Abstract
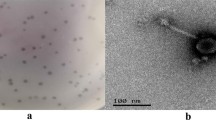
This work describes the characterization and genome annotation of Salmonella phage vB_SalM_8-19 (referred to as 8-19) isolated from sewage samples collected in a pig farm in Jilin, China. This phage was capable of infecting 60% Salmonella strains in our lab stock. The genome of phage 8-19 is composed of linear double-stranded DNA that is 52,648 bp in length with a G + C content of 46.02%; containing 74 ORFs and no tRNA genes. In October 2019, phylogenetic analyses indicated that phage 8-19 might belong to a novel cluster among the other similar phages which have not been specifically classified within some new genus in family Myoviridae. Recently, the International Committee on Taxonomy of Viruses (ICTV) defined phage 8-19 and its related phages as genus Rosemountvirus, family Myoviridae. This new genus, known as Rosemountvirus, is rarely reported in the literature.


Similar content being viewed by others
Code availability
Not applicable.
References
Antunes P, Mourao J, Campos J et al (2016) Salmonellosis: the role of poultry meat. Clin Microbiol Infect Off Publ Eur Soc Clin Microbiol Infect Dis 22:110–121. https://doi.org/10.1016/j.cmi.2015.12.004
Ao TT, Feasey NA, Gordon MA et al (2015) Global burden of invasive nontyphoidal Salmonella disease, 2010. Emerg Infect Dis 21:941–949. https://doi.org/10.3201/eid2106.140999
Arnott A, Wang Q, Bachmann N et al (2018) Multidrug-resistant Salmonella Enterica 4,[5],12:i:-sequence type 34, New South Wales, Australia, 2016–2017. Emerg Infect Dis 24:751–753. https://doi.org/10.3201/eid2404.171619
Bai J, Jeon B, Ryu S (2019) Effective inhibition of Salmonella Typhimurium in fresh produce by a phage cocktail targeting multiple host receptors. Food Microbiol 77:52–60. https://doi.org/10.1016/j.fm.2018.08.011
Bebeacua C, Lorenzo Fajardo JC, Blangy S et al (2013) X-ray structure of a superinfection exclusion lipoprotein from phage TP-J34 and identification of the tape measure protein as its target. Mol Microbiol 89:152–165. https://doi.org/10.1111/mmi.12267
Bolger AM, Lohse M, Usadel B (2014) Trimmomatic: a flexible trimmer for Illumina sequence data. Bioinformatics 30:2114–2120. https://doi.org/10.1093/bioinformatics/btu170
Carvalho C, Costa AR, Silva F et al (2017) Bacteriophages and their derivatives for the treatment and control of food-producing animal infections. Crit Rev Microbiol 43:583–601. https://doi.org/10.1080/1040841X.2016.1271309
Coburn B, Grassl GA, Finlay BB (2007) Salmonella, the host and disease: a brief review. Immunol Cell Biol 85:112–118. https://doi.org/10.1038/sj.icb.7100007
Ekperigin HE, Nagaraja KV (1998) Microbial food borne pathogens. Salmonella. Vet Clin North Am Food Anim Pract 14:17–29. https://doi.org/10.1016/S0749-0720(15)30276-0
Fabrega A, Vila J (2013) Salmonella enterica serovar typhimurium skills to succeed in the host: virulence and regulation. Clin Microbiol Rev 26:308–341. https://doi.org/10.1128/CMR.00066-12
Grant JR, Stothard P (2008) The CGView server: a comparative genomics tool for circular genomes. Nucleic Acids Res 36:W181–W184. https://doi.org/10.1093/nar/gkn179
GRN No. 435 (2013a) Preparation consisting of six bacterial monophage specific to Salmonella enterica (monophage cocktail). https://www.accessdata.fda.gov/scripts/fdcc/index.cfm?set=GRASNotices&id=435&sort=GRN_No&order=DESC&startrow=1&type=basic&search=435. Accessed 22 Feb 2013
GRN No. 468 (2013b) Preparation containing the bacterial monophages, Fo1a and S16, specific to Salmonella. https://www.accessdata.fda.gov/scripts/fdcc/index.cfm?set=GRASNotices&id=468&sort=GRN_No&order=DESC&startrow=1&type=basic&search=468. Accessed 23 Dec 2013
GRN No. 603 (2016a) Preparation containing the bacterial monophages, BP-63 and BP-12, specific to Salmonella. https://www.accessdata.fda.gov/scripts/fdcc/index.cfm?set=GRASNotices&id=603&sort=GRN_No&order=DESC&startrow=1&type=basic&search=603. Accessed 28 Jul 2016
GRN No. 630 (2016b) Preparation containing the bacterial monophages, Fo1a and S16, specific to Salmonella. https://www.accessdata.fda.gov/scripts/fdcc/index.cfm?set=GRASNotices&id=630&sort=GRN_No&order=DESC&startrow=1&type=basic&search=630. Accessed 17 May 2016
GRN No. 752 (2018) Preparation containing the bacterial monophages, BP-63 and LVR16-A, specific to Salmonella. https://www.accessdata.fda.gov/scripts/fdcc/index.cfm?set=GRASNotices&id=752&sort=GRN_No&order=DESC&startrow=1&type=basic&search=752. Accessed 31 Dec 2017
Hagens S, Loessner MJ (2010) Bacteriophage for biocontrol of foodborne pathogens: calculations and considerations. Curr Pharm Biotechnol 11:58–68. https://doi.org/10.2174/138920110790725429
Jajere SM (2019) A review of Salmonella enterica with particular focus on the pathogenicity and virulence factors, host specificity and antimicrobial resistance including multidrug resistance. Vet World 12:504–521. https://doi.org/10.14202/vetworld.2019.504-521
Joensen KG, Scheutz F, Lund O et al (2014) Real-time whole-genome sequencing for routine typing, surveillance, and outbreak detection of verotoxigenic Escherichia coli. J Clin Microbiol 52:1501–1510. https://doi.org/10.1128/JCM.03617-13
Johnson RP, Gyles CL, Huff WE et al (2008) Bacteriophages for prophylaxis and therapy in cattle, poultry and pigs. Anim Health Res Rev 9:201–215. https://doi.org/10.1017/S1466252308001576
Koberg S, Brinks E, Albrecht V et al (2018) Complete genome sequence of the novel virulent phage PMBT28 with lytic activity against thermotolerant Salmonella Enterica subsp. enterica serovar senftenberg ATCC 43845. Genome Announc 6:e00568-e618. https://doi.org/10.1128/genomeA.00568-18
Kropinski AM, Mazzocco A, Waddell TE et al (2009) Enumeration of bacteriophages by double agar overlay plaque assay. Methods Mol Biol 501:69–76. https://doi.org/10.1007/978-1-60327-164-6_7
Kumar S, Stecher G, Li M et al (2018) MEGA X: molecular evolutionary genetics analysis across computing platforms. Mol Biol Evol 35:1547–1549. https://doi.org/10.1093/molbev/msy096
Lang LH (2006) FDA approves use of bacteriophages to be added to meat and poultry products. Gastroenterology 131:1370. https://doi.org/10.1053/j.gastro.2006.10.012
LeLievre V, Besnard A, Schlusselhuber M et al (2019) Phages for biocontrol in foods: what opportunities for Salmonella sp. control along the dairy food chain? Food Microbiol 78:89–98. https://doi.org/10.1016/j.fm.2018.10.009
Lowe TM, Chan PP (2016) tRNAscan-SE on-line: integrating search and context for analysis of transfer RNA genes. Nucleic Acids Res 44:W54–W57. https://doi.org/10.1093/nar/gkw413
Marchler-Bauer A, Bo Y, Han L et al (2017) CDD/SPARCLE: functional classification of proteins via subfamily domain architectures. Nucleic Acids Res 45:D200–D203. https://doi.org/10.1093/nar/gkw1129
Mazzocco A, Waddell TE, Lingohr E et al (2009) Enumeration of bacteriophages using the small drop plaque assay system. Methods Mol Biol 501:81–85. https://doi.org/10.1007/978-1-60327-164-6_9
Milho C, Silva MD, Melo L et al (2018) Control of Salmonella enteritidis on food contact surfaces with bacteriophage PVP-SE2. Biofouling 34:753–768. https://doi.org/10.1080/08927014.2018.1501475
Newase S, Kapadnis BP, Shashidhar R (2019) Isolation and genome sequence characterization of bacteriophage vB_SalM_PM10, a Cba120virus, concurrently infecting Salmonella Enterica serovars typhimurium, typhi, and enteritidis. Curr Microbiol 76:86–94. https://doi.org/10.1007/s00284-018-1588-8
Palomo G, Campos MJ, Ugarte M et al (2013) Dissemination of antimicrobial-resistant clones of Salmonella Enterica among domestic animals, wild animals, and humans. Foodborne Pathog Dis 10:171–176. https://doi.org/10.1089/fpd.2012.1288
Pell LG, Kanelis V, Donaldson LW et al (2009) The phage λ major tail protein structure reveals a common evolution for long-tailed phages and the type VI bacterial secretion system. Proc Natl Acad Sci USA 106:4160–4165. https://doi.org/10.1073/pnas.0900044106
Penades JR, Chen J, Quiles-Puchalt N et al (2015) Bacteriophage-mediated spread of bacterial virulence genes. Curr Opin Microbiol 23:171–178. https://doi.org/10.1016/j.mib.2014.11.019
Saitou N, Nei M (1987) The neighbor-joining method: a new method for reconstructing phylogenetic trees. Mol Biol Evol 4:406–425. https://doi.org/10.1093/oxfordjournals.molbev.a040454
Sambrook J, Russell DW, Nina I et al (2001) Chapter 2: Bacteriophage λ and its vectors. In: Sambrook J, Russell DW (eds) Molecular cloning: A laboratory manual. vol 1. 3rd edn, Cold Spring Harbor, USA, pp 2.56
Scharff RL (2012) Economic burden from health losses due to foodborne illness in the United States. J Food Prot 75:123–131. https://doi.org/10.4315/0362-028X.JFP-11-058
Sritha KS, Bhat SG (2018) Genomics of Salmonella phage ΦStp1: candidate bacteriophage for biocontrol. Virus Genes 54:311–318. https://doi.org/10.1007/s11262-018-1538-3
Voetsch AC, Van Gilder TJ, Angulo FJ et al (2004) FoodNet estimate of the burden of illness caused by nontyphoidal Salmonella infections in the United States. Clin Infect Dis 38:S127–S134. https://doi.org/10.1086/381578
Wang IN, Smith DL, Young R (2000) Holins: the protein clocks of bacteriophage infections. Annu Rev Microbiol 54:799–825. https://doi.org/10.1146/annurev.micro.54.1.799
Yan T, Liang L, Yin P et al (2020) Application of a novel phage LPSEYT for biological control of Salmonella in foods. Microorganisms 8:400. https://doi.org/10.3390/microorganisms8030400
Yuan YY, Zhao FY, Wang LL et al (2019) Complete genome analysis of the novel Enterococcus faecalis phage vB_EfaS_AL3. Arch Virol 164:2599–2603. https://doi.org/10.1007/s00705-019-04341-7
Zankari E, Hasman H, Cosentino S et al (2012) Identification of acquired antimicrobial resistance genes. J Antimicrob Chemother 67:2640–2644. https://doi.org/10.1093/jac/dks261
Zinno P, Devirgiliis C, Ercolini D et al (2014) Bacteriophage P22 to challenge Salmonella in foods. Int J Food Microbiol 191:69–74. https://doi.org/10.1016/j.ijfoodmicro.2014.08.037
Funding
This work was supported by the LiaoNing Revitalization Talents Program of China (Grant number XLYC1907085), the Fundamental Research Funds for the Central Universities of China (Grant number DUT20GJ217), and the Dalian Key Technology Research and Development Program of China (Grant number 2019YF23SN048).
Author information
Authors and Affiliations
Contributions
CC: writing-original draft, visualization, investigation, data curation, software. WBD: resources. CHJ: Writing-review and editing. YYY: investigation. XYP: project administration, funding acquisition, resources. WLL: formal analysis. LSY: funding acquisition. LJB: methodology. XM: project administration. LXY: conceptualization, funding acquisition.
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Ethical approval
This article does not contain any studies with human participants or animals by any of the authors. In this study, the sewage samples were kindly provided and collected from a pig farm of the Jilin Academy of Agricultural Sciences (JAAS, Jilin, China). We are grateful to JAAS for total approvals and permissions to collect the sewage samples.
Consent to participate
Not applicable.
Consent for publication
All authors gave approval for publication.
Availability of data and material
All data needed to evaluate the conclusions in the paper are present in the paper and/or the Supplementary Materials.
Additional information
Communicated by Erko Stackebrandt.
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
About this article
Cite this article
Cong, C., Bing Dong, W., Hui Jing, C. et al. Genome analysis of Salmonella phage vB_SalM_8-19 (genus Rosemountvirus). Arch Microbiol 203, 1345–1356 (2021). https://doi.org/10.1007/s00203-020-02121-5
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00203-020-02121-5