipids. Strain JC1013T constituted m-diaminopimelic acid as diagnostic cell wall amino acids. MK-7 is the predominant menaquinone of strain JC1013T. The genome size of strain JC1013T was 4.6 Mbp and its G + C content was 42.7 mol%. For the affirmation of strain’s taxonomic status, a detailed phylogenomic study was done. Based on the phylogenetic analyses, low ANI (84.6%), AAI (88.5%) values, in-silico DDH (< 29%) value, morphological, physiological and chemo-taxonomical characteristics, strain JC1013T was clearly distinguished from the nearest phylogenetic neighbor, Mesobacillus selenatarsenatis SF-1T to conclude that it is a new species of the genus Mesobacillus. We propose the name as Mesobacillus aurantius with type strain JC1013T (= NBRC 114146T = KACC 21451 T).


Similar content being viewed by others
Abbreviations
- NCBI:
-
National centre for biotechnology information
- GCM:
-
The global catalogue of microorganisms
- ANI:
-
Average nucleotide identity
- AAI:
-
Average amino identity
- dDDH:
-
Digital DNA–DNA hybridization
- BLAST:
-
Basic local alignment search tool
- MUSCLE:
-
Multiple sequence comparison by log-expectation
- TLC:
-
Thin-layer chromatography
- HPLC:
-
High-pressure liquid chromatography
- NBRC:
-
Biological resource center
- NITE:
-
KACC, Korean agricultural culture collection
- CMC:
-
Carboxymethyl cellulose
References
Ahmed I, Yokota A, Fujiwara T (2007) A novel highly boron tolerant bacterium, Bacillus boroniphilus sp. nov., isolated from soil, that requires boron for its growth. Extremophiles 11:217–224
Auch AF, Klenk HP, Göker M (2010) Standard operating procedure for calculating genome-to-genome distances based on high-scoring segment pairs. Stand Genomic Sci 2:142–148
Deepshikha G, Mujahid M, Prasuna M et al (2019) iTRAQ-based quantitative proteomics reveals insights into metabolic and molecular responses of glucose-grown cells of Rubrivivax benzoatilyticus JA2. J Prot 194:49–59
Divyasree B, Suresh G, Sasikala C, Ramana C (2017) Chryseobacterium salipaludis sp. nov., isolated at a wild ass sanctuary. Int J Syst Evol Microbiol 68:542–546
Edgar RC (2004) MUSCLE: multiple sequence alignment with high accuracy and high throughput. Nucleic Acids Res 32:1792–1797
Ehmann A (1977) The van urk-salkowski reagent - a sensitive and specific chromogenic reagent for silica gel thin-layer chromatographic detection and identification of indole derivatives. J Chromatogr 132:267–276
Felsenstein J (1985) Confidence limits on phylogenies: an approach using the bootstrap. Evolution 39:783–791
Gandham S, Lodha T, Sasikala Ch, Ramana ChV (2018) Rhodobacter alkalitolerans sp. nov., isolated from an alkaline brown pond. Arch Microbiol 200:1487–1492
Garabito MJ, Marquez MC, Ventosa A (1998) Halotolerant Bacillus diversity in hypersaline environments. Can J Microbiol 44:95–102
Kanso S, Greene AC, Patel BKC (2002) Bacillus subterraneus sp. nov., an iron- and manganese-reducing bacterium from a deep subsurface Australian thermal aquifer. Int J Syst Evol Microbiol 52:869–874
Kates M (1972) Techniques in Lipidology. In Laboratory Techniques in Biochemistry and Molecular Biology. American Elsevier Publishing Company 3: part 2, 355–356
Kates M (1986) Techniques of lipidology. Isolation, analysis and identification of lipids. In: Burdon RH, van Knippenberg PH (eds) Laboratory techniques in biochemistry and molecular biology, vol 3. part 2. Elsevier, Amsterdam, pp 100–112
Khaneja R, Perez-Fons L, Fakhry S et al (2010) Carotenoids found in Bacillus. J Appl Microbiol 108:1889–1902
Kumar RM, Kaur G, Kumar A et al (2015) Taxonomic description and genome sequence of Bacillus campisalis sp. nov., a member of the genus Bacillus isolated from a solar saltern. Int J Syst Evol Microbiol 65:3235–3240
Kumar S, Karan R, Kapoor S et al (2012) Screening and isolation of halophilic bacteria producing industrially important enzymes. Braz J Microbiol 43:1595–1603
Kumar S, Glen S, Koichiro T (2016) MEGA7: molecular evolutionary genetics analysis version 7.0 for bigger datasets. Mol BiolEvol 33:1870–1874
Kimura M (1980) A simple method for estimating evolutionary rate of base substitutions through comparative studies of nucleotide sequences. J Mol Evol 16:111–120
Lakshmi KVNS, Sasikala Ch, Ashok Kumar GV et al (2011) Phaeovibrio sulfidiphilus gen. nov. sp. nov., phototrophic alphaproteobacterium isolated from brakish water. Int J Syst Evol Microbiol 61:828–832
Levasseur A, Drula E, Lombard V et al (2013) Expansion of the enzymatic repertoire of the CAZy database to integrate auxiliary redox enzymes. Biotechnol Biofuels 6:41–45
Logan NA, Berge O, Bishop AH et al (2009) Proposed minimal standards for describing new taxa of aerobic, endospore-forming bacteria. Int J Syst Evol Microbiol 59:2114–2121
Luo R, Liu B, Xie Y, Li Z, Huang W et al (2012) SOAPdenovo2: an empirically improved memory-efficient short-read de novo assembler. Giga Sci. https://doi.org/10.1186/2047-217X-1-18
Mai-Gisondi G, Maaheimo H, Chong SL et al (2017) Functional comparison of versatile carbohydrate esterases from families CE1, CE6 and CE16 on acetyl-4-O-methylglucuronoxylan and acetyl-galactoglucomannan. Biochimica et Biophysica Acta (BBA)-Gen Subj 9:2398–2405
McKerrow J, Vagg S, McKinney T et al (2000) A simple HPLC method for analysing diaminopimelic acid diastereomers in cell walls of Gram-positive bacteria. Lett Appl Microbiol 30:178–182
Meier-Kolthoff JP, Göker M (2019) TYGS is an automated high-throughput platform for state-of-the-art genome-based taxonomy. Nat Commun 10:2182
Mollan RC, Harmey MA, Bonnely DMX (1973) UV spectra of indoles in strong sulphuric acid. Phytochemistry 12:447–450
Mujahid M, Sasikala C, Ramana ChV (2010) Aniline-induced tryptophan production and identification of indole derivatives from three purple bacteria. Curr Microbiol 61:285–290
Na S, Kim YO, Yoon SH et al (2018) UBCG: up-to-date bacterial core gene set and pipeline for phylogenomic tree reconstruction. J Microbiol 56:280–285
Oren A (2002) Diversity of halophilic microorganisms; environments, phylogeny, physiology and applications. J Ind Microbial Biotechnol 28:56–63
Oren A (2008) Microbial life at high salt concentrations: phylogenetic and metabolic diversity. Saline Syst 4:38–40
Pal D, Mathan Kumar R, Kaur N et al (2017) Bacillus maritimus sp. nov., a novel member of the genus Bacillus isolated from marine sediment. Int J Syst Evol Microbiol 67:60–66
Patel S, Gupta RS (2019) A phylogenomic and comparative genomic framework for resolving the polyphyly of the genus Bacillus: proposal for six new genera of Bacillus species, Peribacillusgen. nov., Cytobacillus gen. nov., Mesobacillus gen. nov., Neobacillus gen. nov., Metabacillus gen. nov., and Alkalihalobacillus gen. nov. Int J Syst Evol Microbiol 70:406–438
Pérez-Ibarra BM, Flores ME, García-Varela M (2007) Isolation and characterization of Bacillus thioparus sp. nov., chemolithoautotrophic, thiosulfate-oxidizing bacterium. FEMS Microbiol Lett 271:289–296
Richter M, Rosselló-Móra R (2009) Shifting the genomic gold standard for the prokaryotic species definition. Proc Natl Acad Sci USA 106:19126–19131
Rodriguez-R LM, Konstantinidis KT (2016) The enveomics collection: a toolbox for 4 specialized analyses of microbial genomes and metagenomes. Peer J Preprints 4:e1900v1
Rosselló-Móra R, Amann R (2015) Past and future species definitions for bacteria and Archaea. Syst Appl Microbiol 38:209–216
Sato T, Yoshida S, Hoshino H et al (2011) Sesquarterpenes (C35 terpenes) biosynthesized via the cyclization of a linear C35 isoprenoid by a tetraprenyl-β-curcumene synthase and a tetraprenyl-β-curcumene cyclase: identification of a new terpene cyclase. J Am Chem Soc 25:9734–9737
Sandmann G (2019) Antioxidant protection from UV and light-stress related to carotenoid structures. Antioxidants 8:219–232
Sasser M (1990) Identification of bacteria by gas chromatography of cellular fatty acids. MIDI Technical note 101. MIDI Inc, Newark, DE
Schleifer KH (1985) Analysis of the chemical composition and primary structure of murein. Meth Microbiol 18:123–156
Schleifer KH, Kandler O (1972) Peptidoglycan types of bacterial cell walls and their taxonomic implications. Bacteriol Rev 36:407–477
Smibert RM, Krieg NR (1981) General characterization. In: Gerhardt P, Murray RGE, Costilow RN, Nester EW, Wood WA et al. (eds.), Manual of Methods for General Microbiology. Washington, DC: Am Society Microbiol 24:409–443
Tindall BJ (1990a) Lipid composition of Halobacterium lacusprofundi. FEMS Microbiol Lett 66:199–202
Tindall BJ (1990b) A comparative study of the lipid composition of Halobacterium saccharovorum from various sources. Syst Appl Microbiol 13:128–130
Wattam AR, Davis JJ, Assaf R et al (2017) Improvements to PATRIC, the all-bacterial bioinformatics database and analysis resource center. Nucleic Acids Res 45:535–542
Wu L, Ma J (2019) The global catalogue of microorganisms (GCM) 10K type strain sequencing project: providing services to taxonomists for standard genome sequencing and annotation. Int J Syst Evol Microbiol 69:895–898
Xie C, Yokota A (2003) Phylogenetic analysis of Lampropedia hyaline based on the 16S rRNA gene sequence. J Gen Appl Microbiol 49:345–349
Yamamura S, Yamashita M, Fujimoto N et al (2007) Bacillus selenatarsenatis sp. nov., a selenate- and arsenate-reducing bacterium isolated from the effluent drain of a glass-manufacturing plant. Int J Syst Evol Microbiol 57:1060–1064
Yoon JH, Kang SS, Lee KC et al (2001) Bacillus jeotgali sp. nov., isolated from jeotgal, Korean traditional fermented seafood. Int J Syst Evol Microbiol 51:1087–1092
Yoon SH, Ha SM, Kwon S et al (2017) Introducing EzBioCloud: a taxonomically united database of 16S rRNA and whole genome assemblies. Int J Syst Evol Microbiol 67:1613–1617
Acknowledgements
Anusha is grateful to the Department of Sciences and Technology, Government of India for the award of JRF/SRF Inspire fellowship. Ch.V.R. thanks the Department of Biotechnology, Government of India, for the award of Tata Innovative fellowship.Ch. S thank UGC, Government of India for Mid-Career award grant. Jagadeeshwari thanks TEQIP for research fellowship. All authors thank the BGI group (GCM 10K type strain sequencing project), China for genome sequencing of strain JC1013T. Financial support received from Council for Scientific and Industrial Research (CSIR), University Grants commission (UGC), All India Council for Technical Education (AICTE) New Delhi are acknowledged. Infrastructure facilities funded by DST-FIST, UGC-SAP (DRS) and TEQIP are acknowledged.
Author information
Authors and Affiliations
Corresponding authors
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Additional information
Communicated by Erko Stackebrandt.
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
The GenBank/EMBL/DDBJ accession number for 16S rRNA gene sequence of strain JC1013T is LS998022. The GenBank/EMBL/DDBJ accession for the whole genome shotgun sequence is JAAVUM000000000.
Supplementary information
Rights and permissions
About this article
Cite this article
Rai, A., Smita, N., Shabbir, A. et al. Mesobacillus aurantius sp. nov., isolated from an orange-colored pond near a solar saltern . Arch Microbiol 203, 1499–1507 (2021). https://doi.org/10.1007/s00203-020-02146-w
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00203-020-02146-w