Abstract
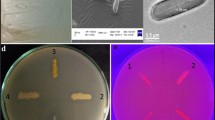
Lipase is an important commercial enzyme with unique and versatile biotechnological applications. This study was conducted to biosynthesize and characterizes alkaliphilic lipase by Exiguobacterium sp. strain AMBL-20T isolated from the glacial water samples of the northeastern (Gilgit-Baltistan) region of Pakistan. The isolated bacterium was identified as Exiguobaterium sp. strain AMBL-20T on the basis of morphological, biochemical, and phylogenetic analysis of 16S rRNA sequences with GenBank accession number MW229267. The bacterial strain was further screened for its lipolytic activity, biosynthesis, and characterization by different parameters with the aim of maximizing lipase activity. Results showed that 2% Olive oil, 0.2% peptone at 25 °C, pH 8, and 24 h of incubation time found optimal for maximum lipase production. The lipase enzyme was partially purified by ammonium sulphate precipitation and its activity was standardized at pH 8 under 30 °C temperature. The enzyme showed functional stability over a range of temperature and pH. Hence, extracellular alkaliphilic lipase from Exiguobacterium sp. is a potential candidate with extraordinary industrial applications, particularly in bio-detergent formulations.





Similar content being viewed by others
References
Abdel-Fattah YR, Soliman NA, Yousef SM, El-Helow ER (2012) Application of experimental designs to optimize medium composition for the production of thermostable lipase/esterase by Geobacillus thermodenitrificans AZ1. J Gen Eng Biotech 10(2):193–200. https://doi.org/10.1016/j.jgeb.2012.08.001
Adetunji AI, Olaniran AO (2018) Optimization of culture conditions for enhanced lipase production by an indigenous Bacillus aryabhattai SE3-PB using response surface methodology. Biotech Biotech Equip 32(6):1514–1526. https://doi.org/10.1080/13102818.2018.1514985
Alam MZ, Malik A (2008) Chromate resistance, transport and bioreduction by Exiguobacterium sp. ZM-2 isolated from agricultural soil irrigated with tannery effluent. J Basic Microbiol 48:416–420. https://doi.org/10.1002/jobm.200800046
Ali CH, Zhang JJ, Mbadinga SM, Liu JF, Yang SZ, Gu JD, Mu BZ (2015) Screening, isolation, and optimization of an extracellular lipase producing Exiguobacterium sp. BBXS-7 segregated from waste cooking oil-contaminated sites. Wulfen J 22:185–201
Amin A (2019) Review of diesel production from renewable resources: catalysis, process kinetics, and technologies. Ain Shams Eng J 10(4):821–839. https://doi.org/10.1016/j.asej.2019.08.001
Anobom CD, Pinheiro AS, De-Andrade RA, Aguieiras EC, Andrade GC, Moura MV, Freire DM (2014) From structure to catalysis: recent developments in the biotechnological applications of lipases. BioMed Res Int. https://doi.org/10.1155/2014/684506
Bacha AB, Al-Assaf A, Moubayed NM, Abid I (2018) Evaluation of a novel thermo-alkaline Staphylococcus aureus lipase for application in detergent formulations. Saud J Biol Sci 25(3):409–417. https://doi.org/10.1016/j.sjbs.2016.10.006
Bhosale HJ, Sukalkar SR, Shaheen ZU, Kadam TA (2011) Production of xylanase by Streptomyces rameus grown on agricultural wastes. J Biotech Bioinf Bioeng 1:505–512
Bhushan I, Yadav A, Parshad R (2011) Enhancement in the production of lipase from Arthrobacter sp. using fed-batch fermentation strategy. Biosci Biotech Res Asia 2(5):522–534. https://dioi.org/30-06/AJOBR-2011/2(5)522-534
Bora L, Bora M (2012) Optimization of extracellular thermophilic highly alkaline lipase from thermophilic Bacillus sp isolated from Hotspring of Arunachal Pradesh. India Braz J Microbiol 43(1):30–42
Bordes F, Barbe S, Escalier P, Mourey L, André I, Marty A, Tranier S (2010) Exploring the conformational states and rearrangements of Yarrowia lipolytica lipase. Biophys J 99(7):2225–2234
Borkar PS, Bodade RG, Rao SR, Khobragade CN (2009) Purification and characterization of extracellular lipase from a new strain: Pseudomonas aeruginosa SRT 9. Braz J Microbiol 40(2):358–366. https://doi.org/10.1590/S1517-83822009000200028
Bornscheuer UT, Bessler C, Srinivas R, Krishna SH (2002) Optimizing lipases and related enzymes for efficient application. TRENDS Biotechnol 20(10):433–437. https://doi.org/10.1016/S0167-7799(02)02046-2
Casas-Godoy L, Duquesne S, Bordes F, Sandoval G, Marty A (2012) Lipases: an overview. In: Lipases and phospholipases. Humana Press, p 3–30 https://doi.org/https://doi.org/10.1007/978-1-4939-8672-9_1,
Cavicchioli R (2016) On the concept of a psychrophile. ISME J 10(4):793. https://doi.org/10.1038/ismej.2015.160
Chang J, Lee YS, Fang SJ, Park IH, Choi YL (2013) Recombinant expression and characterization of an organic-solvent-tolerant a-amylase from Exiguobacterium sp. DAU5. Biotech Appl Biochem 169:1870–1883
Chaturvedi P, Shivaji S (2006) Exiguobacterium indicum sp. nov., a psychrophilic bacterium from the Hamta glacier of the Himalayan mountain ranges of India. Int J Syst Evol Microbiol 56:2765–2770. https://doi.org/10.1099/ijs.0.64508-0
Chen X, Wang L, Zhou J, Wu H, Li D, Cui Y, Lu B (2017) Exiguobacterium sp. A1b/GX59 isolated from a patient with community-acquired pneumonia and bacteremia: genomic characterization and literature review. BMC Infec Dis 17(1):508
Cherif S, Mnif S, Hadrich F et al (2011) A newly high alkaline lipase: an ideal choice for application in detergent formulations. Lip Health Dis 10:221. https://doi.org/10.1186/1476-511X-10-221
Chu IM, Lee C, Li TS (1992) Production and degradation of alkaline protease in batch cultures of Bacillus subtilis ATCC 14416. Enz Microb Technol 14(9):755–761
Crapart S, Fardeau ML, Cayol JL et al (2007) Exiguobacterium profundum sp. nov., a moderately thermophilic, lactic acid-producing bacterium isolated from a deep-sea hydrothermal vent. Int J Syst Evol Microbiol 57:287–292. https://doi.org/10.1099/ijs.0.64639-0
Dastager SG, Mawlankar R, Sonalkar VV et al (2015) Exiguobacterium enclense sp. nov., isolated from sediment. Int J Syst Evol Microbiol 65:1611–1616. https://doi.org/10.1099/ijs.0.000149
de Oliveira LA, Germano MG, Willerding AL, Moreira FW, Chagas AF Jr (2011) Lipase activity among bacteria isolated from Amazonian soils. Enzyme Res. https://doi.org/10.4061/2011/720194
Dhanve RS, Kalyani DC, Phugare SS et al (2009) Coordinate action of exiguobacterial oxidoreductive enzymes in biodegradation of reactive yellow 84A dye. Biodegradation 20:245–255. https://doi.org/10.1007/s10532-008-9217-z
Duza MB, Mastan S (2014) Optimization of lipase production from Bacillus thuringiensis (TS11BP), Achromobacter xylosoxidans J2 (TS2MCN)-isolated from soil sediments near oilseed farm. IOSR J Pharm Biol Sci 9:66–76
Eko Sukohidayat N, Zarei M, Baharin B, Manap M (2018) Purification and characterization of lipase produced by Leuconostoc mesenteroides subsp. mesenteroides ATCC 8293 using an aqueous two-phase system (ATPS) composed of triton X-100 and maltitol. Molecules 23(7):1800. https://doi.org/10.3390/molecules23071800
Fatima H, Khan N, Rehman AU, Hussain Z (2014) Production and partial characterization of lipase from Pseudomonas putida. J Ferment Technol 2:112. https://doi.org/10.4172/2167-7972.1000112
Ferreira-Leitão VS, Cammarota MC, Goncalves Aguieiras EC, de Sá V, Ribeiro L, Fernandez-Lafuente R, Freire DMG (2017) The protagonism of biocatalysis in green chemistry and its environmental benefits. Catalysts 7(1):9. https://doi.org/10.3390/catal7010009
Frühling A, Schumann P, Hippe H, Sträubler B, Stackebrandt E (2002) Exiguobacterium undae sp. nov. and Exiguobacterium antarcticum sp. nov. Int J Syst Evol Microbiol 52(4):1171–1176
Gupta A, Joseph B, Mani A, Thomas G (2008) Biosynthesis and properties of an extracellular thermostable serine alkaline protease from Virgibacillus pantothenticus. World J Microbiol Biotechnol 24(2):237–243. https://doi.org/10.1007/s11274-007-9462-z
Hakim A, Bhuiyan FR, Iqbal A, Emon TH, Ahmed J, Azad AK (2018) Production and partial characterization of dehairing alkaline protease from Bacillus subtilis AKAL7 and Exiguobacterium indicum AKAL11 by using organic municipal solid wastes. Heliyon 4(6):e00646. https://doi.org/10.1016/j.heliyon.2018.e00646
Hasan F, Shah AA, Hameed A (2006) Industrial applications of microbial lipases. Enz Microb Technol 39(2):235–251. https://doi.org/10.1016/j.enzmictec.2005.10.016
Hassan SW, Ali M, Elsayed SM (2018) Production of cold active lipase by free and immobilized marine Bacillus cereus HSS: application in wastewater treatment. Front Microbiol 9:2377. https://doi.org/10.3389/fmicb.2018.02377
Hemlata B, Uzma Z, Tukaram K (2016) Substrate kinetics of thiol activated hyperthermostable alkaline lipase of Bacillus sonorensis 4R and its application in bio-detergent formulation. Biocat Agric Biotechnol 8:104–111. https://doi.org/10.1016/j.bcab.2016.08.0081
Ito S, Kobayashi T, Ara K, Ozaki K, Kawai S, Hatada Y (1998) Alkaline detergent enzymes from alkaliphiles: enzymatic properties, genetics and structures. Extremophiles 2:185–190
Jaeger KE, Eggert T (2002) Lipases for biotechnology. Curr Opin Biotechnol 13(4):390–397. https://doi.org/10.1016/S0958-1669(02)00341-5
Jaeger KE, Reetz MT (1998) Microbial lipases form versatile tools for biotechnology. Trend Biotechnol 16(9):396–403
Jaiswal A, Preet M, Tripti B (2017) Production and optimization of lipase enzyme from mesophiles and thermophiles. J Microb Biochem Technol 9(3):126–131. https://doi.org/10.4172/1948-5948.1000355
Joseph B, Shrivastava N, Ramteke PW (2012) Extracellular cold-active lipase of Microbacterium luteolum isolated from Gangotri glacier, western Himalaya: Isolation, partial purification and characterization. J Gen Engin Biotechnol 10(1):137–144. https://doi.org/10.1016/j.jgeb.2012.02.001
Kasana RC, Pandey CB (2018) Exiguobacterium: an overview of a versatile genus with potential in industry and agriculture. Crit Rev Biotechnol 38(1):141–156. https://doi.org/10.1080/07388551.2017.1312273
Kasana RC, Yadav SK (2007) Isolation of a psychrotrophic Exiguobacterium sp. SKPB5 (MTCC 7803) and characterization of its alkaline protease. Curr Microbiol 54:224–229. https://doi.org/10.1007/s00284-006-0402-1
Katiyar P, Pratibha VH, Baghel VS (2017) Isolation, partial purification and characterization of a cold active lipase from Pseudomonas sp., isolated from satopanth glacier of western Himalaya India. Int J Scient Res Manag 5(7):6106–6112. https://doi.org/10.18535/ijsrm/v5i7.3
Khan FI, Lan D, Durrani R, Huan W, Zhao Z, Wang Y (2017) The lid domain in lipases: structural and functional determinant of enzymatic properties. Front Bioengin Biotechnol 5:16
Kumar PS, Raj JPP, Duraipandiyan V, Ignacimuthu S (2012) Antibacterial activity of some actinomycetes from Tamil Nadu. India Asian Pac J Trop Biomed 2(12):936–943. https://doi.org/10.1016/S2221-1691(13)60003-9
Kumar PP, Jansi RS, Kumar PS, Christhudas IN, Raj JP, Vijayakumar A, Ignacimuthu S (2017) Optimization of biosynthesis parameters, partial purification and characterization of extracellular lipase from soil derived Streptomyces sp. Loyola Lipase-1. Biocat Agricu Biotechnol 12:241–247. https://doi.org/10.1016/j.bcab.2017.10.011
Lakshmi P (2012) Purification and characterization of an alkaline protease from a novel soil isolate, Exiguobacterium acetylicum MTCC 9115 [dissertation]. India: Institute of Science and Technology Jawaharlal Nehru Technological University, Hyderabad; . https://doi.org/https://doi.org/10.1080/07388551.2017.1312273
Lanyı B (1987) Classical and rapid identification methods for medically important bacteria. Meth Microbiol 19:1–67
Lee SH, Chung CW, Yu YJ et al (2009) Effect of alkaline protease- producing Exiguobacterium sp. YS1 inoculation on the solubilization and bacterial community of waste activated sludge. Biores Technol 100:4597–4603. https://doi.org/10.1016/j.biortech.2009.04.056
Lee LP, Karbul HM, Citartan M, Gopinath SC, Lakshmipriya T, Tang TH (2015) Lipase secreting Bacillus species in an oil-contaminated habitat: promising strains to alleviate oil pollution. Biomed Res Int 1:1–9. https://doi.org/10.1155/2015/820575
Marmur J (1961) A procedure for the isolation of deoxyribonucleic acid from micro-organisms. J Mol Biol 3(2):208-IN1. https://doi.org/10.1016/S0022-2836(61)80047-8
Mazhar H, Abbas N, Zamir T, Hussain Z, Ali SS (2018) Optimization study of lipolytic enzyme from Bacillus cereus, PCSIR NL-37. Punjab Uni J Zool 33(2):217–224. https://doi.org/10.17582/journal.pujz/2018.33.2.217.224
Mohanasrinivasan V, Devi CS, Jayasmita D, Selvarajan E, Naine SJ (2018) Purification and characterization of extracellular lipase from Serratia marcescens VITSD2. Proc Nat Acad Sci India Sect B: Biol Sci 88(1):373–381. https://doi.org/10.1007/s40011-016-0763-6
Mojallali L, Shahbani Zahiri H, Rajaei S, Akbari Noghabi K, Haghbeen K (2014) A novel _34-kDa a-amylase from psychrotroph Exiguobacterium sp. SH3: production, purification, and characterization. Biotechnol Appl Biochem 61:118–125. https://doi.org/10.1002/bab.1140
Nardini M, Dijkstra BW (1999) α/β Hydrolase fold enzymes: the family keeps growing. Curr Opin Struct Biol 9(6):732–737
Noble MEM, Cleasby A, Johnson LN, Egmond MR, Frenken LGJ (1993) The crystal structure of triacylglycerol lipase from Pseudomonas glumae reveals a partially redundant catalytic aspartate. FEBS Lett 331(1–2):123–128
Nwachukwu E, Ejike EN, Ejike BU, Onyeanula EO, Chikezie-Abba RO, Okorocha NA, Onukaogu UE (2017) Characterization and optimization of lipase production from soil microorganism (Serratia marcescens). Int J Curr Microbiol Appl Sci 6(12):1215–1231. https://doi.org/10.20546/ijcmas.2017.612.138
Olajuyigbe FM, Ajele JO (2011) Thermostable alkaline protease from Bacillus licheniformis lbbl-11 isolated from traditionally fermented African locust bean (Parkia biglobosa). J Food Biochem 35(1):1–10. https://doi.org/10.1111/j.1745-4514.2010.00362.x
Ortiz C, Ferreira ML, Barbosa O, dos Santos JC, Rodrigues RC, Berenguer-Murcia Á, Fernandez-Lafuente R (2019) Novozym 435: the “perfect” lipase immobilized biocatalyst? Catal Sci Technol 10:2380–2420. https://doi.org/10.1039/C9CY00415G
Pallavi P, Chandra SJ, Reddy VK, Reddy SR (2014) Optimization of cultural parameters for lipase production by Bacillus subtilis Y-IVI. Int J Curr Microbiol Appl Sci 3(12):194–200
Peters GH, van Aalten DM, Svendsen A, Bywater R (1997) Essential dynamics of lipase binding sites: the effect of inhibitors of different chain length. Protein Engin 10(2):149–158
Qiao Y, Peng Q, Yan J (2015) Gene cloning and enzymatic characterization of alkali-tolerant type I pullulanase from Exiguobacterium acetylicum. Lett Appl Microbiol 60:52–59. https://doi.org/10.1111/lam.12333
Rahman RNZRA, Masomian M, Salleh AB, Basri M (2009) A new thermostable lipase by Aneurinibacillus thermoaerophilus strain HZ: nutritional studies. Ann Microbiol 59(1):133
Rajaei S, Heidari R, Zahiri HS (2014) A novel coldadapted pullulanase from Exiguobacterium sp. SH3: production optimization, purification, and characterization. Starch 66:225–234. https://doi.org/10.1002/star.201300030
Ranaldi S, Belle V, Woudstra M, Bourgeas R, Guigliarelli B, Roche P, Fournel A (2010) Amplitude of pancreatic lipase lid opening in solution and identification of spin label conformational subensembles by combining continuous wave and pulsed EPR spectroscopy and molecular dynamics. Biochemistry 49(10):2140–2149
Razzaq A, Shamsi S, Ali A, Ali Q, Sajjad M, Malik A, Ashraf M (2019) Microbial proteases applications. Biotech 7:110. https://doi.org/10.3389/fbioe.2019.00110
Reiner K (2010) Catalase test protocol. American Society for Microbiology. Available at: http://www.microbelibrary.org/library/laboratory-test/3226-catalase-testprotocol.htm
Robinson PK (2015) Enzymes: principles and biotechnological applications. Ess Biochem 59:1–41. https://doi.org/10.1042/bse0590001
Rodrigues DF, Goris J, Vishnivetskaya T (2006) Characterization of Exiguobacterium isolates from the Siberian permafrost, description of Exiguobacterium sibiricum sp. nov. Extremophiles 10:285–294. https://doi.org/10.1007/s00792-005-0497-5
Sachan S, Iqbal MS, Singh A (2018) Extracellular lipase from Pseudomonas aeruginosa JCM5962 (T): isolation, identification, and characterization. Int Microbiol 21(4):197–205. https://doi.org/10.1007/s10123-018-0016-z
Saraswat R, Verma V, Sistla S, Bhushan I (2017) Evaluation of alkali and thermotolerant lipase from an indigenous isolated Bacillus strain for detergent formulation. Elect J Biotechnol 30:33–38. https://doi.org/10.1016/j.ejbt.2017.08.007
Shields P (2010) Cathcart L. Oxidase test protocol. American Society for Microbiology. Available at: http://www.microbelibrary.org/library/laboratory-test/3229-oxidase-test-protocol.htm
Singh NK, Raichand R, Kaur I (2013) Exiguobacterium himgiriensis sp. nov. a novel member of the genus Exiguobacterium, isolated from the Indian Himalayas. Antonie Van Leeuwenhoek 103:789–796. https://doi.org/10.1007/s10482-012-9861-5
Smibert RM, Krieg NR (1994) Phenotypic characterization. In: Gerhardt P (ed) methods for general and molecular bacteriology. American Society for Microbiology, Washington, DC, pp 607–654
Soleymani S, Alizadeh H, Mohammadian H, Rabbani E, Moazen F, Sadeghi HM, Rabbani M (2017) Efficient media for high lipase production: one variable at a time approach. Avic J Med Biotechnol 9(2):82
Sooch BS, Kauldhar BS (2013) Influence of multiple bioprocess parameters on production of lipase from Pseudomonas sp. BWS-5. Braz Arch Biol Technol 56(5):711–721. https://doi.org/10.1590/S1516-89132013000500002
Sumarsih S, Hadi S, Andini DGT, Nafsihana FK (2018) Carbon and nitrogen sources for lipase production of Micrococcus sp. Isolated from palm oil mill effluent-contaminated soil. IOP Conf Ser: Earth and Env Sci 217:1–012029. https://doi.org/10.1088/1755-1315/217/1/012029
Tamura K, Stecher G, Peterson D, Filipski A, Kumar S (2013) MEGA6: molecular evolutionary genetics analysis version 6.0. Mol Biol Evol 30(12):2725–2729. https://doi.org/10.1093/molbev/mst197
Thakur V, Tewari R, Sharma R (2014) Evaluation of production parameters for maximum lipase production by P. stutzeri MTCC 5618 and scale-up in bioreactor. Chin J Biol 2014:14. https://doi.org/10.1155/2014/208462
Tripathi R, Singh J, Kumar Bharti R, Thakur IS (2014) Isolation, purification and characterization of lipase from Microbacterium sp. and its application in biodiesel production. Energy Procedia 54:518–529. https://doi.org/10.1016/j.egypro.2014.07.293
Ugo AK, Amara AV, Igwe CN, Kenechuwku U (2017) Microbial lipases: a prospect for biotechnological industrial catalysis for green products: a review. J Ferment Tech 6(144):2
Vázquez L, González N, Reglero G, Torres C (2016) Solvent-free lipase-catalyzed synthesis of diacylgycerols as low-calorie food ingredients. Front Bioengin Biotechnol 4:6. https://doi.org/10.3389/fbioe.2016.00006
Veerapagu M, Narayanan AS, Ponmurugan K, Jeya K (2013) Screening selection identification production and optimization of bacterial lipase from oil spilled soil. Asian J Pharm Clin Res 6(3):62–67
Vishnivetskaya TA, Kathariou S, Tiedje JM (2009) The Exiguobacterium genus: biodiversity and biogeography. Extremophiles 13:541–555. https://doi.org/10.1007/s00792-009-0243-5
Vitorino LC, Bessa LA (2017) Technological microbiology: development and applications. Front Microbiol 8:827. https://doi.org/10.3389/fmicb.2017.00827
Weisburg WG, Barns SM, Pelletier DA, Lane DJ (1991) 16S ribosomal DNA amplification for phylogenetic study. J Bacteriol 173:697–703
Zarinviarsagh M, Ebrahimipour G, Sadeghi H (2017) Lipase and biosurfactant from Ochrobactrum intermedium strain MZV101 isolated by washing powder for detergent application. Lip Health Dis 16(1):177. https://doi.org/10.1186/s12944-017-0565-8
Acknowledgements
The authors are thankful to the Department of Biological Sciences, International Islamic University Islamabad (IIUI), and HEC Pakistan for the provision of essential research services to complete this research work.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Additional information
Communicated by Erko Stackebrandt.
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
About this article
Cite this article
Yasin, M.T., Ali, Y., Ahmad, K. et al. Alkaline lipase production by novel meso-tolerant psychrophilic Exiguobacterium sp. strain (AMBL-20) isolated from glacier of northeastern Pakistan. Arch Microbiol 203, 1309–1320 (2021). https://doi.org/10.1007/s00203-020-02133-1
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00203-020-02133-1