Abstract
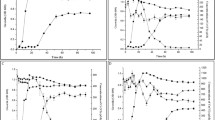
The nitrated compounds 2,4-dinitrotoluene (2,4-DNT), 2,4,6-trinitrotoluene (TNT), and pentaerythritol tetranitrate (PETN) are toxic xenobiotics widely used in various industries. They often coexist as environmental contaminants. The aims of this study were to evaluate the transformation of 100 mg L−1 of TNT, 2,4-DNT, and PETN by Raoultella planticola M30b and Rhizobium radiobacter M109c and identify enzymes that may participate in the transformation. These strains were selected from 34 TNT transforming bacteria. Cupriavidus metallidurans DNT was used as a reference strain for comparison purposes. Strains DNT, M30b and M109c transformed 2,4-DNT (100%), TNT (100, 94.7 and 63.6%, respectively), and PETN (72.7, 69.3 and 90.7%, respectively). However, the presence of TNT negatively affects 2,4-DNT and PETN transformation (inhibition > 40%) in strains DNT and M109c and fully inhibited (100% inhibition) 2,4-DNT transformation in R. planticola M30b.
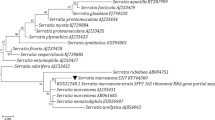
Genomes of R. planticola M30b and R. radiobacter M109c were sequenced to identify genes related with 2,4-DNT, TNT or PETN transformation. None of the tested strains presented DNT oxygenase, which has been previously reported in the transformation of 2,4-DNT. Thus, unidentified novel enzymes in these strains are involved in 2,4-DNT transformation. Genes encoding enzymes homologous to the previously reported TNT and PETN-transforming enzymes were identified in both genomes. R. planticola M30b have homologous genes of PETN reductase and xenobiotic reductase B, while R. radiobacter M109c have homologous genes to GTN reductase and PnrA nitroreductase. The ability of these strains to transform explosive mixtures has a potentially biotechnological application in the bioremediation of contaminated environments.







Similar content being viewed by others
Data availability
All relevant data appear in the article and its Supporting Information files. The raw data of the transformation experiments appear in the Online Resource file “Raw data.xlsx”. Strains R. planticola M30b and R. radiobacter M109c are available in the bacteria collection of the Microbiology Department of the School of Science at Pontificia Universidad Javeriana (CMPUJ, Bogota, Colombia) under codes CMPUJ470 and CMPUJ471, respectively. CMPUJ collection are register in World Federation for Culture Collections. The genome sequences of strains R. planticola M30b and R. radiobacter M109c are deposited in the GeneBank database under accession Nos. GCA_006757685.1 and GCA_006757705.1, respectively. Sequences of the fosmids M30b_B4 and M109c_G12 are deposited in the GeneBank database under accession Nos. MT009031 and MT009032, respectively.
References
Agrawal J, Hodgson R (2007) Synthetic routes to nitrate esters. In: Agrawal J, Hodgson R (eds) Organic Chemistry of Explosives, 1st edn. Wiley, England, pp 87–124
Andrews S (2010) FastQC: a quality control tool for high throughput sequence data [cited 2019 Jan 15]. Available from: http://www.bioinformatics.babraham.ac.uk/projects/fastqc
Arbeli Z, Garcia-Bonilla E, Pardo C, Peña L, Ramos E, Velásquez T et al (2016) Persistence of pentolite (PETN and TNT) in soil microcosms and microbial enrichment cultures. Environ Sci Pollut Res Int 23:9144–9155
Avila-Arias H, Avellaneda H, Garzón V, Rodríguez G, Arbeli Z, Garcia-Bonilla E, Villegas-Plazas S, Roldan F (2017) Screening for biosurfactant production by 2,4,6-trinitrotoluene-transforming bacteria. J Appl Microbiol 123:401–413
Aziz R, Bartels D, Best A, DeJongh M, Disz T, Edwards R, Formsma K, Gerdes S, Glass E, Kubal M, Meyer F, Olsen G, Olson R, Osterman A, Overbeek R, McNeil L, Paarmann D, Paczian T, Parrello B, Pusch G, Reich C, Stevens R, Vassieva O, Vonstein V, Wilke A, Zagnitko O (2008) The RAST Server: rapid annotations using subsystems technology. BMC genomics 9:75
Binks P, French C, Nicklin S, Bruce N (1996) Degradation of pentaerythritol tetranitrate by Enterobacter cloacae PB2. Appl Environ Microbiol 62:1214–1219
Bolger A, Lohse M, Usadel B (2014) Trimmomatic: a flexible trimmer for Illumina sequence data. Bioinformatics 30(15):2114–2120
Caballero A, Lázaro J, Ramos J, Esteve-Núñez A (2005) PnrA, a new nitroreductase-family enzyme in the TNT-degrading strain Pseudomonas putida JLR11. Environ Microbiol 7(8):1211–1219
Claus H, Perret N, Bausinger T, Fels G, Preu J, Konig H (2007) TNT transformation products are affected by the growth conditions of Raoultella terrigena. Biotechnol Lett 29:411–419
Duarte M, Jauregui R, Vilchez-Vargas R, Junca H, Pieper D (2014) AromaDeg, a novel database for phylogenomics of aerobic bacterial degradation of aromatics. Database 2014:1–12
Ebert S, Fischer P, Knackmuss H (2001) Converging catabolism of 2,4,6-trinitrophenol (picric acid) and 2,4-dinitrophenol by Nocardioides simplex FJ2-1A. Biodegradation 12:367–376
EPA (2006) Method 8330B (SW-846): Nitroaromatics, nitramines, and nitrate esters by high performance liquid chromatography (HPLC), United States, pp 1–31
EPA (2010) Provisional Peer-Reviewed Toxicity Values for Pentaerythritol tetranitrate (PETN), United States pp.1–22. EPA/690/R-10/021F
EPA (2014) Technical fact sheet – 2,4,6-Trinitrotoluene (TNT), United States pp.1–8. EPA 505-F-14–009 Office of Solid Waste and Emergency Response
EPA (2017) Technical fact sheet – Dinitrotoluenes (DNT), United States pp.1–8. EPA 505-F-17–010 Office of Land and Emergency Management
Esteve-Núñez A, Caballero A, Ramos J (2001) Biological degradation of 2,4,6-trinitrotoluene. Microbiol Mol Biol Rev 65:335–352
Fernández M, Duque E, Pizarro-Tobías P, van Dillewijn P, Wittich R, Ramos J (2009) Microbial responses to xenobiotic compounds. Identification of genes that allow Pseudomonas putida KT2440 to cope with 2,4,6-trinitrotoluene. Microb Biotechnol 2(2):287–294
Fida T, Palamuru S, Pandey G, Spain J (2014) Aerobic biodegradation of 2,4-dinitroanisole by Nocardioides sp. strain JS1661. Appl Environ Microbiol 80(24):7725–7731
French C, Nicklin S, Bruce N (1998) Aerobic degradation of 2,4,6-trinitrotoluene by Enterobacter cloacae PB2 and by pentaerythritol tetranitrate reductase. Appl Environ Microbiol 64(8):2864–2868
George S, Huggins-Clark G, Brooks L (2001) Use of a Salmonella microsuspension bioassay to detect the mutagenicity of munitions compounds at low concentrations. Mutat Res 490(1):45–56
Gorecki S, Nesslany F, Hubé D, Mullot J, Vasseur P, Marchioni E et al (2017) Human health risks related to the consumption of foodstuffs of plant and animal origin produced on a site polluted by chemical munitions of the First World War. Sci Total Environ 599–600:314–323
Gumuscu B, Tekinay T (2013) Effective biodegradation of 2,4,6-trinitrotoluene using a novel bacterial strain isolated from TNT-contaminated soil. Int Biodeterior Biodegradation 85:35–41
HaÏdour A, Ramos J (1996) Identification of products resulting from the biological reduction of 2,4,6-trinitrotoluene, 2,4-dinitrotoluene, and 2,6-dinitrotoluene by Pseudomonas sp. Environ Sci Technol 30:2365–2370
Han S (2008) In situ bioremediation and natural attenuation of dinitrotoluenes and trinitrotoluene Dissertarion Georgia Institute of Technology, Atlanta
Hudcova T, Halecky M, Kozliak E, Stiborova M, Paca J (2011) Aerobic degradation of 2,4-dinitrotoluene by individual bacterial strains and defined mixed population in submerged cultures. J Hazard Mater 192(2):605–613
Husserl J (2011) Biodegradation of nitroglycerin as a growth substrate: A basis for natural attenuation and bioremediation. Dissertation, Georgia Institute of Technology, Atlanta
Jackson R, Rylott E, Fournier D, Hawari J, Bruce N (2007) Exploring the biochemical properties and remediation applications of the unusual explosive-degrading P450 system XplA/B. Proc Natl Acad Sci USA 104:16822–16827
Johnson G, Jain R, Spain J (2002) Origins of the 2,4-dinitrotoluene pathway. J Bacteriol 184:4219–4232
Khan M, Lee J, Park J (2013) A toxicological review on potential microbial degradation intermediates of 2,4,6-trinitrotoluene, and its implications in bioremediation. KSCE Journal of Civil Engineering 17(6):1223–1231
Khan M, Lee J, Yoo K, Kim S, Park J (2015) Improved TNT detoxification by starch addition in a nitrogen-fixing Methylophilus-dominant aerobic microbial consortium. J Hazard Mater 300:873–881
Küce P, Coral G, Kantar Ç (2015) Biodegradation of 2,4-dinitrotoluene (DNT) by Arthrobacter sp. K1 isolated from a crude oil contaminated soil. Ann Microbiol 65(1):467–476
Kundu D, Hazra C, Chaudhari A (2015) Biodegradation of 2,4-dinitrotoluene with Rhodococcus pyridinivorans NT2: characteristics, kinetic modeling, physiological responses and metabolic pathway. RSC Advances 5(49):38818–38829
Labidi M, Ahmad D, Halasz A, Hawari J (2001) Biotransformation and partial mineralization of the explosive 2,4,6-trinitrotoluene (TNT) by rhizobia. Can J Microbiol 47:559–566
Luo R, Liu B, Xie Y, Li Z, Huang W, Yuan J et al (2012) SOAPdenovo2: an empirically improved memory-efficient short-read de novo assembler. GigaScience 1(1):1–18
Martin J, Comfort S, Shea P, Kokjohn T, Drijbe R (1997) Denitration of 2,4,6-trinitrotoluene by Pseudomonas savastanoi. Can J Microbiol 43:447–455
Muter O, Potapova K, Limane B, Sproge K, Jakobsone I, Cepurnieks G, Bartkevics V (2012) The role of nutrients in the biodegradation of 2,4,6-trinitrotoluene in liquid and soil. J Environ Manage 98:51–55
Nishino S, Paoli G, Spain J (2000) Aerobic degradation of dinitrotoluenes and pathway for bacterial degradation of 2,6-dinitrotoluene. Appl Environ Microbiol 66:2139–2147
Overbeek R, Olson R, Pusch G, Olsen G, Davis J, Disz T et al (2014) The SEED and the Rapid Annotation of microbial genomes using Subsystems Technology (RAST). Nucleic Acids Res 42:D206-214
Parales R, Spain J, Jhonson G (2005) Bacterial degradation of DNT and TNT mixtures. Davis (CA): University of California, Georgia Institute of Technology, Air Force; Proyect No.: CU1212. Sponsored by Strategic Environmental Research & Development Program
Phelan J, Web S (2002) Chemical sensing for buried landmines - fundamental processes influencing trace chemical detection. Mine detection dogs training, operations and odor detection. Geneva international center for humanitarian demining, Geneva, pp 209–286
Rice E, Baird R, Eaton A, Clesceri L (2012) 4500-NO2-. In: Rice E (ed) Standard methods for the examination of water and wastewater, 22nd edn. APHA, Washington, DC
Riefler R, Smets B (2002) NAD(P)H:flavin mononucleotide oxidoreductase inactivation during 2,4,6-trinitrotoluene reduction. Appl Environ Microbiol 68:1690–1696
Sabbioni G, Rumler R (2007) Biomonitoring of workers cleaning up ammunition waste sites. Biomarkers 12(6):559–573
Sagi-Ben Moshe S, Ronen Z, Dahan O, Weisbrod N, Groisman L, Adar E, Nativ R (2009) Sequential biodegradation of TNT, RDX and HMX in a mixture. Environ Pollut 157:2231–2238
Sanjust E, Rinaldi A, Rescigno A, Porcu M, Alberti G, Rinaldi A, Finazzi-Agro A (1995) A hydroxyquinone with amine oxidase activity: preparation and properties. Biochem Biophys Res Commun 208:825–834
Seemann T (2017) Shovill: Faster SPAdes assembly of Illumina reads [cited 2019 Jan 15]. Available from: https://github.com/tseemann/shovill
Shemer B, Yagur-Kroll S, Hazan C, Belkin S (2018) Aerobic transformation of 2,4-dinitrotoluene by Escherichia coli and its implications for the detection of trace explosives. Appl Environ Microbiol 84(4):e01729-e11717
Snape J, Walkley N, Morby A, Nicklin S, White G (1997) Purification, properties, and sequence of glycerol trinitrate reductase from Agrobacterium radiobacter. J Bacteriol 179(24):7796–7802
Snellinx Z, Taghavi S, Vangronsveld J, Van der Lelie D (2003) Microbial consortia that degrade 2,4-DNT by interspecies metabolism: Isolation and characterisation. Biodegradation 14(1):19–29
Spanggord R, Mortelmans K, Griffin A, Simmon V (1982) Mutagenicity in Salmonella typhimurium and structure-activity relationships of wastewater components emanating from the manufacture of trinitrotoluene. Environ Mutagen 4(2):163–179
Spanggord R, Spain J, Nishino S, Mortelmans K (1991) Biodegradation of 2,4-dinitrotoluene by a Pseudomonas sp. Appl Environ Microbiol 57:3200–3205
Stenuit B, Agathos S (2010) Microbial 2,4,6-trinitrotoluene degradation: could we learn from (bio)chemistry for bioremediation and vice versa? Appl Microbiol Biotechnol 88:1043–1064
Stenuit B, Agathos S (2011) Biodegradation and bioremediation of TNT and other nitro explosives. In: Murray M (ed) Comprehensive Biotechnology, 2nd edn. Academic Press, Burlington, VT, pp 167–181
Suen W, Spain J (1993) Cloning and characterization of Pseudomonas sp. strain DNT genes for 2,4-dinitrotoluene degradation. J Bacteriol 175(6):1831–1837
Thijs S, Van Hamme J, Gkorezis P, Rineau F, Weyens N, Vangronsveld J (2014) Draft genome sequence of Raoultella ornithinolytica TNT, a trinitrotoluene-denitrating and plant growth-promoting strain isolated from explosive-contaminated soil. Genome Announc 2(3):e00491-e1414
Van Dillewijn P, Wittich R, Caballero A, Ramos J (2008) Type II hydride transferases from different microorganisms yield nitrite and diarylamines from polynitroaromatic compounds. Appl Environ Microbiol 74:6820–6823
White G, Snape J, Nicklin S (1996) Biodegradation of glycerol trinitrate and pentaerythritol tetranitrate by Agrobacterium radiobacter. Appl Environ Microbiol 62(2):637–642
Williams R, Rathbone D, Scrutton N, Bruce N (2004) Biotransformation of explosives by the old yellow enzyme family of flavoproteins. Appl Environ Microbiol 70(6):3566–3574
Wittich R, Haïdour A, Van Dillewijn P, Ramos J (2008) OYE flavoprotein reductases initiate the condensation of TNT-derived intermediates to secondary diarylamines and nitrite. Environ Sci Technol 42:734–739
Xiao Y, Wu J, Liu H, Wang S, Liu S, Zhou N (2006) Characterization of genes involved in the initial reactions of 4-chloronitrobenzene degradation in Pseudomonas putida ZWL73. Appl Microbiol Biotechnol 73(1):166–171
Zhang C, Hughes J, Nishino S, Spain J (2000) Slurry-phase biological treatment of 2,4-dinitrotoluene and 2,6-dinitrotoluene: role of bioaugmentation and effects of high dinitrotoluene concentrations. Environ Sci Technol 34:2810–2816
Zhang M, Liu G, Song K, Wang Z, Zhao Q, Li S, Ye Z (2015) Biological treatment of 2,4,6-trinitrotoluene (TNT) red water by immobilized anaerobic–aerobic microbial filters. Chem Eng J 259:876–884
Zhuang L (2007) Remediation of pentaerythritol tetranitrate (PETN) contaminated water and soil. Dissertation. Earth Sciences, University of Waterloo. Ontario, Canada
Zhuang L, Gui L, Gillham R (2012) Biodegradation of pentaerythritol tetranitrate (PETN) by anaerobic consortia from a contaminated site. Chemosphere 89:810–816
Acknowledgments
The authors thank Dr. Jim Spain for providing the reference strain and Drs. Johanna Husserl and Howard Junca for contributing their knowledge and advice towards the development of this study.
Funding
This work was funded by INDUMIL (Colombian Military Industry—https://www.indumil.gov.co/). Colciencias (Colombian Innovation Agency—https://minciencias.gov.co/) awarded Hernan Avellaneda "Convocatoria 727 de 2015". The funders had no role in study design, data collection and analysis, decision to publish, or manuscript preparation. Soil sampling was approved by the Colombian National Authority of Environmental Licenses (Autoridad Nacional de Licencias Ambientales—ANLA) under Scientific Research Permit on Biodiversity No. 545 dated September 18, 2009 and amended by Resolution No. 450 dated May 15, 2013.
Author information
Authors and Affiliations
Contributions
All authors contributed to the study conception and design. Material preparation, data collection, and analysis were performed by Hernán Avellaneda. The first draft of the manuscript was written by Hernán Avellaneda and all authors commented on previous versions of the manuscript. All authors read and approved the final manuscript.
Corresponding author
Ethics declarations
Conflict of interest
The authors declare no conflicts of interest.
Ethical approval
Pontificia Universidad Javeriana ethics committee approved the present study in session held on April 11th.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Avellaneda, H., Arbeli, Z., Teran, W. et al. Transformation of TNT, 2,4-DNT, and PETN by Raoultella planticola M30b and Rhizobium radiobacter M109 and exploration of the associated enzymes. World J Microbiol Biotechnol 36, 190 (2020). https://doi.org/10.1007/s11274-020-02962-8
Received:
Accepted:
Published:
DOI: https://doi.org/10.1007/s11274-020-02962-8