Abstract
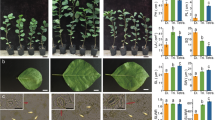
Studies have shown that polyploidy has pronounced effects on plants in multiple aspects, including genome structure, gene expression, metabolism and so on, which finally induce changes in phenotype. Many of these changes occurred immediately after polyploidization and retained or lost later. Therefore, it is meaningful for crop breeders to gain knowledge of these changes when it is relatively stable. In this study, we investigated the phenotypes of a highly self-pollinated synthesized allotetraploid in Cucumis, named with Cucumis × hytivus J. F. Chen & J. H. Kirkbr. (C. ×hytivus for short). Results showed that although many phenotypes of C. ×hytivus were intermediate between its parents, total leaf area and cell size exhibited parent-of-origin and dosage effect, respectively. Additionally, C. ×hytivus exhibited divergent biomass allocation strategy compared to the parents, developing more leaves. Beside the commonalities across different polyploid systems, the combination of hybridity and genome duplication in allopolyploids may lead to a diverse possibility of phenotypic changes. In spite of the reduced light absorption by less photosynthetic pigments in young leaves, allopolyploidy caused limited adverse effect on the photosynthesis of C. ×hytivus, which may benefit from the increased leaf thickness and potentially facilitated the survival and speciation of this novel species. The present study offers novel insights into the varied phenotypic effects of polyploidy and is a valuable reference for the crop improvement through polyploidization.






Similar content being viewed by others
Reference
Bassene JB, Froelicher Y, Dhuique-Mayer C, Mouhaya W, Ferrer RM, Ancillo G, Morillon R, Navarro L, Ollitrault P (2009) Non-additive phenotypic and transcriptomic inheritance in a citrus allotetraploid somatic hybrid between C. reticulata and C. limon: the case of pulp carotenoid biosynthesis pathway. Plant Cell Rep 28:1689
Bretagnolle F, Thompson JD, Lumaret R (1995) The influence of seed size variation on seed germination and seedling vigour in diploid and tetraploid dactylis glomerata L. Ann Bot 76:607–615
Cao M, Zou X, Warren M, Zhu H (2006) Tropical forests of xishuangbanna, China. Biotropica 38:306–309
Chen L, Chen J (2008) Changes of cytosine methylation induced by wide hybridization and allopolyploidy in Cucumis. Genome 51:789–799
Chen JF, Kirkbride JH (2000) A new synthetic species of Cucumis (Cucurbitaceae) from interspecific hybridization and chromosome doubling. Brittonia 52:315–319
Chen J, Zhang SL, Zhang X (1994) The Xishuangbanna gourd (Cucumis sativus var. xishuangbannesis Qi et Yuan), a traditionally cultivated plant of the Hanai People, Xishuangbanna, Yunnan, China. Cucurbit Genet Coop Rep 17:18–20
Chen JF, Staub J, Adelberg J, Lewis S, Kunkle B (2002) Synthesis and preliminary characterization of a new species (amphidiploid) in Cucumis. Euphytica 123:315–322
Chen J, Staub J, Qian C, Jiang J, Luo X, Zhuang F (2003) Reproduction and cytogenetic characterization of interspecific hybrids derived from Cucumis hystrix Chakr. × Cucumis sativus L. Theor Appl Genet 106:688–695
Chen L, Lou Q, Zhuang Y, Chen J, Zhang X, Wolukau JN (2007) Cytological diploidization and rapid genome changes of the newly synthesized allotetraploids Cucumis × hytivus. Planta 225:603–614
Chen ZJ (2010) Molecular mechanisms of polyploidy and hybrid vigor. Trends Plant Sci 15:57–71
Coate JE, Doyle JJ (2013) Genomics and transcriptomics of photosynthesis in polyploids. In: Chen ZJ, Birchler JA (eds) Polyploid and hybrid genomics. Wiley, Oxford, pp 153–169
Dijkhuizen A, Kennard WC, Havey MJ, Staub JE (1996) RFLP variation and genetic relationships in cultivated cucumber. Euphytica 90:79–87
Drake PL, Froend RH, Franks PJ (2013) Smaller, faster stomata: scaling of stomatal size, rate of response, and stomatal conductance. J Exp Bot 64:495–505
Gates DM, Keegan HJ, Schleter JC, Weidner VR (1965) Spectral properties of plants. Appl Optic 4:11–20
Ha M, Kim ED, Chen ZJ (2009a) Duplicate genes increase expression diversity in closely related species and allopolyploids. Proc Natl Acad Sci USA 106(7):2295–2300
Ha M, Lu J, Tian L, Ramachandran V, Kasschau KD, Chapman EJ, Carrington JC, Chen X, Wang XJ, Chen ZJ (2009) Small RNAs serve as a genetic buffer against genomic shock in Arabidopsis interspecific hybrids and allopolyploids. Proc Natl Acad Sci USA 106(42):17835–17840
Huang S, Li R, Zhang Z et al (2009) The genome of the cucumber, Cucumis sativus L. Nat Genet 41:1275–1281
Ježilová E, Nožková-Hlaváčková V, Duchoslav M (2014) Photosynthetic characteristics of three ploidy levels of Allium oleraceum L. (Amaryllidaceae) differing in ecological amplitude. Plant Spec Biol 30:212–224
Jiao Y, Wickett NJ, Ayyampalayam S, Chanderbali AS, Landherr L, Ralph PE, Tomsho LP, Hu Y, Liang H, Soltis PS, Soltis DE, Clifton SW, Schlarbaum SE, Schuster SC, Ma H, Leebens-Mack J, dePamphilis CW (2011) Ancestral polyploidy in seed plants and angiosperms. Nature 473:97–100
Jiao W, Yuan J, Jiang S, Liu Y, Wang L, Liu M, Zheng D, Ye W, Wang X, Chen ZJ (2018) Asymmetrical changes of gene expression, small RNAs and chromatin in two resynthesized wheat allotetraploids. Plant J 93:828–842
Jung Y, Kawaura K, Kishii M, Sakuma S, Ogihara Y (2015) Comparison of genome-wide gene expression patterns in the seedlings of nascent allohexaploid wheats produced by two combinations of hybrids. Genes Genet Syst 90(2):79–88
Lashermes P, Combes MC, Hueber Y, Severac D, Dereeper A (2014) Genome rearrangements derived from homoeologous recombination following allopolyploidy speciation in coffee. Plant J 78(4):674–685
Masterson J (1994) Stomatal size in fossil plants: evidence for polyploidy in majority of angiosperms. Science 264:421–424
Maxwell K, Johnson GN (2000) Chlorophyll fluorescence-a practical guide. J Exp Bot 51:659–668
McClintock B (1984) The significance of responses of the genome to challenge. Science 226:792–801
Miller M, Zhang C, Chen ZJ (2012) Ploidy and hybridity effects on growth vigor and gene expression in Arabidopsis thaliana hybrids and their parents. G3 (Bethesda) 2: 505–513
Murchie E, Lawson T (2013) Chlorophyll fluorescence analysis: a guide to good practice and understanding some new applications. J Exp Bot 64:3983–3998
Ni Z, Kim ED, Ha M, Lackey E, Liu J, Zhang Y, Sun Q, Chen ZJ (2009) Altered circadian rhythms regulate growth vigour in hybrids and allopolyploids. Nature 457:327–331
Osborn TC, Pires JC, Birchler JA, Auger DL, Chen ZJ, Lee HS, Comai L, Madlung A, Doerge RW, Colot V, Martienssen RA (2003) Understanding mechanisms of novel gene expression in polyploids. Trends Genet 19:141–147
Poorter H, Niklas KJ, Reich PB, Oleksyn J, Poot P, Mommer L (2012) Biomass allocation to leaves, stems and roots: meta-analyses of interspecific variation and environmental control. New Phytol 193:30–50
Sarilar V, Palacios PM, Rousselet A, Ridel C, Falque M, Eber F, Chèvre AM, Joets J, Brabant P, Alix K (2013) Allopolyploidy has a moderate impact on restructuring at three contrasting transposable element insertion sites in resynthesized Brassica napus allotetraploids. New Phytol 198:593–604
Van de Peer Y, Mizrachi E, Marchal K (2017) The evolutionary significance of polyploidy. Nat Rev Genet 18:411–424
Villar R, Veneklaas EJ, Jordano P, Lambers H (1998) Relative growth rate and biomass allocation in 20 Aegilops (Poaceae) species. New Phytol 140:425–437
Vogelmann TC (1993) Plant tissue optics. Ann Rew Plant Biol 44:231–251
Wilson JB (1988) A review of evidence on the control of shoot: root ratio, in relation to models. Ann Bot 61:433–449
Yoo MJ, Liu X, Pires JC, Soltis PS, Soltis DE (2014) Nonadditive gene expression in polyploids. Ann Rev Genet 48:485–517
Yu X, Hyldgaard B, Rosenqvist E, Ottosen CO, Chen J (2015) Interspecific hybridization in Cucumis leads to the divergence of phenotypes in response to low light and extended photoperiods. Front Plant Sci 6:802
Yu X, Wang X, Hyldgaard B, Zhu Z, Zhou R, Kjær KH, Ouzounis T, Lou Q, Li J, Cai Q, Rosenqvist E, Ottosen CO, Chen J (2018) Allopolyploidization in Cucumis contributes to delayed leaf maturation with repression of redundant homoeologous genes. Plant J 94:393–404
Zhang K, Wang X, Cheng F (2019) Plant polyploidy: origin, evolution, and its influence on crop domestication. Hort Plant J 5:231–239
Zhou R, Yu X, Kjær KH, Rosenqvist E, Ottosen CO, Wu Z (2015) Screening and validation of tomato genotypes under heat stress using Fv/Fm to reveal the physiological mechanism of heat tolerance. Environ Exp Bot 118:1–11
Zhuang Y, Chen JF (2009) Changes of gene expression in early generations of the synthetic allotetraploid Cucumis × hytivus Chen et Kirkbride. Genet Res Crop Evol 56:1071–1076
Zhuang Y, Chen JF, Jahn M (2009) Expression and sequence variation of the cucumber Por gene in the synthesized allotetraploid Cucumis × hytivus. Mol Biol Rep 36:1725–1731
Acknowledgements
We thank Eva Rosenqvist for the help in conducting the Leaf spectral measurements. This work was supported by the National Natural Science Foundation of China (31902006), the Fundamental Research Funds for the Central Universities (KJQN202029) and the Natural Science Foundation of Jiangsu Province, China (BK20180536).
Author information
Authors and Affiliations
Contributions
XY and JC designed the experiments. XY, YZ and PW performed the experiments and analyzed the data; XY wrote the paper. All authors reviewed and contributed to draft the manuscript. All authors read and approved the final manuscript.
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Additional information
Publisher's note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
About this article
Cite this article
Yu, X., Zhai, Y., Wang, P. et al. Morphological, anatomical and photosynthetic consequences of artificial allopolyploidization in Cucumis. Euphytica 217, 5 (2021). https://doi.org/10.1007/s10681-020-02735-2
Received:
Accepted:
Published:
DOI: https://doi.org/10.1007/s10681-020-02735-2