Abstract
Several studies now show that certain proteins exhibit selective preference toward liquid ordered (L\(_o\)) or toward liquid disordered (L\(_d\)) regions of the heterogeneous membrane and some of them have preference for the L\(_o\)-L\(_d\) interface. Spatially heterogenous organization of lipids, enriched in specific protein molecules, function as platforms for signaling and are involved in several other physiologically critical functions. In this review, we collate together some of the experimental observations of cases where proteins preferentially segregate into different phases and highlight the importance of these preferential localization in terms of underlying functions. We also try to understand the structural features and chemical makeup of the membrane-interacting motifs of these proteins. Finally, we put forth some preliminary analysis on class I viral fusion proteins, some of which are known to partition at the L\(_o\)-L\(_d\) interface, and through them we try to understand the evolutionary design principles of phase segregating proteins. Put together, this review summarizes the existing studies on preferential partitioning of proteins into different membrane phases while emphasizing the need to understand the molecular design-level features that can help us “engineer” functionally rich peptides and proteins with a programmed membrane partitioning.
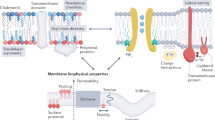
Graphic Abstract






Similar content being viewed by others
References
Baoukina S, Mendez-Villuendas E, Tieleman DP (2012) Molecular view of phase coexistence in lipid monolayers. J Am Chem Soc 134(42):17543. https://doi.org/10.1021/ja304792p
Barnoud J, Rossi G, Marrink SJ, Monticelli L (2014) Hydrophobic compounds reshape membrane domains. PLoS Comput Biol 10(10):e1003873
Barrera NP, Zhou M, Robinson CV (2013) The role of lipids in defining membrane protein interactions: insights from mass spectrometry. Trends Cell Biol 23(1):1. https://doi.org/10.1016/j.tcb.2012.08.007
Baumgart T, Hess ST, Webb WW (2003) Imaging coexisting fluid domains in biomembrane models coupling curvature and line tension. Nature 425(6960):821. https://doi.org/10.1038/nature02013
Beck-García K, Beck-García E, Bohler S, Zorzin C, Sezgin E, Levental I, Alarcón B, Schamel WW (2015) Nanoclusters of the resting T cell antigen receptor (TCR) localize to non-raft domains. Biochim Biophys Acta BBA 4(1853):802
Bretscher MS (1972) Asymmetrical lipid bilayer structure for biological membranes. Nat New Biol 236(61):11. https://doi.org/10.1038/newbio236011a0
Bretscher MS, Munro S (1993) Cholesterol and the Golgi apparatus. Science 261(5126):1280
Brown DA, Rose JK (1992) Sorting of GPI-anchored proteins to glycolipid-enriched membrane subdomains during transport to the apical cell surface. Cell 68(3):533. https://doi.org/10.1016/0092-8674(92)90189-j
Brügger B (2014) Lipidomics: analysis of the lipid composition of cells and subcellular organelles by electrospray ionization mass spectrometry. Annu Rev Biochem 83:79. https://doi.org/10.1146/annurev-biochem-060713-035324
Cebecauer M, Amaro M, Jurkiewicz P, Sarmento MJ, Sachl R, Cwiklik L, Hof M (2018) Membrane lipid nanodomains. Chem Rev 118(23):11259
Centi A, Dutta A, Parekh SH, Bereau T (2020) Inserting Small Molecules across Membrane Mixtures: Insight from the Potential of Mean Force. Biophys J 118(6):1321. https://doi.org/10.1016/j.bpj.2020.01.039
Colin J, Gregory-Pauron L, Lanhers MC, Claudepierre T, Corbier C, Yen FT, Malaplate-Armand C, Oster T (2016) Membrane raft domains and remodeling in aging brain. Biochimie 130:178. https://doi.org/10.1016/j.biochi.2016.08.014
Cornell CE, Mileant A, Thakkar N, Lee KK, Keller SL (2020) Direct imaging of liquid domains in membranes by cryo-electron tomography. Proc Natl Acad Sci USA 117(33):19713. https://doi.org/10.1073/pnas.2002245117
Cs Huang, Zhou J, Feng AK, Lynch CC, Klumperman J, DeArmond SJ, Mobley WC (1999) Nerve growth factor signaling in caveolae-like domains at the plasma membrane. J Biol Chem 274(51):36707
Cuddy LK, Winick-Ng W, Rylett RJ (2014) Regulation of the high-affinity choline transporter activity and trafficking by its association with cholesterol-rich lipid rafts. J Neurochem 128(5):725. https://doi.org/10.1111/jnc.12490
Danielli JF, Davson H (1935) A contribution to the theory of permeability of thin films. J Cell Comp Physiol 5(4):495. https://doi.org/10.1002/jcp.1030050409
Danylchuk DI, Sezgin E, Chabert P, Klymchenko AS (2020) Redesigning solvatochromic probe laurdan for imaging lipid order selectively in cell plasma membranes. Anal Chem 92(21):14798–14805
Diaz-Rohrer BB, Levental KR, Simons K, Levental I (2014) Membrane raft association is a determinant of plasma membrane localization. Proc Natl Acad Sci USA 111(23):8500. https://doi.org/10.1073/pnas.1404582111
Dinic J, Riehl A, Adler J, Parmryd I (2015) The T cell receptor resides in ordered plasma membrane nanodomains that aggregate upon patching of the receptor. Sci Rep 5:10082. https://doi.org/10.1038/srep10082
Dotti CG, Simons K (1990) Polarized sorting of viral glycoproteins to the axon and dendrites of hippocampal neurons in culture. Cell 62(1):63. https://doi.org/10.1016/0092-8674(90)90240-F
Dowhan W (2017) Understanding phospholipid function: why are there so many lipids? J Biol Chem 292(26):10755. https://doi.org/10.1074/jbc.X117.794891
Edidin M (2003) The state of lipid rafts: from model membranes to cells. Annu Rev Biophys Biomol Struct 32(1):257. https://doi.org/10.1146/annurev.biophys.32.110601.142439
Gorter E, Grendel F (1925) On Biomolecular layers of lipoids on the chromicytes of the blood. J Exp Med 41(4):439–443. https://doi.org/10.1084/jem.41.4.439
Gowrishankar K, Ghosh S, Saha S, Mayor RCS, Rao M (2012) Active remodeling of cortical actin regulates spatiotemporal organization of cell surface molecules. Cell 149(6):1353
Hanzal-Bayer MF, Hancock JF (2007) Lipid rafts and membrane traffic. FEBS Lett 581(11):2098. https://doi.org/10.1016/j.febslet.2007.03.019
Heberle FA, Doktorova M, Scott HL, Skinkle AD, Waxham MN, Levental I (2020) Direct label-free imaging of nanodomains in biomimetic and biological membranes by cryogenic electron microscopy. Proc Natl Acad Sci USA 117(33):19943. https://doi.org/10.1073/pnas.2002200117
Iyer SS, Negi A, Srivastava A (2020) Interpretation of phase boundary fluctuation spectra in biological membranes with nanoscale organization. J Chem Theory Comput 16(4):2736
Iyer SS, Srivastava A (2020) Degeneracy in molecular scale organization of biological membranes. Soft Matter 16(29):6752. https://doi.org/10.1039/D0SM00619J
Jolly MK, Ward C, Eapen MS, Myers S, Hallgren O, Levine H, Sohal SS (2018) Epithelial-mesenchymal transition, a spectrum of states: Role in lung development, homeostasis, and disease. Dev Dyn 247(3):346. https://doi.org/10.1002/dvdy.24541
Jy Jia, Lamer S, Schümann M, Schmidt MR, Krause E, Haucke V (2006) Quantitative proteomics analysis of detergent-resistant membranes from chemical synapses: evidence for cholesterol as spatial organizer of synaptic vesicle cycling. Mol Cell Proteom 5(11):2060
Kalappurakkal JM, Sil P, Mayor S (2020) Toward a new picture of the living plasma membrane. Protein Sci 29(6):1355. https://doi.org/10.1002/pro.3874
Kedia S, Ramakrishna P, Netrakanti PR, Jose M, Sibarita JB, Nadkarni S, Nair D (2020) Real-time nanoscale organization of amyloid precursor protein. Nanoscale 12:8200. https://doi.org/10.1039/D0NR00052C
King C, Sengupta P, Seo AY, Lippincott-Schwartz J (2020) ER membranes exhibit phase behavior at sites of organelle contact. Proc Natl Acad Sci USA 117(13):7225
Kleinzeller A, McIntosh TJ, Pohorille A, New MH, Schweighofer K, Wilson MA, Paula S, Deamer DW, Verkman A, Hoffmann EK, et al (1999) Membrane permeability: 100 years since Ernest Overton. Curr Top Mem 48:1e22
Kusumi A, Nakada C, Ritchie K, Murase K, Suzuki K, Murakoshi H, Kasai RS, Kondo J, Fujiwara T (2005) Paradigm shift of the plasma membrane concept from the two-dimensional continuum fluid to the partitioned fluid: high-speed single-molecule tracking of membrane molecules. Annu Rev Biophys Biomol Struct 34(1):351. https://doi.org/10.1146/annurev.biophys.34.040204.144637
Kyte J, Doolittle RF (1982) A simple method for displaying the hydropathic character of a protein. J Mol Biol 157(1):105
Langmuir I (1917) The constitution and fundamental properties of solids and liquids. J Franklin Inst 183(1):102. https://doi.org/10.1021/ja02268a002
Larsen JB, Jensen MB, Bhatia VK, Pedersen SL, Bjornholm T, Iversen L, Uline M, Szleifer I, Jensen KJ, Hatzakis NS et al (2015) Membrane curvature enables N-Ras lipid anchor sorting to liquid-ordered membrane phases. Nat Chem Biol 11(3):192. https://doi.org/10.1038/nchembio.1733
Larsen JB, Jensen MB, Bhatia VK, Pedersen SL, Bjørnholm T, Iversen L, Uline M, Szleifer I, Jensen KJ, Hatzakis NS et al (2015) Membrane curvature enables N-Ras lipid anchor sorting to liquid-ordered membrane phases. Nat Chem Biol 11(3):192. https://doi.org/10.1038/nchembio.1733
Levental I, Grzybek M, Simons K (2010) Greasing their way: lipid modifications determine protein association with membrane rafts. Biochemistry 49(30):6305. https://doi.org/10.1021/bi100882y
Levental ILK, Heberle F (2020) Lipid rafts: controversies resolved, mysteries remain. Trends Cell Biol 30(5):341. https://doi.org/10.1016/j.tcb.2020.01.009
Levental KR, Lorent JH, Lin X, Skinkle AD, Surma MA, Stockenbojer EA, Gorfe AA, Levental I (2016) Polyunsaturated lipids regulate membrane domain stability by tuning membrane order. Biophys J 110(8):1800. https://doi.org/10.1016/j.bpj.2016.03.012
Li L, Chen H, Wang M, Chen F, Gao J, Sun S, Li Y, Gao D (2017) NCAM-140 translocation into lipid rafts mediates the neuroprotective effects of GDNF. Mol Neurobiol 54(4):2739. https://doi.org/10.1007/s12035-016-9749-x
Lichtenberg D, Goñi FM, Heerklotz H (2005) Detergent-resistant membranes should not be identified with membrane rafts. Trends Biochem Sci 30(8):430. https://doi.org/10.1016/j.tibs.2005.06.004
Lin X, Gorfe AA, Levental I (2018) Protein partitioning into ordered membrane domains: insights from simulations. Biophys J 114(8):1936. https://doi.org/10.1016/j.bpj.2018.03.020
Lin Q, London E (2013) Altering hydrophobic sequence lengths shows that hydrophobic mismatch controls affinity for ordered lipid domains (rafts) in the multitransmembrane strand protein perfringolysin O. J Biol Chem 288(2):1340. https://doi.org/10.1074/jbc.M112.415596
Lingwood D, Simons K (2010) Lipid rafts as a membrane-organizing principle. Science 327(5961):46
Lorent JH, Diaz-Rohrer B, Lin X, Spring K, Gorfe AA, Levental KR, Levental I (2017) Structural determinants and functional consequences of protein affinity for membrane rafts. Nat Commu 8(1):1. https://doi.org/10.1038/s41467-017-01328-3
Lorent JH, Levental I (2015) Structural determinants of protein partitioning into ordered membrane domains and lipid rafts. Chem Phys Lipids 192:23. https://doi.org/10.1016/j.chemphyslip.2015.07.022
Lorent J, Levental K, Ganesan L, Rivera-Longsworth G, Sezgin E, Doktorova M, Lyman E, Levental I (2020) Plasma membranes are asymmetric in lipid unsaturation, packing and protein shape. Nat Chem Biol 16(6):644. https://doi.org/10.1038/s41589-020-0529-6
Magnani F, Tate CG, Wynne S, Williams C, Haase J (2004) Partitioning of the serotonin transporter into lipid microdomains modulates transport of serotonin. J Biol Chem 279(37):38770–38778
Mayor S, Rao M (2004) Rafts: scale-dependent, active lipid organization at the cell surface. Traffic 5(4):231. https://doi.org/10.1111/j.1600-0854.2004.00172.x
van Meer G, de Kroon AI (2011) Lipid map of the mammalian cell. J Cell Sci 124(1):5. https://doi.org/10.1242/jcs.071233
Miller JL (2018) Membrane phase demixing seen in living cells. Phys Today 71:21
Mouritsen O, Bloom M (1984) Mattress model of lipid-protein interactions in membranes. Biophys J 46(2):141. https://doi.org/10.1016/S0006-3495(84)84007-2
Nair D, Hosy E, Petersen JD, Constals A, Giannone G, Choquet D, Sibarita JB (2013) Super-resolution imaging reveals that AMPA receptors inside synapses are dynamically organized in nanodomains regulated by PSD95. J Neurosci 33(32):13204. https://doi.org/10.1523/JNEUROSCI.2381-12.2013
Nicolson GL (2014) The fluid-Mosaic model of membrane structure: still relevant to understanding the structure, function and dynamics of biological membranes after more than 40 years. Biochim Biophys Acta BBA 1838(6):1451
Perouansky M (2015) The Overton in Meyer-Overton: a biographical sketch commemorating the 150th anniversary of Charles Ernest Overton’s birth. Br J Anaesth 114(4):537. https://doi.org/10.1093/bja/aev069
Pike LJ (2006) Rafts defined: a report on the Keystone Symposium on Lipid Rafts and Cell Function. J Lipid Res 47(7):1597. https://doi.org/10.1194/jlr.E600002-JLR200
Pollard TD, Earnshaw WC, Lippincott-Schwartz J, Johnson G (2016) Cell biology E-book. Elsevier Health Sciences
Puchkov D, Haucke V (2013) Greasing the synaptic vesicle cycle by membrane lipids. Trends Cell Biol 23(10):493. https://doi.org/10.1016/j.tcb.2013.05.002
Raghu H, Sodadasu PK, Malla RR, Gondi CS, Estes N, Rao JS (2010) Localization of uPAR and MMP-9 in lipid rafts is critical for migration, invasion and angiogenesis in human breast cancer cells. BMC Cancer 10(1):1. https://doi.org/10.1186/1471-2407-10-647
Scheinpflug K, Wenzel M, Krylova O, Bandow JE, Dathe M, Strahl H (2017) Antimicrobial peptide cWFW kills by combining lipid phase separation with autolysis. Sci Rep 7(1):1
Sengupta P, Seo AY, Pasolli HA, Song YE, Johnson MC, Lippincott-Schwartz J (2019) A lipid-based partitioning mechanism for selective incorporation of proteins into membranes of HIV particles. Nat Cell Biol 21(4):452. https://doi.org/10.1038/s41556-019-0300-y
Sharma P, Varma R, Sarasij IRC, Gousset K, Krishnamoorthy G, Rao M, Mayor S (2004) Nanoscale organization of multiple GPI-anchored proteins in living cell membranes. Cell 116(4):577. https://doi.org/10.1016/S0092-8674(04)00167-9
Shaw TR, Ghosh S, Veatch SL (2020) Critical Phenomena in Plasma Membrane Organization and Function, Critical phenomena in plasma membrane organization and function
Shevchenko A, Simons K (2010) Lipidomics: coming to grips with lipid diversity. Nat Rev Mol Cell Biol 11(8):593. https://doi.org/10.1038/nrm2934
Sievers F, Wilm A, Dineen D, Gibson TJ, Karplus K, Li W, Lopez R, McWilliam H, Remmert M, Söding J et al (2011) Fast, scalable generation of high-quality protein multiple sequence alignments using Clustal Omega. Mol Syst Biol 7(1):539. https://doi.org/10.1038/msb.2011.75
Simons K, Ikonen E (1997) Functional rafts in cell membranes. Nature 387:4. https://doi.org/10.1038/42408
Simons K, Van Meer G (1988) Lipid sorting in epithelial cells. Biochemistry 27(17):6197. https://doi.org/10.1021/bi00417a001
Simons K, Vaz WL (2004) Model systems, lipid rafts, and cell membranes. Annu Rev Biophys Biomol Struct 33:269
Singer SJ, Nicolson GL (1972) The fluid mosaic model of the structure of cell membranes. Science 175(4023):720. https://doi.org/10.1126/science.175.4023.720
Stone MB, Shelby SA, Núñez MF, Wisser K, Veatch SL (2017) Protein sorting by lipid phase-like domains supports emergent signaling function in B lymphocyte plasma membranes. Elife 6:e19891. https://doi.org/10.7554/eLife.19891
Tisza MJ, Zhao W, Fuentes JS, Prijic S, Chen X, Levental I, Chang JT (2016) Motility and stem cell properties induced by the epithelial-mesenchymal transition require destabilization of lipid rafts. Oncotarget 7(32):51553
Tulodziecka K, Diaz-Rohrer BB, Farley MM, Chan RB, Di Paolo G, Levental KR, Waxham MN, Levental I (2016) Remodeling of the postsynaptic plasma membrane during neural development. Mol Biol Cell 27(22):3480. https://doi.org/10.1091/mbc.E16-06-0420
Vinklárek IS, Vel’as L, Riegerová P, Skála K, Mikhalyov I, Gretskaya N, Hof M, Šachl R (2019) Experimental evidence of the existence of interleaflet coupled nanodomains: an MC-FRET study. J Phys Chem Lett 10(9):2024. https://doi.org/10.1021/acs.jpclett.9b00390
Wang TY, Leventis R, Silvius JR (2000) Fluorescence-based evaluation of the partitioning of lipids and lipidated peptides into liquid-ordered lipid microdomains: a model for molecular partitioning into lipid rafts. Biophys J 79(2):919
Wang TY, Leventis R, Silvius JR (2001) Partitioning of lipidated peptide sequences into liquid-ordered lipid domains in model and biological membranes. Biochemistry 40(43):13031
Yang K, Han X (2016) Lipidomics: techniques, applications, and outcomes related to biomedical sciences. Trends Biochem Sci 41(11):954. https://doi.org/10.1016/j.tibs.2016.08.010
Yang ST, Kiessling V, Tamm LK (2016) Line tension at lipid phase boundaries as driving force for HIV fusion peptide-mediated fusion. Nat Commun 7(1):1. https://doi.org/10.1038/ncomms11401
Yang ST, Kreutzberger AJB, Kiessling V, Ganser-Pornillos BK, White JM, Tamm LK (2017) HIV virions sense plasma membrane heterogeneity for cell entry. Sci Adv 3(6):e1700338
Acknowledgements
A.S. thanks the Indian Institute of Science and the Ministry of Human Resource Development of India for the start up grant and the Department of Science and Technology of India for the early career grant. This research was also supported by the Department of Biotechnology, Government of India in the form of IISc-DBT partnership program. Support from FIST program sponsored by the Department of Science and Technology and UGC, India – Centre for Advanced Studies and Ministry of Human Resource Development, India is gratefully acknowledged by the authors.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
About this article
Cite this article
Bodosa, J., Iyer, S.S. & Srivastava, A. Preferential Protein Partitioning in Biological Membrane with Coexisting Liquid Ordered and Liquid Disordered Phase Behavior: Underlying Design Principles. J Membrane Biol 253, 551–562 (2020). https://doi.org/10.1007/s00232-020-00150-1
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00232-020-00150-1