Abstract
Filamentous actin (F-actin) provides cells with mechanical support and promotes the mobility of intracellular structures. Although F-actin is traditionally considered to be cytoplasmic, here we reveal that nuclear F-actin participates in the replication stress response. Using live and super-resolution imaging, we find that nuclear F-actin is polymerized in response to replication stress through a pathway regulated by ATR-dependent activation of mTORC1, and nucleation through IQGAP1, WASP and ARP2/3. During replication stress, nuclear F-actin increases the nuclear volume and sphericity to counteract nuclear deformation. Furthermore, F-actin and myosin II promote the mobility of stressed-replication foci to the nuclear periphery through increasingly diffusive motion and directed movements along the nuclear actin filaments. These actin functions promote replication stress repair and suppress chromosome and mitotic abnormalities. Moreover, we find that nuclear F-actin is polymerized in vivo in xenograft tumours after treatment with replication-stress-inducing chemotherapeutic agents, indicating that this pathway has a role in human disease.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout






Similar content being viewed by others
Data availability
All data supporting the findings of this study are available from the corresponding author on reasonable request. Source data are provided with this paper.
Code availability
Custom MATLAB code for performing nuclear registration with Imaris XT (Bitplane) is available at http://www.matebiro.com/software/nuclearactin. Custom Python code for generating the sliding window MSD analysis is available at GitHub (https://github.com/marknormanread/motility_analysis_lamm2020). Both codes are released under the GNU general public license (v.3).
References
Zeman, M. K. & Cimprich, K. A. Causes and consequences of replication stress. Nat. Cell Biol. 16, 2–9 (2014).
Saldivar, J. C., Cortez, D. & Cimprich, K. A. The essential kinase ATR: ensuring faithful duplication of a challenging genome. Nat. Rev. Mol. Cell Biol. 18, 622–636 (2017).
Cortez, D. Replication-coupled DNA repair. Mol. Cell 74, 866–876 (2019).
Lopes, M. et al. The DNA replication checkpoint response stabilizes stalled replication forks. Nature 412, 557–561 (2001).
Kumar, A. et al. ATR mediates a checkpoint at the nuclear envelope in response to mechanical stress. Cell 158, 633–646 (2014).
Svitkina, T. The actin cytoskeleton and actin-based motility. Cold Spring Harb. Perspect. Biol. 10, a018267 (2018).
Moore, H. M. & Vartiainen, M. K. F-actin organizes the nucleus. Nat. Cell Biol. 19, 1386–1388 (2017).
Baarlink, C., Wang, H. & Grosse, R. Nuclear actin network assembly by formins regulates the SRF coactivator MAL. Science 340, 864–867 (2013).
Plessner, M., Melak, M., Chinchilla, P., Baarlink, C. & Grosse, R. Nuclear F-actin formation and reorganization upon cell spreading. J. Biol. Chem. 290, 11209–11216 (2015).
Baarlink, C. et al. A transient pool of nuclear F-actin at mitotic exit controls chromatin organization. Nat. Cell Biol. 19, 1389–1399 (2017).
Belin, B. J., Lee, T. & Mullins, R. D. DNA damage induces nuclear actin filament assembly by formin-2 and Spire-(1/2) that promotes efficient DNA repair. eLife 4, e07735 (2015).
Caridi, C. P. et al. Nuclear F-actin and myosins drive relocalization of heterochromatic breaks. Nature 559, 54–60 (2018).
Schrank, B. R. et al. Nuclear ARP2/3 drives DNA break clustering for homology-directed repair. Nature 559, 61–66 (2018).
Imseng, S., Aylett, C. H. & Maier, T. Architecture and activation of phosphatidylinositol 3-kinase related kinases. Curr. Opin. Struct. Biol. 49, 177–189 (2018).
Saxton, R. A. & Sabatini, D. M. mTOR signaling in growth, metabolism, and disease. Cell 169, 361–371 (2017).
Mok, K. W., Chen, H., Lee, W. M. & Cheng, C. Y. rpS6 regulates blood-testis barrier dynamics through Arp3-mediated actin microfilament organization in rat sertoli cells. An in vitro study. Endocrinology 156, 1900–1913 (2015).
Mok, K. W., Mruk, D. D. & Cheng, C. Y. rpS6 regulates blood-testis barrier dynamics through Akt-mediated effects on MMP-9. J. Cell Sci. 127, 4870–4882 (2014).
Padrick, S. B., Doolittle, L. K., Brautigam, C. A., King, D. S. & Rosen, M. K. Arp2/3 complex is bound and activated by two WASP proteins. Proc. Natl Acad. Sci. USA 108, E472–E479 (2011).
Parisis, N. et al. Initiation of DNA replication requires actin dynamics and formin activity. EMBO J. 36, 3212–3231 (2017).
Lamm, N., Rogers, S. & Cesare, A. J. The mTOR pathway: implications for DNA replication. Prog. Biophys. Mol. Biol. 147, 17–25 (2019).
Essers, J. et al. Nuclear dynamics of PCNA in DNA replication and repair. Mol. Cell. Biol. 25, 9350–9359 (2005).
Petermann, E., Orta, M. L., Issaeva, N., Schultz, N. & Helleday, T. Hydroxyurea-stalled replication forks become progressively inactivated and require two different RAD51-mediated pathways for restart and repair. Mol. Cell 37, 492–502 (2010).
Schonbrun, M. et al. TOR complex 2 controls gene silencing, telomere length maintenance, and survival under DNA-damaging conditions. Mol. Cell. Biol. 29, 4584–4594 (2009).
Shimada, K. et al. TORC2 signaling pathway guarantees genome stability in the face of DNA strand breaks. Mol. Cell 51, 829–839 (2013).
Matsuoka, S. et al. ATM and ATR substrate analysis reveals extensive protein networks responsive to DNA damage. Science 316, 1160–1166 (2007).
Shen, C. et al. TOR signaling is a determinant of cell survival in response to DNA damage. Mol. Cell. Biol. 27, 7007–7017 (2007).
Kim, S. et al. Amino acid signaling to mTOR mediated by inositol polyphosphate multikinase. Cell Metab. 13, 215–221 (2011).
Wang, Y. H. et al. DNA damage causes rapid accumulation of phosphoinositides for ATR signaling. Nat. Commun. 8, 2118 (2017).
Maiso, P. et al. Defining the role of TORC1/2 in multiple myeloma. Blood 118, 6860–6870 (2011).
Latres, E. et al. Insulin-like growth factor-1 (IGF-1) inversely regulates atrophy-induced genes via the phosphatidylinositol 3-kinase/Akt/mammalian target of rapamycin (PI3K/Akt/mTOR) pathway. J. Biol. Chem. 280, 2737–2744 (2005).
Tekletsadik, Y. K., Sonn, R. & Osman, M. A. A conserved role of IQGAP1 in regulating TOR complex 1. J. Cell Sci. 125, 2041–2052 (2012).
Bashour, A. M., Fullerton, A. T., Hart, M. J. & Bloom, G. S. IQGAP1, a Rac- and Cdc42-binding protein, directly binds and cross-links microfilaments. The. J. Cell Biol. 137, 1555–1566 (1997).
Le Clainche, C. et al. IQGAP1 stimulates actin assembly through the N-WASP-Arp2/3 pathway. J. Biol. Chem. 282, 426–435 (2007).
Johnson, M. A., Sharma, M., Mok, M. T. & Henderson, B. R. Stimulation of in vivo nuclear transport dynamics of actin and its co-factors IQGAP1 and Rac1 in response to DNA replication stress. Biochim. Biophys. acta 1833, 2334–2347 (2013).
Arber, S. et al. Regulation of actin dynamics through phosphorylation of cofilin by LIM-kinase. Nature 393, 805–809 (1998).
Dopie, J., Skarp, K. P., Rajakyla, E. K., Tanhuanpaa, K. & Vartiainen, M. K. Active maintenance of nuclear actin by importin 9 supports transcription. Proc. Natl Acad. Sci. USA 109, E544–E552 (2012).
Lossaint, G. et al. FANCD2 binds MCM proteins and controls replisome function upon activation of S phase checkpoint signaling. Mol. Cell 51, 678–690 (2013).
Sfeir, A. et al. Mammalian telomeres resemble fragile sites and require TRF1 for efficient replication. Cell 138, 90–103 (2009).
Titus, M. A. Myosin-driven intracellular transport. Cold Spring Harb. Perspect. Biol. 10, a021972 (2018).
de Lanerolle, P. & Serebryannyy, L. Nuclear actin and myosins: life without filaments. Nat. Cell Biol. 13, 1282–1288 (2011).
Higuchi, H. & Takemori, S. Butanedione monoxime suppresses contraction and ATPase activity of rabbit skeletal muscle. J. Biochem. 105, 638–643 (1989).
Straight, A. F. et al. Dissecting temporal and spatial control of cytokinesis with a myosin II inhibitor. Science 299, 1743–1747 (2003).
Burrell, R. A. et al. Replication stress links structural and numerical cancer chromosomal instability. Nature 494, 492–496 (2013).
Kedrin, D. et al. Intravital imaging of metastatic behavior through a mammary imaging window. Nat. Methods 5, 1019–1021 (2008).
Nobis, M. et al. A RhoA-FRET biosensor mouse for intravital imaging in normal tissue homeostasis and disease contexts. Cell Rep. 21, 274–288 (2017).
Madaan, K., Kaushik, D. & Verma, T. Hydroxyurea: a key player in cancer chemotherapy. Expert Rev. Anticancer Ther. 12, 19–29 (2012).
Wagner, J. M. & Karnitz, L. M. Cisplatin-induced DNA damage activates replication checkpoint signaling components that differentially affect tumor cell survival. Mol. Pharmacol. 76, 208–214 (2009).
Masamsetti, V. P. et al. Replication stress induces mitotic death through parallel pathways regulated by WAPL and telomere deprotection. Nat. Commun. 10, 4224 (2019).
Singh, M. et al. Lamin A/C depletion enhances DNA damage-induced stalled replication fork arrest. Mol. Cell. Biol. 33, 1210–1222 (2013).
Guerreiro, I. & Kind, J. Spatial chromatin organization and gene regulation at the nuclear lamina. Curr. Opin. Genet. Dev. 55, 19–25 (2019).
Seeber, A., Hauer, M. H. & Gasser, S. M. Chromosome dynamics in response to DNA damage. Annu. Rev. Genet. 52, 295–319 (2018).
Chung, D. K. et al. Perinuclear tethers license telomeric DSBs for a broad kinesin- and NPC-dependent DNA repair process. Nat. Commun. 6, 7742 (2015).
Freudenreich, C. H. & Su, X. A. Relocalization of DNA lesions to the nuclear pore complex. FEMS Yeast Res. 16, fow095 (2016).
Jacinto, E. et al. Mammalian TOR complex 2 controls the actin cytoskeleton and is rapamycin insensitive. Nat. Cell Biol. 6, 1122–1128 (2004).
Sarbassov, D. D. et al. Rictor, a novel binding partner of mTOR, defines a rapamycin-insensitive and raptor-independent pathway that regulates the cytoskeleton. Curr. Biol. 14, 1296–1302 (2004).
Cohen, A., Kupiec, M. & Weisman, R. Gad8 protein is found in the nucleus where it Interacts with the Mlui cell cycle box-binding factor (MBF) transcriptional complex to regulate the response to DNA replication stress. J. Biol. Chem. 291, 9371–9381 (2016).
Swiech, L. et al. CLIP-170 and IQGAP1 cooperatively regulate dendrite morphology. J. Neurosci. 31, 4555–4568 (2011).
Maekawa, M. et al. Signaling from Rho to the actin cytoskeleton through protein kinases ROCK and LIM-kinase. Science 285, 895–898 (1999).
Ubhi, T. & Brown, G. W. Exploiting DNA replication stress for cancer treatment. Cancer Res. 79, 1730–1739 (2019).
Kitzing, T. M. et al. Positive feedback between Dia1, LARG, and RhoA regulates cell morphology and invasion. Genes Dev. 21, 1478–1483 (2007).
Shcherbakova, D. M. et al. Bright monomeric near-infrared fluorescent proteins as tags and biosensors for multiscale imaging. Nat. Commun. 7, 12405 (2016).
Van, Ly,D. et al. Telomere loop dynamics in chromosome end protection. Mol. Cell 71, 510–525 (2018).
Lamm, N. et al. Genomic instability in human pluripotent stem cells arises from replicative stress and chromosome condensation defects. Cell Stem Cell 18, 253–261 (2016).
Maya-Mendoza, A., Petermann, E., Gillespie, D. A., Caldecott, K. W. & Jackson, D. A. Chk1 regulates the density of active replication origins during the vertebrate S phase. EMBO J. 26, 2719–2731 (2007).
Sakaue-Sawano, A. et al. Visualizing spatiotemporal dynamics of multicellular cell-cycle progression. Cell 132, 487–498 (2008).
Gligorijevic, B., Kedrin, D., Segall, J. E., Condeelis, J. & van Rheenen, J. Dendra2 photoswitching through the mammary imaging window. J. Vis. Exp. 28, 1278 (2009).
Ritsma, L. et al. Surgical implantation of an abdominal imaging window for intravital microscopy. Nat. Protoc. 8, 583–594 (2013).
Warren, S. C. et al. Removing physiological motion from intravital and clinical functional imaging data. eLife 7, e35800 (2018).
Acknowledgements
We thank S. Page and staff at the Australian Cancer Research Foundation Telomere Analysis Centre at the Children’s Medical Research Institute for microscopy infrastructure; D. Croucher and V. Wickramshinghe for their feedback. M.B. acknowledges Bitplane for an Imaris Developer license. N.L. is supported by a Cancer Institute NSW early career fellowship and Kids Cancer Alliance grant, a Hebrew University Smorgon Foundation Fellowship, and bridging funds from University of Sydney. M.N.R. is supported by the University of Sydney Centre for Excellence in Advanced Food Enginomics. M.N. is supported by a fellowship from Cancer Institute NSW and grants from the St Vincent’s Clinic Foundation and the Australian NHMRC. V.P.M. is supported by an Australian post-graduate award from the University of Sydney. P.T. is supported by a fellowship and grants from the Australian NHMRC and support from Len Ainsworth and Sutton group. M.B. is supported by funding from the European Molecular Biology Laboratory, Australia. A.J.C. is supported by grants from the Australian NHMRC (nos. 1053195, 1106241 and 1104461), the Cancer Council NSW (RG 15-12), the Cancer Institute NSW (11/FRL/5-02) and philanthropy from S. Brown (Sydney, Australia).
Author information
Authors and Affiliations
Contributions
N.L. and A.J.C. conceived the study. N.L., M.N., D.V.L., S.G.P. and V.P.M. performed experimentation. M.N. and P.T. provided intravital imaging. M.N.R. and M.B. developed analysis tools. N.L., M.N.R., M.N., M.B. and A.J.C. analysed the data. N.L., M.B. and A.J.C. created figures and wrote the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
Extended Data Fig. 1 Nuclear F-actin is induced and dissociated in S-phase with the introduction and removal of replication stress.
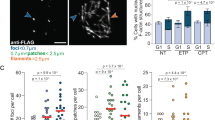
a-d, Nuclear-actin-CB expressing IMR90 (a, b), IMR90 E6E7 (c, d), and U-2OS (c, d) cultures treated with 0.4 µM APH or 500 µM HU for eight hours and stained for Lamin A/C (mean ± s.e.m, n = 3 biological replicates quantifying ≥ 58 nuclei per replicate, two-sided Fisher’s exact test). Images are single Z-plane super-resolution micrographs through the nuclear volume. e, Replication rate and fork distance in IMR90 cells ± 0.4 µM APH assayed by molecular combing [replication rate n = 181 (DMSO) and 146 (APH), and fork distance n = 80 (DMSO) and 70 (APH), sampled from three biological replicates compiled into a Tukey box plot, unpaired two-tailed t-test]. f, Immunoblots of nuclear and cytosolic extracts from cultures treated ± 0.4 µM APH for eight hours. Volume of nuclear to cytoplasmic extract is 5:1. g, Normalised time course of pyrene-labelled actin assembly using the extracts from (f). h, Super-resolution single Z-plane images from a 0.4 µM APH treated U-2OS cell expressing nuclear-actin-CB fixed and stained for Lamin A/C and Phalloidin. i, Time in S-phase before nuclear F-actin was detected in 0.4 µM APH treated U-2OS cells expressing PCNA-CB and nuclear-WT-actin or nuclear-actin-CB (mean ± s.e.m., n = 11 cells per condition counted in 1 experiment). j, Live microscopy of nuclear-actin-CB and PCNA-CB transfected U-2OS cell treated with 0.4 µM APH for 24 hours before drug washout. k, Images of nuclear-actin-CB and nuclear-WT-actin expressing U-2OS cells with nucleolar actin. l, Western blots of nuclear and cytosolic extracts from U-2OS cultures ± nuclear-actin-CB expression. Nuclear to cytoplasmic extract volume is 5:1. For f, g, j, k, and l, data representative of three biological replicates are shown. For all panels: scale bar represents 5 µm, ****p < 0.0001. Source data and unprocessed blots are provided in Source Data Extended Data Fig. 1.
Extended Data Fig. 2 mTOR signalling, IQGAP1 localization to nuclear F-actin, and Cofilin1 phosphorylation occur with replication stress.
a, Western blots of whole cell extracts from IMR90 E6E7 cells treated with 0.4 µM APH ± 200 nM INK128 or 100 nM VE822 for the indicated duration. b, Western blots of whole cell extracts from siRNA transfected U-2OS cells treated with 0.4 µM APH for eight hours. Cells were siRNA transfected 48 hours prior to extraction. c, Single z-plane from super-resolution microscopy through the nucleus of a nuclear-actin-CB and PCNA-CB expressing U-2OS cell treated with 0.4 µM APH for 24 hours then fixed and stained for IQGAP1. d, Western blots of whole cell extracts from IMR90 or U-2OS cells treated ± 0.4 µM APH for 24 hours. e, Representative still images from live microscopy of nuclear-actin-CB and PCNA-CB expressing U-2OS cells where nuclear F-actin briefly assembled before losing structural integrity. Cells were treated with 0.4 µM APH + 200 nM INK128 or 100 nM VE822. Time is hr:min relative to the first image of the series. Cells displaying this phenotype were excluded from quantitative analyses shown elsewhere. All data in this figure are representative of at least two biological replicates. For all panels: scale bar represents 5 µm. Source data and unprocessed blots are provided in Source Data Extended Data Fig. 2.
Extended Data Fig. 3 Replication stress induces alteration of S-phase nuclear architecture through ATR, mTOR, and ARP2/3.
a, Single Z-plane super-resolution micrographs of fixed nuclear-actin-CB and PCNA-CB expressing IMR90 fibroblasts treated with 0.4 µM APH ± 200 nM LatB, 200 nM INK128, or 100 nM VE822 for 24 hours. b, c, Frequency of nuclear F-actin positive S-phase (b) and non-S-phase (c) nuclei from the experiment in (a) (mean ± s.e.m., n = 3 biological replicates scoring ≥ 51 S-phase and ≥ 32 non-S-phase nuclei per replicate, two-sided Fisher’s exact test). d-f, Western blots of whole cell extracts from siRNA transfected U-2OS cells. Where applicable cells were treated with 0.4 µM APH for 8 hours. Data are representative of three biological replicates. g, Percentage of F-actin positive S-phase nuclei determined by live imaging of siRNA transfected U-2OS cells expressing nuclear-actin-CB and PCNA-CB. Cells were treated with vehicle or 0.4 µM APH (mean ± s.e.m., n = 3 biological replicates analyzing ≥ 40 cells per replicate, Fisher’s exact test). h, Examples of 70° branched actin filaments in 0.4 µM APH treated U-2OS cells. i, S-phase nuclear volume from fixed nuclear-actin-CB and PCNA-CB expressing IMR90 cells [mean ± s.e.m., n = 125 (DMSO), 206 (APH), 125 (APH+LatB), 125 (APH+INK128), and 126 (APH+VE822) cells sampled from three biological replicates compiled into a Tukey box plot, unpaired two-tailed t-test]. j, Time course of normalised S-phase nuclear volume in nuclear-actin-CB and PCNA-CB expressing U-2OS cells treated with 0.4 µM APH and transfected with the indicated siRNAs [mean ± s.e.m., n = 34 (DMSO), 36 (APH), 15 (APH + mTOR siRNA), 16 (APH + ATR siRNA), and 15 (APH + ARP2/3 siRNA) nuclei sampled from two biological replicates, one-way ANOVA]. All panels: scale bar represents 5 µm, ns = not significant, *p < 0.05, ** p < 0.01, ****p < 0.0001. Source data and unprocessed blots are provided in Source Data Extended Data Fig. 3.
Extended Data Fig. 4 Stressed replication foci localize to F-actin and the nuclear periphery.
a, PCNA-CB expressing U-2OS cell cycle (hr:min). b, Nuclear-actin-CB and PCNA-CB expressing late-S U-2OS cell treated with 0.4 µM APH for 24 hours and stained for RAD51. Arrows indicate RAD51-positive (orange) and -negative (white) actin-associated PCNA-CB foci. c, Co-localization between FANCD2 or RAD51 and late S-phase F-actin associated PCNA-CB foci (mean ± s.e.m., n = 30 cells pooled from three biological replicates). d, Early S-phase nuclear-actin-CB and PCNA-CB expressing U-2OS cell treated with 0.4 µM APH for 24 hours and stained for FANCD2. e, F-actin associated PCNA foci relative to FANCD2 in early and mid S-phase (mean ± s.e.m., n = 3 biological replicates scoring 21 cells total). f, Top: Super-resolution microscopy of single Z-planes from 3D images of nuclear-actin-CB and PCNA-CB expressing IMR90 cells treated with 0.4 µM APH ± 200 nM LatB, 200 nM INK128, or 100 nM VE822 for 24 hours. Bottom: PCNA-CB foci from the above 3D images collapsed into 2D and colour-coded for nuclear peripheral distance. g, Quantitation of (f) [mean ± s.e.m., n = 1684 (DMSO), 2155 (APH), 1695 (APH+LatB), 2001 (APH+INK128), and 2015 (APH+VE822) foci sampled from ≥ 12 nuclei across three biological replicates, Chi square test]. h, Left panels: super-resolution microscopy of single Z-planes from 3D images of U-2OS cells treated with 0.4 µM APH ± 200 nM LatB, labelled with EdU and stained for TRF2. Telomeres were assigned to one of six equal volume zones from nuclear centre to periphery. Right: Telomeres collapsed into 2D and colour-coded for nuclear volumetric zone. i, Quantitation of (h) [mean ± s.e.m., n = 2099 (DMSO), 2096 (APH), 3001 (APH+LatB) telomeres from ≥ 29 nuclei across three biological replicates, Chi square test]. All panels: Scale bar represents 5 µm, ****p < 0.0001. Source data provided in Source Data Extended Data Fig. 4.
Extended Data Fig. 5 ATR, mTOR and F-actin promote late replication foci mobility including movement along nuclear F-actin.
All data were collected from U-2OS cells expressing nuclear-actin-CB and PCNA-CB treated with or without DMSO, 0.4 µM APH, 200 nM LatB, 200 nM INK128, 100 nM VE-822, 10 mM BDM, 50 µM Blebbistatin, or 100 μM MyoVin-I. a, Average speed of late S-phase PCNA-CB foci [mean ± s.e.m., n = 30 (DMSO and APH), 26 (APH+LatB), 29 (APH+INK128), and 26 (APH+VE822) foci sampled from ≥ 7 nuclei compiled into a dot plot, unpaired two-tailed t-test]. b, c, Mean Squared Displacement (MSD) of replication foci trajectories. The lines in (b) represent individuals cells, with the inset showing MSD curves in log-log space used to calculate linear regression slopes for each nucleus, (c) are the linear regression slope values from (b) [mean ± s.d., n = 4 (APH and APH+LatB) and 5 (APH+INK128 and APH+VE822) nuclei, two-tailed unpaired t-test]. d, Examples from live-imaging of diffusive late PCNA-CB foci movement along or towards nuclear F-actin. Expanded regions are shown on the right. Arrows depict PCNA-CB foci movement. e, Still images from live imaging of cells treated as indicated. Time is hr:min relative to the first image. f, Quantitation of S-phase nuclear volume in the experiment in (e) [mean ± s.e.m, n = 34 (DMSO), 36 (APH), 18 (APH+BDM), 19 (APH+Blebbistatin), and 18 (APH+MyoVin-I) nuclei sampled from two biological replicates, one-way ANOVA]. g, h, MSD analysis of replication foci trajectories in cells treated as in (f) and displayed as in (b, c) (mean ± s.e.m., n = 4 (APH and APH+Myo-VinI) and 5 (APH+BDM and APH+Blebbistatin) nuclei, unpaired two-tailed t-test). For all panels: Scale bar represents 5 µm, ns = not significant, *p < 0.05, ** p < 0.01, ****p < 0.0001. Source data are provided in Source Data Extended Data Fig. 5.
Extended Data Fig. 6 Inhibiting actin polymerization induces cellular and molecular outcomes consistent with unrepaired replication stress.
a, Molecular combing assays to measure recovery replication rate in IMR90 E6E7 cells. Cells were treated with 0.1 µM APH and IdU for three hours, before washout and labelling with CldU as shown in Fig. 5a. Following APH washout cells were treated with or without 200 nM LatB, 200 nM INK128, 100 nM VE-822. Scale bar represents 10 µm. b, Quantitation of the experiment shown in (a) [mean ± s.e.m., n = 142 (DMSO), 132 (LatB), 138 (INK128), and 138 (VE822) forks sampled from two biological replicates compiled into a Tukey box plot, two-tailed Student’s t-test]. c, Cytogenetic chromosome preparations from U-2OS cells treated with LatB or APH for 24 hours before sample collection. Expanded images are shown below with examples of chromosome fragility indicated with red arrows. Scale bar represents 7.5 µm. d, Quantitation of the experiment depicted in (c) (n = 2 biological replicates, quantifying 78 mitoses per condition). e, Cell cycle phase duration from IMR90 E6E7-FUCCI cells treated with 200 nM [mean ± s.e.m., G1: n = 47 (DMSO and LatB), S/G2: n = 42 (DMSO) and 49 (LatB), M: n = 56 (DMSO) and 48 (LatB) cells sampled from three biological replicates and compiled into a Tukey box plot, unpaired two-tailed t-test). f, Micronuclei (left panels) and anaphase abnormalities (right panels) in IMR90 E6E7 cells following treatment with LatB. Scale bar represents 5 µm. g, Frequency of micronuclei and anaphase abnormalities in IMR90 E6E7 cells following treatment with escalating dosages of LatB (n = 2 biological replicates scoring ≥ 65 cells per replicate) For d, g, replicate means are shown as dot points, bar represents the overall mean. For all panels, ns = not significant, *p < 0.05, ****p < 0.0001. Source data are provided in Source Data Extended Data Fig. 6.
Supplementary information
Supplementary Video 1
Normal cell cycle in a U-2OS cell expressing nuclear-actin-CB and PCNA-CB. Representative maximum projection video captured using spinning-disk confocal live microscopy of a nuclear-actin-CB- and PCNA-CB-expressing U-2OS cell. Cells were transfected 72 h before imaging and treated with DMSO starting 24 h before the imaging session. Time is shown as hours:minutes:seconds relative to the first image of the series.
Supplementary Video 2
Replication stress induces nuclear F-actin. Representative maximum projection video captured using spinning-disk confocal live microscopy of a nuclear-actin-CB- and PCNA-CB-expressing U-2OS cell treated with 0.4 µM APH. Cultures were transfected 72 h before imaging and treated with APH starting 24 h before the imaging session. Time is shown as hours:minutes:seconds relative to the first image of the series.
Supplementary Video 3
LatB prevents nuclear actin polymerization in APH-treated cells. Representative maximum projection video captured using spinning-disk confocal live microscopy of a nuclear-actin-CB- and PCNA-CB-expressing U-2OS cell treated with 0.4 µM APH and 200 nM LatB. Cultures were transfected 72 h before imaging and treated with APH and LatB starting 24 h before the imaging session. Time is shown as hours:minutes:seconds relative to the first image of the series.
Supplementary Video 4
Inhibiting mTOR or ATR prevents nuclear actin polymerization in APH-treated cells. Representative maximum projection video captured using spinning-disk confocal live microscopy of a nuclear-actin-CB- and PCNA-CB-expressing U-2OS cell treated with 0.4 µM APH and 200 nM INK128, or 0.4 µM APH and 100 nM VE-822. Cultures were transfected 72 h before imaging and treated with the indicated compounds 24 h before the imaging session. Time is shown as hours:minutes:seconds relative to the first image of the series.
Supplementary Video 5
Inhibiting mTOR can result in the loss of nuclear F-actin structural integrity in cells under replication stress. Experimental conditions are identical to the cell treated with 0.4 µM APH and 200 nM INK128 in Supplementary Video 4. The F-actin structural collapse phenotype is demonstrated.
Supplementary Video 6
Demonstration of nuclear registration for PCNA chromobody foci analysis. The same cell is shown before and after registration. The video is a maximum projection video captured using spinning-disk confocal live microscopy of a nuclear-actin-CB- and PCNA-CB-expressing U-2OS cell. Cultures were transfected 72 h before imaging. Time is shown as hours:minutes:seconds relative to the first image of the series.
Supplementary Video 7
Replication stress induces increased mobility of late replication foci. Representative registered maximum projection video from spinning-disk confocal live microscopy of a late S-phase nuclear-actin-CB- and PCNA-CB-expressing U-2OS cell treated with 0.4 µM APH. Experimental conditions were the same as in Supplementary Video 2. The nucleus is registered as shown in Supplementary Video 6 and foci were tracked. Time is shown as hours:minutes:seconds relative to the first image of the series.
Supplementary Video 8
Mobility of stressed late replication foci is dependent on actin polymerization. Representative registered maximum projection video from spinning-disk confocal live microscopy of a late S-phase nuclear-actin-CB- and PCNA-CB-expressing U-2OS cell treated with 0.4 µM APH and 200 nM LatB. Experimental conditions were the same as in Supplementary Video 3. The nucleus is registered as shown in Supplementary Video 6 and foci were tracked. Time is shown as hours:minutes:seconds relative to the first image of the series.
Supplementary Video 9
Replication stress induces directed movement of late replication foci along actin fibres. Experimental and analysis conditions were the same as in Supplementary Video 7. Directed movement of a late replication foci along a nuclear F-actin filament is shown.
Source data
Source Data Fig. 1
Statistical source data Fig 1.
Source Data Fig. 1
Unprocessed western blots Fig. 1.
Source Data Fig. 2
Statistical source data Fig. 2.
Source Data Fig. 2
Unprocessed western blots Fig. 2.
Source Data Fig. 3
Statistical source data Fig. 3.
Source Data Fig. 3
Unprocessed western blots Fig. 3.
Source Data Fig. 4
Statistical source data Fig. 4.
Source Data Fig. 5
Statistical source data Fig. 5.
Source Data Fig. 5
Unprocessed western blots Fig. 5.
Source Data Fig. 6
Statistical source data Fig. 6.
Source Data Extended Data Fig. 1
Statistical source data Extended Data Fig. 1.
Source Data Extended Data Fig. 1
Unprocessed western blots Extended Data Fig. 1.
Source Data Extended Data Fig. 2
Unprocessed western blots Extended Data Fig. 2.
Source Data Extended Data Fig. 3
Statistical source data Extended Data Fig. 3.
Source Data Extended Data Fig. 3
Unprocessed western blots Extended Data Fig. 3.
Source Data Extended Data Fig. 4
Statistical source data Extended Data Fig. 4.
Source Data Extended Data Fig. 5
Statistical source data Extended Data Fig. 5.
Source Data Extended Data Fig. 6
Statistical source data Extended Data Fig. 6.
Rights and permissions
About this article
Cite this article
Lamm, N., Read, M.N., Nobis, M. et al. Nuclear F-actin counteracts nuclear deformation and promotes fork repair during replication stress. Nat Cell Biol 22, 1460–1470 (2020). https://doi.org/10.1038/s41556-020-00605-6
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41556-020-00605-6
This article is cited by
-
DNA replication and replication stress response in the context of nuclear architecture
Chromosoma (2024)
-
Genome maintenance meets mechanobiology
Chromosoma (2024)
-
RHOJ controls EMT-associated resistance to chemotherapy
Nature (2023)
-
Nuclear myosin VI maintains replication fork stability
Nature Communications (2023)
-
Cancer cells remodel nuclear actin filaments to resist chemotherapy
Nature (2023)