Abstract
Phenotypic and genetic heterogeneities are conserved features of prokaryotic populations. During periods of stress, this programmed diversity increases the likelihood that variants within the population will survive the adverse conditions, allowing for proliferation. Phenotypic heterogeneity can have a mutational or indeed a non-mutational basis as observed in bet-hedging strategies adopted by antibiotic-tolerant persister cells. Genetic variants can arise by phase variation (slip-strand mispairing, promoter inversions etc.), nucleotide polymorphisms resulting from replication errors or larger rearrangements such as deletions and insertions. In the face of selective pressures, these alterations may be neutral, beneficial or deleterious.
We recently described the genetic basis of tolerance to a normally toxic metabolite, d-serine (d-ser) in enterohaemorrhagic E. coli (EHEC). Here we summarize our work in the context of population dynamics, provide further discussion on the distinction between these tolerance mechanisms and the importance of heterogeneity for maximising adaptive potential.
Similar content being viewed by others
Enterohaemorrhagic E. coli displays significant growth arrest upon exposure to millimolar concentrations of d-ser and this is associated with activation of an unusual SOS-like response characterized by induction of RecA expression (Connolly and Roe 2016). Moreover, its primary colonization apparatus—the locus of enterocyte effacement (LEE)-encoded type 3 secretion system is transcriptionally repressed by D-ser (Connolly et al. 2015), further supporting the notion that EHEC has evolved to become particularly incompatible with exposure to this amino acid (Fig. 1a). In our recent work (O’Boyle et al. 2020), we examined the effect of repeated exposure of EHEC to physiological concentrations of d-ser that were able to inhibit growth, thereby promoting adaptive evolution and the development of tolerance to d-ser. Whole-genome sequencing (WGS) and transcriptomics were then used to reveal the genetic basis of tolerance.
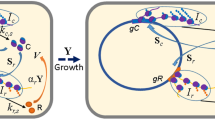
EHEC overcomes d-ser induced growth arrest by zipping the envelope shut or eating the poison. Upon exposure to millimolar d-ser (green ball and stick molecules) concentrations, wild-type EHEC (green cell) suffer growth arrest and activation of stress response (a). Growth arrest can be overcome by genetic disruption or transcriptional knock-down in inner membrane d-ser transporters (yellow cell), leading to zipping the cell envelope shut (b). Alternatively, mutations leading to the activation of the previously silent d-ser deaminase DsdA (red cell) allow EHEC to eat d-ser (the poison) and surmount the inhibition of growth normally caused by d-ser (c)
Complete d-ser tolerance was obtained through two distinct adaptive strategies: either by “zipping the cell envelope shut (Fig. 1b)” (blocking cytoplasmic accumulation of D-ser via the disruption of inner membrane transporters) or “eating the poison (Fig. 1c)” (constitutive activation of a d-ser deaminase that was normally locked in a silent state) (Fig. 1). “Zipping the envelope shut” was achieved primarily through inactivating single nucleotide polymorphisms (SNPs) in the d-ser transporter CycA (Cosloy 1973), allowing for growth in the presence of d-ser. These mutations appeared after as little as two successive 24-h batch cultures with distinct SNPs being observed in replicate evolution experiments. Importantly, inactivation of CycA alone resulted in partial tolerance and was subject to a refinement, observed after eight days of successive batch culture when stable, constitutive knock down in transcription of a second d-ser transporter SstT (Ogawa et al. 1998), resulted in growth rates in d-ser indistinguishable from those observed in the absence of d-ser (i.e., gain of complete tolerance). A similar refinement of an evolutionary innovation was seen during the evolution of aerobic citrate utilization under nutrient limitation in E. coli. Primary activation of citT by a rearrangement that led to control by the aerobically active rnk promoter facilitated the Cit + phenotype, but secondary refinement arising from a hyperactivating mutation in the dctA dicarboxylate transporter gene was required for full citrate utilisation (Quandt et al. 2014, 2015).
The alternative strategy identified in our study was to “eat the poison”. Diarrheagenic E. coli very rarely carry an intact dsdCXA locus (Moritz and Welch 2006; Connolly et al. 2015). Indeed, to date, we have never obtained a “wild-type” E. coli O157:H7 isolate containing both a functional-type three secretion system and the complete dsdCXA locus. Due to lack of the transcriptional activator DsdC, the deaminase encoded by dsdA does not respond to d-ser and remains silent (Connolly et al. 2015). We observed distinct genetic deletions allowing for full tolerance to d-ser by constitutive activation of d-ser deaminase via alternative upstream promoters. Again, the timing of emergence of these mutants was interesting. One full fitness mutant was acquired after only 2 days of successive batch culture while another was isolated on the tenth and final day of the experiment. Mutants were isolated on d-ser-containing plates to allow for discrimination of the tolerance phenotype. We believe that the mutation giving rise to full tolerance at day two was likely to have emerged after plating, as this mutant was not detected at any subsequent timepoint, and further full-fitness mutants adopting either transport or catabolic routes were not isolated until the eighth day of the experiment. Parallel adaptive evolution experiments with multiple populations can reveal differing transcriptional landscapes with convergent growth outcomes (Fong et al. 2005). In our work, we observed that this is also possible within a single population with full-fitness adaptive mutants displaying the “zip the envelope shut” and the “eat the poison” transcriptional profile and phenotype being isolated simultaneously on the final day of sampling. This is indicative of the co-existence of multiple adaptive lineages, a feature which has previously been described during long-term adaptation (Good et al. 2017) and highlights the inherent flexibility of E. coli populations in surmounting stress.
It should be noted that the sampling carried out in our study was far from exhaustive and it is likely that many additional evolutionary trajectories towards complete tolerance exist. The selective pressure imparted during differential identification of tolerant clones by plating on medium containing d-ser created desirable bias towards full-tolerance isolates, facilitating the analysis of fewer clones by WGS; however, this bias also prevented the analysis of neutral mutations. One can imagine further mutations within the population requiring more complex refinements before gaining a full tolerance phenotype. Deep sequencing of the mixed population during adaptation would more precisely reveal the dynamics of emergence and fixation of such mutations over time. Similarly, a comprehensive picture of the paths towards fitness can only be achieved through experimentation with replicate populations (Conrad et al. 2011). Our study involved only a short replicate experiment over a 48-h period as a proof of principle that distinct mutations in similar genes are selected for in multiple populations. It is quite likely that further replicates would identify novel adaptive routes to d-ser tolerance. For example, mutations were not observed in another transporter (YhaO/DlsT) that we have previously shown to functionally transport d-ser (Connolly et al. 2016). While further replicates may implicate disruption of DlsT in conferring tolerance to d-ser, our previous work showed that exposure to d-ser is required for maximal expression of DlsT and it is possible that without cytoplasmic accumulation of d-ser via CycA, expression of DlsT does not reach sufficient levels to cause growth inhibition in EHEC. The strength of selection has been shown to influence the evolutionary trajectory associated with overcoming a given selective pressure (Sanz-García et al. 2020). It should be noted that the 1 mM concentration used in our study was selected on the basis of being an approximation of the physiological concentration found in the bladder (Anfora et al. 2007), an environment unfavourable for EHEC. It is possible that different alleles would be adaptive in varying concentrations, an important consideration when trying to obtain a comprehensive appreciation of adaptive flexibility.
Finally, it should be noted that the adaptive routes described in our study confer advantages specifically in the in vitro system in which they were observed. As yet, it is unclear whether d-ser can exert such selective pressures within the host. It is also unclear what the consequences of these mutations would be for fitness within the host niche. Dietary consumption of compounds capable of inhibiting bacterial growth has been associated with the evolution of strains with enhanced pathogenicity (Collins et al. 2018). As adaptation to d-ser facilitated not only increased growth but also unexpectedly, resistance to repression of the type 3 secretion system essential for colonization, d-ser could apply selective pressure to pathogenic E. coli with important consequences for the outcome of infections.
References
Anfora AT, Haugen BJ, Roesch P et al (2007) Roles of serine accumulation and catabolism in the colonization of the murine urinary tract by Escherichia coli CFT073. Infect Immun 75:5298–5304. https://doi.org/10.1128/IAI.00652-07
Collins J, Robinson C, Danhof H et al (2018) Dietary trehalose enhances virulence of epidemic Clostridium difficile. Nature 553:291–294. https://doi.org/10.1038/nature25178
Connolly JPR, Roe AJ (2016) Intracellular D-serine accumulation promotes genetic diversity via modulated induction of RecA in enterohemorrhagic Escherichia coli. J Bacteriol 198:3318LP – 3328. https://doi.org/10.1128/JB.00548-16
Connolly JPR, Goldstone RJ, Burgess K et al (2015) The host metabolite d-serine contributes to bacterial niche specificity through gene selection. ISME J 9:1039–1051. https://doi.org/10.1038/ismej.2014.242
Connolly JPR, Gabrielsen M, Goldstone RJ et al (2016) A highly conserved bacterial d-serine uptake system links host metabolism and virulence. PLOS Pathog 12:e1005359. https://doi.org/10.1371/journal.ppat.1005359
Conrad TM, Lewis NE, Palsson BØ (2011) Microbial laboratory evolution in the era of genome-scale science. Mol Syst Biol 7:509. https://doi.org/10.1038/msb.2011.42
Cosloy SD (1973) D-serine transport system in Escherichia coli K-12. J Bacteriol 114:679 LP – 684
Fong SS, Joyce AR, Palsson BØ (2005) Parallel adaptive evolution cultures of Escherichia coli lead to convergent growth phenotypes with different gene expression states. Genome Res 15:1365–1372. https://doi.org/10.1101/gr.3832305
Good BH, McDonald MJ, Barrick JE et al (2017) The dynamics of molecular evolution over 60,000 generations. Nature 551:45–50. https://doi.org/10.1038/nature24287
Moritz RL, Welch RA (2006) The Escherichia coli argW-dsdCXA genetic island is highly variable, and E coli K1 strains commonly possess two copies of dsdCXA. J Clin Microbiol 44:4038–4048. https://doi.org/10.1128/JCM.01172-06
O’Boyle N, Connolly JPR, Tucker NP, Roe AJ (2020) Genomic plasticity of pathogenic Escherichia coli mediates d-serine tolerance via multiple adaptive mechanisms. Proc Natl Acad Sci. https://doi.org/10.1073/pnas.2004977117
Ogawa W, Kim Y-M, Mizushima T, Tsuchiya T (1998) Cloning and expression of the gene for the Na+-coupled serine transporter from Escherichia coli and characteristics of the transporter. J Bacteriol 180:6749LP – 6752
Quandt EM, Deatherage DE, Ellington AD et al (2014) Recursive genomewide recombination and sequencing reveals a key refinement step in the evolution of a metabolic innovation in Escherichia coli. Proc Natl Acad Sci 111:2217LP – 2222. https://doi.org/10.1073/pnas.1314561111
Quandt EM, Gollihar J, Blount ZD et al (2015) Fine-tuning citrate synthase flux potentiates and refines metabolic innovation in the Lenski evolution experiment. Elife 4:e09696. https://doi.org/10.7554/eLife.09696
Sanz-García F, Sánchez MB, Hernando-Amado S, Martínez JL (2020) Evolutionary landscapes of Pseudomonas aeruginosa towards ribosome-targeting antibiotic resistance depend on selection strength. Int J Antimicrob Agents. https://doi.org/10.1016/j.ijantimicag.2020.105965
Acknowledgements
We thank Eliza Wolfson (@eliza_coli) for providing the illustration used in this article. Funding was provided by a Tenovus Scotland Small Pilot Grant awarded to Nicky O’Boyle and a Biotechnology and Biological Sciences Research Council grant [BB/R006539/1] awarded to Andrew J. Roe.
Author information
Authors and Affiliations
Corresponding authors
Additional information
Communicated by M. Kupiec.
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
O’Boyle, N., Roe, A.J. Heterogeneity in populations of enterohaemorrhagic Escherichia coli undergoing d-serine adaptation. Curr Genet 67, 221–224 (2021). https://doi.org/10.1007/s00294-020-01130-7
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00294-020-01130-7