Abstract
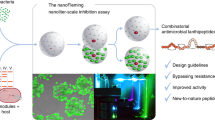
The discovery of antibiotics was one of the fundamental stages in the development of humanity, leading to a dramatic increase in the life expectancy of millions of people all over the world. The uncontrolled use of antibiotics resulted in the selection of resistant strains of bacteria, limiting the effectiveness of antimicrobial therapy nowadays. Antimicrobial peptides (AMPs) were considered promising candidates for next-generation antibiotics for a long time. However, the practical application of AMPs is restricted by their low therapeutic indices, impaired pharmacokinetics, and pharmacodynamics, which is predetermined by their peptide structure. Nevertheless, the DNA-encoded nature of AMPs enables creating broad repertoires of artificial biodiversity of antibiotics, making them versatile templates for the directed evolution of antibiotic activity. Lantibiotics are a unique class of AMPs with an expanded chemical space. A variety of post-translational modifications, mechanisms of action on bacterial membranes, and DNA-encoded nature make them a convenient molecular template for creating highly representative libraries of antimicrobial compounds. Isolation of new drug candidates from this synthetic biodiversity is extremely attractive but requires high-throughput screening of antibiotic activity. The combination of synthetic biology and ultrahigh-throughput microfluidics allows implementing the concept of directed evolution of lantibiotics for accelerated creation of new promising drug candidates.








Similar content being viewed by others
Abbreviations
- AMP:
-
antimicrobial peptides
- Dha:
-
dehydroalanine
- Dhb:
-
dehydrobutyrine
- PE:
-
phosphatidylethanolamine
- RiPPs:
-
ribosomally synthesized and post-translationally modified peptides
- UEV domain:
-
ubiquitin E2 variant domain
REFERENCES
Davies, J., and Davies, D. (2010) Origins and evolution of antibiotic resistance, Microbiol. Mol. Biol. Rev., 74, 417-433, doi: https://doi.org/10.1128/mmbr.00016-10.
O’Neill, J. (2014) Antimicrobial Resistance: Tackling A Crisis For The Health And Wealth Of Nations, Review on Antimicrobial Resistance, London.
Czepiel, J., Drozdz, M., Pituch, H., Kuijper, E. J., Perucki, W., et al. (2019) Clostridium difficile infection: review, Eur. J. Clin. Microbiol. Infect. Dis., 38, 1211-1221, doi: https://doi.org/10.1007/s10096-019-03539-6.
Wang, J., Xiong, Z., Meng, H., Wang, Y., and Wang, Y. (2012) Synthetic biology triggers new era of antibiotics development, Subcell. Biochem., 64, 95-114, doi: https://doi.org/10.1007/978-94-007-5055-5_5.
Foulston, L. (2019) Genome mining and prospects for antibiotic discovery, Curr. Opin. Microbiol., 51, 1-8, doi: https://doi.org/10.1016/j.mib.2019.01.001.
Terekhov, S. S., Smirnov, I. V., Malakhova, M. V., Samoilov, A. E., Manolov, A. I., et al. (2018) Ultrahigh-throughput functional profiling of microbiota communities, Proc. Natl. Acad. Sci. USA, 115, 9551-9556, doi: https://doi.org/10.1073/pnas.1811250115.
Stokes, J. M., Yang, K., Swanson, K., Jin, W., Cubillos-Ruiz, A., et al. (2020) A deep learning approach to antibiotic discovery, Cell, 180, 688-702, doi: https://doi.org/10.1016/j.cell.2020.01.021.
Lau, J. L., and Dunn, M. K. (2018) Therapeutic peptides: historical perspectives, current development trends, and future directions, Bioorg. Med. Chem., 26, 2700-2707, doi: https://doi.org/10.1016/j.bmc.2017.06.052.
Fosgerau, K., and Hoffmann, T. (2015) Peptide therapeutics: current status and future directions, Drug Discov. Today, 20, 122-128, doi: https://doi.org/10.1016/j.drudis.2014.10.003.
Lei, J., Sun, L., Huang, S., Zhu, C., Li, P., et al. (2019) The antimicrobial peptides and their potential clinical applications, Am. J. Transl. Res., 11, 3919-3931.
Kang, S. J., Park, S. J., Mishig-Ochir, T., and Lee, B. J. (2014) Antimicrobial peptides: therapeutic potentials, Expert. Rev. Anti. Infect. Ther., 12, 1477-1486, doi: https://doi.org/10.1586/14787210.2014.976613.
Ortega, M. A., and van der Donk, W. A. (2016) New insights into the biosynthetic logic of ribosomally synthesized and post-translationally modified peptide natural products, Cell Chem. Biol., 23, 31-44, doi: https://doi.org/10.1016/j.chembiol.2015.11.012.
McIntosh, J. A., Donia, M. S., and Schmidt, E. W. (2009) Ribosomal peptide natural products: bridging the ribosomal and nonribosomal worlds, Nat. Prod. Rep., 26, 537-559, doi: https://doi.org/10.1039/b714132g.
Muller, M. M. (2018) Post-translational modifications of protein backbones: unique functions, mechanisms, and challenges, Biochemistry, 57, 177-185, doi: https://doi.org/10.1021/acs.biochem.7b00861.
Hudson, G. A., and Mitchell, D. A. (2018) RiPP antibiotics: biosynthesis and engineering potential, Curr. Opin. Microbiol., 45, 61-69, doi: https://doi.org/10.1016/j.mib.2018.02.010.
Mullane, K., Lee, C., Bressler, A., Buitrago, M., Weiss, K., et al. (2015) Multicenter, randomized clinical trial to compare the safety and efficacy of LFF571 and vancomycin for Clostridium difficile infections, Antimicrob. Agents Chemother., 59, 1435-1440, doi: https://doi.org/10.1128/aac.04251-14.
Poorinmohammad, N., Bagheban-Shemirani, R., and Hamedi, J. (2019) Genome mining for ribosomally synthesised and post-translationally modified peptides (RiPPs) reveals undiscovered bioactive potentials of actinobacteria, Antonie Van Leeuwenhoek, 112, 1477-1499, doi: https://doi.org/10.1007/s10482-019-01276-6.
Velasquez, J. E., and van der Donk, W. A. (2011) Genome mining for ribosomally synthesized natural products, Curr. Opin. Chem. Biol., 15, 11-21, doi: https://doi.org/10.1016/j.cbpa.2010.10.027.
Arnison, P. G., Bibb, M. J., Bierbaum, G., Bowers, A. A., Bugni, T. S., et al. (2013) Ribosomally synthesized and post-translationally modified peptide natural products: overview and recommendations for a universal nomenclature, Nat. Prod. Rep., 30, 108-160, doi: https://doi.org/10.1039/C2NP20085F.
Repka, L. M., Chekan, J. R., Nair, S. K., and van der Donk, W. A. (2017) Mechanistic understanding of lanthipeptide biosynthetic enzymes, Chem. Rev., 117, 5457-5520, doi: https://doi.org/10.1021/acs.chemrev.6b00591.
Kleerebezem, M. (2004) Quorum sensing control of lantibiotic production; nisin and subtilin autoregulate their own biosynthesis, Peptides, 25, 1405-1414, doi: https://doi.org/10.1016/j.peptides.2003.10.021.
Willey, J. M., Willems, A., Kodani, S., and Nodwell, J. R. (2006) Morphogenetic surfactants and their role in the formation of aerial hyphae in Streptomyces coelicolor, Mol. Microbiol., 59, 731-742, doi: https://doi.org/10.1111/j.1365-2958.2005.05018.x.
Bierbaum, G., Gotz, F., Peschel, A., Kupke, T., van de Kamp, M., and Sahl, H. G. (1996) The biosynthesis of the lantibiotics epidermin, gallidermin, Pep5 and epilancin K7, Antonie Van Leeuwenhoek, 69, 119-127, doi: https://doi.org/10.1007/BF00399417.
Zhang, Q., Yu, Y., Velasquez, J. E., and van der Donk, W. A. (2012) Evolution of lanthipeptide synthetases, Proc. Natl. Acad. Sci. USA, 109, 18361-18366, doi: https://doi.org/10.1073/pnas.1210393109.
Shi, Y., Yang, X., Garg, N., and van der Donk, W. A. (2011) Production of lantipeptides in Escherichia coli, J. Am. Chem. Soc., 133, 2338-2341, doi: https://doi.org/10.1021/ja109044r.
Garg, N., Salazar-Ocampo, L. M., and van der Donk, W. A. (2013) In vitro activity of the nisin dehydratase NisB, Proc. Natl. Acad. Sci. USA, 110, 7258-7263, doi: https://doi.org/10.1073/pnas.1222488110.
Ortega, M. A., Hao, Y., Zhang, Q., Walker, M. C., van der Donk, W. A., and Nair, S. K. (2015) Structure and mechanism of the tRNA-dependent lantibiotic dehydratase NisB, Nature, 517, 509-512, doi: https://doi.org/10.1038/nature13888.
Siezen, R. J., Kuipers, O. P., and de Vos, W. M. (1996) Comparison of lantibiotic gene clusters and encoded proteins, Antonie Van Leeuwenhoek, 69, 171-184, doi: https://doi.org/10.1007/BF00399422.
Goto, Y., Okesli, A., and van der Donk, W. A. (2011) Mechanistic studies of Ser/Thr dehydration catalyzed by a member of the LanL lanthionine synthetase family, Biochemistry, 50, 891-898, doi: https://doi.org/10.1021/bi101750r.
Hollenstein, K., Dawson, R. J., and Locher, K. P. (2007) Structure and mechanism of ABC transporter proteins, Curr. Opin. Struct. Biol., 17, 412-418, doi: https://doi.org/10.1016/j.sbi.2007.07.003.
Kuipers, A., de Boef, E., Rink, R., Fekken, S., Kluskens, L. D., et al. (2004) NisT, the transporter of the lantibiotic nisin, can transport fully modified, dehydrated, and unmodified prenisin and fusions of the leader peptide with non-lantibiotic peptides, J. Biol. Chem., 279, 22176-22182, doi: https://doi.org/10.1074/jbc.M312789200.
Lagedroste, M., Smits, S. H. J., and Schmitt, L. (2017) Substrate specificity of the secreted nisin leader peptidase NisP, Biochemistry, 56, 4005-4014, doi: https://doi.org/10.1021/acs.biochem.7b00524.
Velasquez, J. E., Zhang, X., and van der Donk, W. A. (2011) Biosynthesis of the antimicrobial peptide epilancin 15X and its N-terminal lactate, Chem. Biol., 18, 857-867, doi: https://doi.org/10.1016/j.chembiol.2011.05.007.
Huang, E., and Yousef, A. E. (2015) Biosynthesis of paenibacillin, a lantibiotic with N-terminal acetylation, by Paenibacillus polymyxa, Microbiol. Res., 181, 15-21, doi: https://doi.org/10.1016/j.micres.2015.08.001.
Allgaier, H., Jung, G., Werner, R. G., Schneider, U., and Zähner, H. (1985) Elucidation of the structure of epidermin, a ribosomally synthesized, tetracyclic heterodetic polypeptide antibiotic, Angew. Chem. Internat. Ed. Engl., 24, 1051-1053, doi: https://doi.org/10.1002/anie.198510511.
De Arauz, L. J., Jozala, A. F., Mazzola, P. G., and Vessoni Penna, T. C. (2009) Nisin biotechnological production and application: a review, Trends Food Sci. Technol., 20, 146-154, doi: https://doi.org/10.1016/j.tifs.2009.01.056.
Lepak, A. J., Marchillo, K., Craig, W. A., and Andes, D. R. (2015) In vivo pharmacokinetics and pharmacodynamics of the lantibiotic NAI-107 in a neutropenic murine thigh infection model, Antimicrob. Agents Chemother., 59, 1258-1264, doi: https://doi.org/10.1128/AAC.04444-14.
Thomsen, T. T., Mojsoska, B., Cruz, J. C., Donadio, S., Jenssen, H., Lobner-Olesen, A., and Rewitz, K. (2016) The Lantibiotic NAI-107 efficiently rescues Drosophila melanogaster from infection with methicillin-resistant Staphylococcus aureus USA300, Antimicrob. Agents Chemother., 60, 5427-5436, doi: https://doi.org/10.1128/AAC.02965-15.
Castiglione, F., Lazzarini, A., Carrano, L., Corti, E., Ciciliato, I., et al. (2008) Determining the structure and mode of action of microbisporicin, a potent lantibiotic active against multiresistant pathogens, Chem. Biol., 15, 22-31, doi: https://doi.org/10.1016/j.chembiol.2007.11.009.
Chatterjee, C., Miller, L. M., Leung, Y. L., Xie, L., Yi, M., Kelleher, N. L., and van der Donk, W. A. (2005) Lacticin 481 synthetase phosphorylates its substrate during lantibiotic production, J. Am. Chem. Soc., 127, 15332-15333, doi: https://doi.org/10.1021/ja0543043.
Dong, S. H., Tang, W., Lukk, T., Yu, Y., Nair, S. K., and van der Donk, W. A. (2015) The enterococcal cytolysin synthetase has an unanticipated lipid kinase fold, eLife, 4, e07607, doi: https://doi.org/10.7554/eLife.07607.
Ma, H., Gao, Y., Zhao, F., and Zhong, J. (2015) Individual catalytic activity of two functional domains of bovicin HJ50 synthase BovM, Wei Sheng Wu Xue Bao, 55, 50-58.
Shimafuji, C., Noguchi, M., Nishie, M., Nagao, J., Shioya, K., Zendo, T., Nakayama, J., and Sonomoto, K. (2015) In vitro catalytic activity of N-terminal and C-terminal domains in NukM, the post-translational modification enzyme of nukacin ISK-1, J. Biosci. Bioeng., 120, 624-629, doi: https://doi.org/10.1016/j.jbiosc.2015.03.020.
Tang, W., Jimenez-Oses, G., Houk, K. N., and van der Donk, W. A. (2015) Substrate control in stereoselective lanthionine biosynthesis, Nat. Chem., 7, 57-64, doi: https://doi.org/10.1038/nchem.2113.
Thibodeaux, C. J., Ha, T., and van der Donk, W. A. (2014) A price to pay for relaxed substrate specificity: a comparative kinetic analysis of the class II lanthipeptide synthetases ProcM and HalM2, J. Am. Chem. Soc., 136, 17513-17529, doi: https://doi.org/10.1021/ja5089452.
Cubillos-Ruiz, A., Berta-Thompson, J. W., Becker, J. W., van der Donk, W. A., and Chisholm, S. W. (2017) Evolutionary radiation of lanthipeptides in marine cyanobacteria, Proc. Natl. Acad. Sci. USA, 114, E5424-E5433, doi: https://doi.org/10.1073/pnas.1700990114.
Nishie, M., Sasaki, M., Nagao, J., Zendo, T., Nakayama, J., and Sonomoto, K. (2011) Lantibiotic transporter requires cooperative functioning of the peptidase domain and the ATP binding domain, J. Biol. Chem., 286, 11163-11169, doi: https://doi.org/10.1074/jbc.M110.212704.
Kuipers, A., Meijer-Wierenga, J., Rink, R., Kluskens, L. D., and Moll, G. N. (2008) Mechanistic dissection of the enzyme complexes involved in biosynthesis of lacticin 3147 and nisin, Appl. Environ. Microbiol., 74, 6591-6597, doi: https://doi.org/10.1128/AEM.01334-08.
Caetano, T., Barbosa, J., Moesker, E., Sussmuth, R. D., and Mendo, S. (2014) Bioengineering of lanthipeptides in Escherichia coli: assessing the specificity of lichenicidin and haloduracin biosynthetic machinery, Res. Microbiol., 165, 600-604, doi: https://doi.org/10.1016/j.resmic.2014.07.006.
Galvin, M., Hill, C., and Ross, R. P. (1999) Lacticin 3147 displays activity in buffer against gram-positive bacterial pathogens which appear insensitive in standard plate assays, Lett. Appl. Microbiol., 28, 355-358, doi: https://doi.org/10.1046/j.1365-2672.1999.00550.x.
Oman, T. J., and van der Donk, W. A. (2009) Insights into the mode of action of the two-peptide lantibiotic haloduracin, ACS Chem. Biol., 4, 865-874, doi: https://doi.org/10.1021/cb900194x.
Xin, B., Zheng, J., Xu, Z., Li, C., Ruan, L., Peng, D., and Sun, M. (2015) Three novel lantibiotics, ticins A1, A3, and A4, have extremely stable properties and are promising food biopreservatives, Appl. Environ. Microbiol., 81, 6964-6972, doi: https://doi.org/10.1128/aem.01851-15.
Crowther, G. S., Baines, S. D., Todhunter, S. L., Freeman, J., Chilton, C. H., and Wilcox, M. H. (2013) Evaluation of NVB302 versus vancomycin activity in an in vitro human gut model of Clostridium difficile infection, J. Antimicrob. Chemother., 68, 168-176, doi: https://doi.org/10.1093/jac/dks359.
Louie, T. J., Emery, J., Krulicki, W., Byrne, B., and Mah, M. (2009) OPT-80 eliminates Clostridium difficile and is sparing of bacteroides species during treatment of C. difficile Infection, Antimicrob. Agents Chemother., 53, 261-263, doi: https://doi.org/10.1128/aac.01443-07.
Knerr, P. J., and van der Donk, W. A. (2012) Discovery, biosynthesis, and engineering of lantipeptides, Annu. Rev. Biochem., 81, 479-505, doi: https://doi.org/10.1146/annurev-biochem-060110-113521.
Kodani, S., Hudson, M. E., Durrant, M. C., Buttner, M. J., Nodwell, J. R., and Willey, J. M. (2004) The SapB morphogen is a lantibiotic-like peptide derived from the product of the developmental gene ramS in Streptomyces coelicolor, Proc. Natl. Acad. Sci. USA, 101, 11448-11453, doi: https://doi.org/10.1073/pnas.0404220101.
Meindl, K., Schmiederer, T., Schneider, K., Reicke, A., Butz, D., et al. (2010) Labyrinthopeptins: a new class of carbacyclic lantibiotics, Angew. Chem. Int. Ed. Engl., 49, 1151-1154, doi: https://doi.org/10.1002/anie.200905773.
Krawczyk, B., Ensle, P., Müller, W. M., and Süssmuth, R. D. (2012) Deuterium labeled peptides give insights into the directionality of class III lantibiotic synthetase LabKC, J. Am. Chem. Soc., 134, 9922-9925, doi: https://doi.org/10.1021/ja3040224.
Müller, W. M., Schmiederer, T., Ensle, P., and Süssmuth, R. D. (2010) In vitro biosynthesis of the prepeptide of type-III lantibiotic labyrinthopeptin A2 including formation of a C-C bond as a post-translational modification, Angew. Chem. Int. Ed. Engl., 49, 2436-2440, doi: https://doi.org/10.1002/anie.200905909.
Völler, G. H., Krawczyk, J. M., Pesic, A., Krawczyk, B., Nachtigall, J., and Süssmuth, R. D. (2012) Characterization of new class III lantibiotics–erythreapeptin, avermipeptin and griseopeptin from Saccharopolyspora erythraea, Streptomyces avermitilis and Streptomyces griseus demonstrates stepwise N-terminal leader processing, Chembiochem, 13, 1174-1183, doi: https://doi.org/10.1002/cbic.201200118.
Iorio, M., Sasso, O., Maffioli, S. I., Bertorelli, R., Monciardini, P., et al. (2014) A glycosylated, labionin-containing lanthipeptide with marked antinociceptive activity, ACS Chem. Biol., 9, 398-404, doi: https://doi.org/10.1021/cb400692w.
Chen, S., Xu, B., Chen, E., Wang, J., Lu, J., Donadio, S., Ge, H., and Wang, H. (2019) Zn-dependent bifunctional proteases are responsible for leader peptide processing of class III lanthipeptides, Proc. Natl. Acad. Sci. USA, 116, 2533-2538, doi: https://doi.org/10.1073/pnas.1815594116.
Kodani, S., Lodato, M. A., Durrant, M. C., Picart, F., and Willey, J. M. (2005) SapT, a lanthionine-containing peptide involved in aerial hyphae formation in the streptomycetes, Mol. Microbiol., 58, 1368-1380, doi: https://doi.org/10.1111/j.1365-2958.2005.04921.x.
Ferir, G., Petrova, M. I., Andrei, G., Huskens, D., Hoorelbeke, B., et al. (2013) The lantibiotic peptide labyrinthopeptin A1 demonstrates broad anti-HIV and anti-HSV activity with potential for microbicidal applications, PLoS One, 8, e64010, doi: https://doi.org/10.1371/journal.pone.0064010.
Goto, Y., Li, B., Claesen, J., Shi, Y., Bibb, M. J., and van der Donk, W. A. (2010) Discovery of unique lanthionine synthetases reveals new mechanistic and evolutionary insights, PLoS Biol., 8, e1000339, doi: https://doi.org/10.1371/journal.pbio.1000339.
Zhang, Q., Doroghazi, J. R., Zhao, X., Walker, M. C., and van der Donk, W. A. (2015) Expanded natural product diversity revealed by analysis of lanthipeptide-like gene clusters in actinobacteria, Appl. Environ. Microbiol., 81, 4339-4350, doi: https://doi.org/10.1128/AEM.00635-15.
Iftime, D., Jasyk, M., Kulik, A., Imhoff, J. F., Stegmann, E., Wohlleben, W., Süssmuth, R. D., and Weber, T. (2015) Streptocollin, a type IV lanthipeptide produced by Streptomyces collinus Tü 365, Chembiochem., 16, 2615-2623, doi: https://doi.org/10.1002/cbic.201500377.
Hegemann, J. D., and van der Donk, W. A. (2018) Investigation of substrate recognition and biosynthesis in class IV lanthipeptide systems, J. Am. Chem. Soc., 140, 5743-5754, doi: https://doi.org/10.1021/jacs.8b01323.
Rogers, L. A. (1928) The inhibiting effect of Streptococcus Lactis on Lactobacillus Bulgaricus, J. Bacteriol., 16, 321-325, doi: https://doi.org/10.1128/JB.16.5.321-325.1928.
Mattick, A. T. R., Hirsch, A., and Berridge, N. J. (1947) Further observations on an inhibitory substance (nisin) from lactic streptococci, Lancet, 250, 5-8, doi: https://doi.org/10.1016/S0140-6736(47)90004-4.
Van Heijenoort, Y., Gomez, M., Derrien, M., Ayala, J., and van Heijenoort, J. (1992) Membrane intermediates in the peptidoglycan metabolism of Escherichia coli: possible roles of PBP 1b and PBP 3, J. Bacteriol., 174, 3549-3557, doi: https://doi.org/10.1128/jb.174.11.3549-3557.1992.
Breukink, E., and de Kruijff, B. (2006) Lipid II as a target for antibiotics, Nat. Rev. Drug Discov., 5, 321-323, doi: https://doi.org/10.1038/nrd2004.
Piper, C., Draper, L. A., Cotter, P. D., Ross, R. P., and Hill, C. (2009) A comparison of the activities of lacticin 3147 and nisin against drug-resistant Staphylococcus aureus and Enterococcus species, J. Antimicrob. Chemother., 64, 546-551, doi: https://doi.org/10.1093/jac/dkp221.
Dischinger, J., Basi Chipalu, S., and Bierbaum, G. (2014) Lantibiotics: promising candidates for future applications in health care, Int. J. Med. Microbiol., 304, 51-62, doi: https://doi.org/10.1016/j.ijmm.2013.09.003.
Hsu, S.-T. D., Breukink, E., Tischenko, E., Lutters, M. A. G., de Kruijff, B., et al. (2004) The nisin–lipid II complex reveals a pyrophosphate cage that provides a blueprint for novel antibiotics, Nat. Struct. Mol. Biol., 11, 963-967, doi: https://doi.org/10.1038/nsmb830.
Hasper, H. E., de Kruijff, B., and Breukink, E. (2004) Assembly and stability of nisin-lipid II pores, Biochemistry, 43, 11567-11575, doi: https://doi.org/10.1021/bi049476b.
Märki, F., Hänni, E., Fredenhagen, A., and van Oostrum, J. (1991) Mode of action of the lanthionine-containing peptide antibiotics duramycin, duramycin B and C, and cinnamycin as indirect inhibitors of phospholipase A2, Biochem. Pharmacol., 42, 2027-2035, doi: https://doi.org/10.1016/0006-2952(91)90604-4.
Vance, J. E., and Tasseva, G. (2013) Formation and function of phosphatidylserine and phosphatidylethanolamine in mammalian cells, Biochim. Biophys. Acta, 1831, 543-554, doi: https://doi.org/10.1016/j.bbalip.2012.08.016.
Gbaguidi, B., Hakizimana, P., Vandenbussche, G., and Ruysschaert, J. M. (2007) Conformational changes in a bacterial multidrug transporter are phosphatidylethanolamine-dependent, Cell. Mol. Life Sci., 64, 1571-1582, doi: https://doi.org/10.1007/s00018-007-7031-0.
Epand, R. M., and Epand, R. F. (2009) Lipid domains in bacterial membranes and the action of antimicrobial agents, Biochim. Biophys. Acta, 1788, 289-294, doi: https://doi.org/10.1016/j.bbamem.2008.08.023.
Vestergaard, M., Berglund, N. A., Hsu, P. C., Song, C., Koldso, H., Schiott, B., and Sansom, M. S. P. (2019) Structure and dynamics of cinnamycin-lipid complexes: mechanisms of selectivity for phosphatidylethanolamine lipids, ACS Omega, 4, 18889-18899, doi: https://doi.org/10.1021/acsomega.9b02949.
Choung, S. Y., Kobayashi, T., Takemoto, K., Ishitsuka, H., and Inoue, K. (1988) Interaction of a cyclic peptide, Ro09-0198, with phosphatidylethanolamine in liposomal membranes, Biochim. Biophys. Acta, 940, 180-187, doi: https://doi.org/10.1016/0005-2736(88)90193-9.
Hasim, S., Allison, D. P., Mendez, B., Farmer, A. T., Pelletier, D. A., Retterer, S. T., Campagna, S. R., Reynolds, T. B., and Doktycz, M. J. (2018) Elucidating duramycin’s bacterial selectivity and mode of action on the bacterial cell envelope, Front. Microbiol., 9, 219, doi: https://doi.org/10.3389/fmicb.2018.00219.
Jones, A. M., and Helm, J. M. (2009) Emerging treatments in cystic fibrosis, Drugs, 69, 1903-1910, doi: https://doi.org/10.2165/11318500-000000000-00000.
Elvas, F., Stroobants, S., and Wyffels, L. (2017) Phosphatidylethanolamine targeting for cell death imaging in early treatment response evaluation and disease diagnosis, Apoptosis, 22, 971-987, doi: https://doi.org/10.1007/s10495-017-1384-0.
Emoto, K., Kobayashi, T., Yamaji, A., Aizawa, H., Yahara, I., Inoue, K., and Umeda, M. (1996) Redistribution of phosphatidylethanolamine at the cleavage furrow of dividing cells during cytokinesis, Proc. Natl. Acad. Sci. USA, 93, 12867-12872, doi: https://doi.org/10.1073/pnas.93.23.12867.
Zhao, M. (2011) Lantibiotics as probes for phosphatidylethanolamine, Amino Acids, 41, 1071-1079, doi: https://doi.org/10.1007/s00726-009-0386-9.
Hofmann, F. T., Szostak, J. W., and Seebeck, F. P. (2012) In vitro selection of functional lantipeptides, J. Am. Chem. Soc., 134, 8038-8041, doi: https://doi.org/10.1021/ja302082d.
Zhou, X. X., Li, W. F., Ma, G. X., and Pan, Y. J. (2006) The nisin-controlled gene expression system: construction, application and improvements, Biotechnol. Adv., 24, 285-295, doi: https://doi.org/10.1016/j.biotechadv.2005.11.001.
Aso, Y., Nagao, J.-I., Koga, H., Okuda, K.-I., Kanemasa, Y., Sashihara, T., Nakayama, J., and Sonomoto, K. (2004) Heterologous expression and functional analysis of the gene cluster for the biosynthesis of and immunity to the lantibiotic, nukacin ISK-1, J. Biosci. Bioeng., 98, 429-436, doi: https://doi.org/10.1016/S1389-1723(05)00308-7.
Böttiger, T., Schneider, T., Martínez, B., Sahl, H.-G., and Wiedemann, I. (2009) Influence of Ca2+ ions on the activity of lantibiotics containing a mersacidin-like lipid II binding motif, Appl. Environ. Microbiol., 75, 4427-4434, doi: https://doi.org/10.1128/aem.00262-09.
Li, Y. (2011) Recombinant production of antimicrobial peptides in Escherichia coli: a review, Protein Expr. Purif., 80, 260-267, doi: https://doi.org/10.1016/j.pep.2011.08.001.
Mesa-Pereira, B., Rea, M. C., Cotter, P. D., Hill, C., and Ross, R. P. (2018) Heterologous expression of biopreservative bacteriocins with a view to low cost production, Front. Microbiol., 9, 1654, doi: https://doi.org/10.3389/fmicb.2018.01654.
Si, T., Tian, Q., Min, Y., Zhang, L., Sweedler, J. V., van der Donk, W. A., and Zhao, H. (2018) Rapid screening of lanthipeptide analogs via in-colony removal of leader peptides in Escherichia coli, J. Am. Chem. Soc., 140, 11884-11888, doi: https://doi.org/10.1021/jacs.8b05544.
Geng, M., and Smith, L. (2018) Modifying the lantibiotic mutacin 1140 for increased yield, activity, and stability, Appl. Environ. Microbiol., 84, doi: https://doi.org/10.1128/AEM.00830-18.
Field, D., Connor, P. M., Cotter, P. D., Hill, C., and Ross, R. P. (2008) The generation of nisin variants with enhanced activity against specific gram-positive pathogens, Mol. Microbiol., 69, 218-230, doi: https://doi.org/10.1111/j.1365-2958.2008.06279.x.
Yoganathan, S., Sit, C. S., and Vederas, J. C. (2011) Chemical synthesis and biological evaluation of gallidermin-siderophore conjugates, Org. Biomol. Chem., 9, 2133-2141, doi: https://doi.org/10.1039/c0ob00846j.
Li, Q., Montalban-Lopez, M., and Kuipers, O. P. (2018) Increasing the antimicrobial activity of nisin-based lantibiotics against Gram-negative pathogens, Appl. Environ. Microbiol., 84, doi: https://doi.org/10.1128/AEM.00052-18.
Yang, X., Lennard, K. R., He, C., Walker, M. C., Ball, A. T., Doigneaux, C., Tavassoli, A., and van der Donk, W. A. (2018) A lanthipeptide library used to identify a protein–protein interaction inhibitor, Nat. Chem. Biol., 14, 375-380, doi: https://doi.org/10.1038/s41589-018-0008-5.
Nixon, A. E., Sexton, D. J., and Ladner, R. C. (2014) Drugs derived from phage display: from candidate identification to clinical practice, MAbs, 6, 73-85, doi: https://doi.org/10.4161/mabs.27240.
Urban, J. H., Moosmeier, M. A., Aumuller, T., Thein, M., Bosma, T., et al. (2017) Phage display and selection of lanthipeptides on the carboxy-terminus of the gene-3 minor coat protein, Nat. Commun., 8, 1500, doi: https://doi.org/10.1038/s41467-017-01413-7.
Hetrick, K. J., Walker, M. C., and van der Donk, W. A. (2018) Development and application of yeast and phage display of diverse lanthipeptides, ACS Cent. Sci., 4, 458-467, doi: https://doi.org/10.1021/acscentsci.7b00581.
Convery, N., and Gadegaard, N. (2019) 30 years of microfluidics, Micro Nano Eng., 2, 76-91, doi: https://doi.org/10.1016/j.mne.2019.01.003.
Fallah-Araghi, A., Baret, J. C., Ryckelynck, M., and Griffiths, A. D. (2012) A completely in vitro ultrahigh-throughput droplet-based microfluidic screening system for protein engineering and directed evolution, Lab. Chip., 12, 882-891, doi: https://doi.org/10.1039/c2lc21035e.
Mahler, L., Wink, K., Beulig, R. J., Scherlach, K., Tovar, M., et al. (2018) Detection of antibiotics synthetized in microfluidic picolitre-droplets by various actinobacteria, Sci. Rep., 8, 13087, doi: https://doi.org/10.1038/s41598-018-31263-2.
Terekhov, S. S., Smirnov, I. V., Stepanova, A. V., Bobik, T. V., Mokrushina, Y. A., et al. (2017) Microfluidic droplet platform for ultrahigh-throughput single-cell screening of biodiversity, Proc. Natl. Acad. Sci. USA, 114, 2550-2555, doi: https://doi.org/10.1073/pnas.1621226114.
Funding
The research was financially supported by the Russian Foundation for Basic Research (project no. 18-29-08054) and by the Russian Science Foundation (project no. 17-74-30019).
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
The authors declare no conflicts of interest in financial or any other sphere. This article does not contain any studies with human participants or animals performed by any of the authors.
Rights and permissions
About this article
Cite this article
Pipiya, S.O., Terekhov, S.S., Mokrushina, Y.A. et al. Engineering Artificial Biodiversity of Lantibiotics to Expand Chemical Space of DNA-Encoded Antibiotics. Biochemistry Moscow 85, 1319–1334 (2020). https://doi.org/10.1134/S0006297920110048
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1134/S0006297920110048